Sahil Patel Week 2 Assignment
Sahil Patel Electronic Lab Notebook
Purpose of this assignment
The purpose of this assignment is to gain a deeper understanding of the experimental method of DNA microarrays and solving analytical problems using this new information.
Answer the following questions related to Chapter 4 of Campbell & Heyer (2003).
1. Choose two genes from Figure 4.6b and draw a graph to represent the change in transcription over time.
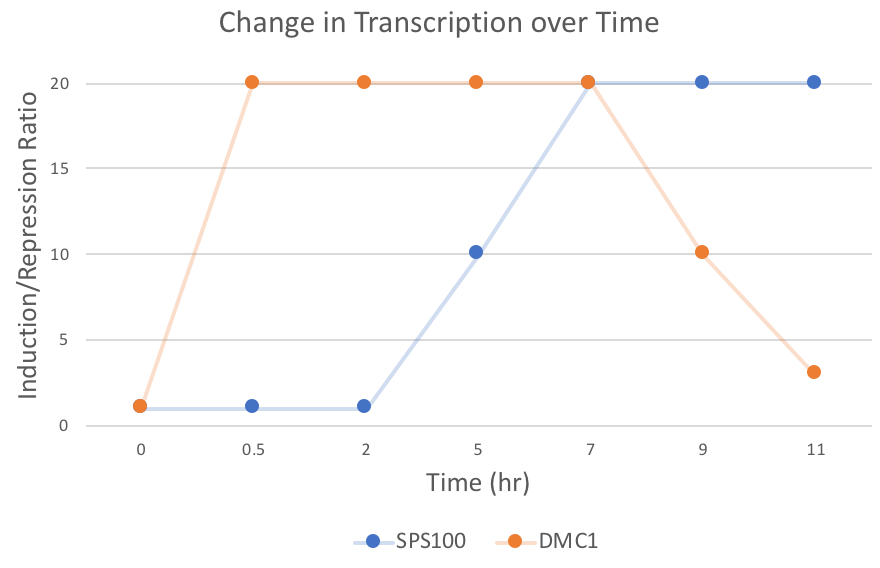
2. Look at Figure 4.7, which depicts the loss of oxygen over time and the transcriptional response of three genes. These data are the ratios of transcription for genes X, Y, and Z during the depletion of oxygen. Using the color scale from Figure 4.6, determine the color for each ratio in Figure 4.7b. (Use the nomenclature "bright green", "medium green", "dim green", "black", "dim red", "medium red", or "bright red" for your answers.)
- Gene X: Hour 1 - Black, Hour 3 - Dim Red, Hour 5 - Black, Hour 7 - Medium Green
- Gene Y: Hour 1 - Black, Hour 3 - Dim/Medium Red, Hour 5 - Black, Hour 7 - Bright Green
- Gene Z: Hour 1 - Black, Hour 3 - Dim Red, Hour 5 - Dim Red, Hour 7 - Dim/Medium Red
3. Were any of the genes in Figure 4.7b transcribed similarly? If so, which ones were transcribed similarly to which ones?
- Out of the three genes, only Genes X and Y transcribed similarly. Both followed the same trend of an initial increase in rate of transcription followed by a steep decrease due to the gradual loss of oxygen.
4. Why would most spots be yellow at the first time point? I.e., what is the technical reason that spots show up as yellow - where does the yellow color come from? And, what would be the biological reason that the experiment resulted in most spots being yellow?
- They yellow dots would show up in the first time point because it is the initial blot at t=0 and so no transcription has taken place yet as well as no oxygen depletion, thus a 1:1 ratio of Green:Red shows up as a yellow spot.
5. Go to the Saccharomyces Genome Database and search for the gene TEF4; you will see it is involved in translation. Look at the time point labeled OD 3.7 in Figure 4.12, and find the TEF4 spot. Over the course of this experiment, was TEF4 induced or repressed? Hypothesize why TEF4’s change in expression was part of the cell’s response to a reduction in available glucose (i.e., the only available food).
- TEF4 was repressed over the course of this experiment. This can be concluded because this gene had an increasingly bright green dot as the transcription process progressed. The reduction in its available glucose, its source of energy, slowed down its rate of gene expression, showing that glucose is a necessary component in order to stop the gene from being repressed.
6. Why would TCA cycle genes be induced if the glucose supply is running out?
- When glucose supply is running out, the TCA cycle genes would be induced in order to produce electron carriers and ATP ultimately leading to energy production.
7. What mechanism could the genome use to ensure genes for enzymes in a common pathway are induced or repressed simultaneously?
- The genome could use regulatory transcription factors that are able to induce or repress simultaneously in order to ensure that the desired outcome is established.
8. Consider a microarray experiment where cells deleted for the repressor TUP1 were subjected to the same experiment of a timecourse of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later timepoints (glucose depleted) are labeled red. What color would you expect the spots that represented glucose-repressed genes to be in the later time points of this experiment?
- The spots that represent the glucose-repressed genes would be green in the later time points in this experiments.
9. Consider a microarray experiment where cells that overexpress the transcription factor Yap1p were subjected to the same experiment of a timecourse of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later timepoints (glucose depleted) are labeled red. What color would you expect the spots that represented Yap1p target genes to be in the later time points of this experiment
- Similarly to question 8, at later timepoints after glucose has been depleted, the spots would also be red.
10. Using the microarray data, how could you verify that you had truly deleted TUP1 or overexpressed YAP1 in the experiments described in questions 8 and 9?
- You could verify the results by comparing the intensity on the microarray of the mutant and wild type (control). The mutant should be a much brighter red than the control.
Acknowledgments
I would like to acknowledge my homework partner, Brianna N. Samuels, for helping me with questions on organization. We met face to face in the life science building study room and also communicated via text.
Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
References
- Campbell, A.M, & Heeyer, L.J. (2003). "Chapter 4: Basic Research with DNA Microarrays," in Discovering Genomics, Proteomics, and Bloinformatics, Cold Spring Harbor Laboratory Press, pp.107-124
- Dahlquist, K. & Fitzpatrick, B. (2019). "BIOL388/S19: Week 2 Assignment Page" Week 2 Assignment Page
Assignment Links
BIOL368 Assignments
Week 1: Instructions, Class Journal and User Page
Week 2: Instructions, Class Journal and Individual Assignment
Week 3: Instructions, Class Journal and Individual Assignment
Week 4: Instructions, Class Journal and Individual Assignment
Week 5: Instructions, Class Journal and Individual Assignment
Week 6: Instructions, Class Journal and Individual Assignment
Week 8: Instructions, Class Journal and Individual Assignment
Week 9: Instructions, Class Journal and Individual Assignment
Week 10: Instructions, Class Journal and Individual Assignment
Week 11: Instructions, Class Journal and Individual Assignment
Week 13: Instructions, Class Journal and Individual Assignment
Week 14: Instructions, Class Journal and Individual Assignment
BIOL388 Assignments
Week 1: Instructions, Class Journal and User Page
Week 2: Instructions, Class Journal and Individual Assignment
Week 3: Instructions, Class Journal and Individual Assignment
Week 4/5: Instructions, Class Journal and Individual Assignment
Week 6: Instructions, Class Journal and Individual Assignment
Week 7: Instructions, Class Journal and Individual Assignment
Week 9: Instructions, Class Journal and Individual Assignment
Week 10: Instructions, Class Journal and Individual Assignment
Week 11: Instructions, Class Journal and Individual Assignment
Week 12: Instructions, Class Journal and Individual Assignment
Week 14/15: Instructions and Individual Assignment