Avalekander Week 6
Purpose
The purpose of this assignment is to run the gene regulatory network in GRNmap, analyze it, and then propose in silico experiments to be run using the model.
Methods
Creating the GRNmap Input Workbook
Now that the gene regulatory network that you want to model was identified, the next step is to generate the input Excel workbook that you will run in the GRNmap modeling software.
production_rates sheet
- The production rates were provided in a Microsoft Access database, which was downloaded.
- Then the values were copied and pasted one-by-one from the "production_rates" table, or perform a query to grab them as a group.
- To perform the query, the following steps were followed.
- The list of genes was imported to a new table in the database. The "External Data" tab was clicked and then the Excel icon with the "up" arrow on it was selected.
- The "Browse" button was clicked and the Excel file was selected containing the network that was used to upload to GRNsight.
- Click the button next to "Import the source data into a new table in the current database" and click "OK".
- In the next window, select the "network" worksheet, if it hasn't already been automatically selected for you. Click "Next".
- In the next window, make sure the "First Row Contains Column Headings" is checked. Click "Next".
- In the next window, the left-most column will be highlighted. Change the "Field Name" to "id" if it doesn't say that already. Click "Next".
- In the next window, select the button for "Choose my own primary key." and choose the "id" field from the drop down next to it. Click "Next".
- In the next field, make sure it says "Import to Table: network". Click Finish.
- In the next window do not save the import steps, so just click "Close".
- A table called "network" should appear in the list of tables at the left of the window.
- Go to the "Create" tab. Click on the icon for "Query Design".
- In the window that appears, click on the "network" table and click "Add". Click on the "production_rates" table and click "Add". Click "Close".
- The two tables should appear in the main part of the window. Tell Access which fields in the two tables correspond to each other. Click on the word "id" in the network table and drag your mouse to the "standard_name" field in the "production_rates" table, and release. A line will appear between those two words.
- Right-click on the line between those words and select "Join Properties" from the menu that appears. Select Option "2: Include ALL records from 'network' and only those records from 'production_rates' where the joined fields are equal." Click "OK".
- Click on the "id" word in the "network" table and drag it to the bottom of the screen to the first column next to the word "Field" and release.
- Click on the "production_rate" field in the "production_rates" table and drag it to the bottom of the screen to the second column next to the word "Field" and release.
- Right-click anywhere in the gray area near the two tables. In the menu that appears, select "Query Type > Make Table Query...".
- In the window that appears, name your table "production_rates_1". Make sure that "Current Database" is selected and Click "OK".
- Go to the "Query Tools: Menus" tab. Click on the exclamation point icon. A window will appear that tells how many rows you are pasting into a new table. Click "Yes".
- Your new "production_rates_1" table will appear in the list at the left. Double-click on that table name to open it.
- You can copy the data in this table and paste it back into your Excel workbook. Make sure that when you paste that you use "Paste Special > Paste values" so that the Access formatting doesn't get carried along. You can also choose to export this table to Excel going to the "External Data" tab and selecting the Excel icon with the arrow pointing to the right. Select the workbook you want to export the table to, making sure that "Preserve Access formatting" is not checked. Click "OK", click "Close".
- All genes should be listed in the same order in all the sheets in the Excel workbook.
degradation_rates sheet
- This sheet contains degradation rates for all genes in the network.
- Currently, the Dahlquist Lab is using data based on published mRNA half-life data from Neymotin et al. (2006).
- The half-life data values were converted to the degradation rates by taking the natural log of the half-life and dividing by 2.
- The sheet should contain two columns (from left to right) entitled "id", and "degradation_rate".
- The id is an identifier that the user will use to identify a particular gene.
- The "degradation_rate" column should then contain the absolute value of the degradation rate for the corresponding gene as described above, rounded to four decimal places.
- To obtain these values, you will use the same file, [Microsoft Access database, which you can Expression-and-Degradation-rate-database_2019.accdb that you used to obtain the production rates in the first worksheet.
Expression Data Sheets for Individual Yeast Strains
- Expression data was provided for dhap4, dzap1, dgln3, and the wt.
- Each strain will have its own sheet in the workbook.
- Each sheet should be given a unique name that follows the convention "STRAIN_log2_expression", where the word "STRAIN" is replaced by the strain designation, which will appear in the optimization_diagnostics sheet.
- The sheet should have the following columns in this order:
- "id": list of all genes. The genes should be listed in the same order in all the sheets in the Excel workbook.
- The next series of columns should contain the expression data for each gene at a given timepoint given as log2 ratios (log2 fold changes). The column header should be the time at which the data were collected, without any units. For example, the 15 minute timepoint would have a column header "15" and the 30 minute timepoint would have the column header "30". GRNmap supports replicate data for each of the timepoints. Replicate data for the same timepoint should be in columns immediately next to each other and have the same column headers.
- If data are provided for multiple strains, each strain should have data for the same timepoints, although the number of replicates can vary.
- Include the data for the 15, 30, and 60 minute timepoints, but not the 90 or 120 minute timepoints.
- The data used is contained in the Expression-and-Degradation-rate-database_2019.accdb file that you used to obtain the production and degradation rates.
network sheet
- The network derived from the YEASTRACT database for the Week 5 assignment was copied and pasted into this sheet directly. The description below just explains what is already in this worksheet.
- This sheet contains an adjacency matrix representation of the gene regulatory network.
- The columns correspond to the transcription factors and the rows correspond to the target genes controlled by those transcription factors.
- A “1” means there is an edge connecting them and a “0” means that there is no edge connecting them.
- The upper-left cell (A1) should contain the text “cols regulators/rows targets”. This text is there as a reminder of the direction of the regulatory relationships specified by the adjacency matrix.
- The rest of row 1 should contain the names of the transcription factors that are controlling the other genes in the network, one transcription factor name per column.
- The rest of column A should contain the names of the target genes that are being controlled by the transcription factors heading each of the columns in the matrix, one target gene name per row.
- The transcription factor names should correspond to the "id" in the other sheets in the workbook. They should be capitalized the same way and occur in the same order along the top and side of the matrix. The matrix needs to be symmetric, i.e., the same transcription factors should appear along the top and left side of the matrix. The genes should be listed in the same order in all the sheets in the Excel workbook.
- Each cell in the matrix should then contain a zero (0) if there is no regulatory relationship between those two transcription factors, or a one (1) if there is a regulatory relationship between them. Again, the columns correspond to the transcription factors and the rows correspond to the target genes controlled by those transcription factors.
network_weights sheet
- These are the initial guesses for the estimation of the weight parameters, w.
- Since these weights are initial guesses which will be optimized by GRNmap, the content of this sheet is identical to the "network" sheet.
optimization_parameters sheet
- The optimization_parameters sheet should have two columns (from left to right) entitled, "optimization_parameter" and "value".
- Copy this worksheet from the sample workbook provided. The only row that you need to modify is row 15, "Strain". Include just the strain designations for which you have a corresponding STRAIN_log2_expression sheet. If you don't have the dgln3, dhap4, or dzap1 expression sheets, then delete those from this row.
- Explanations of what the optimization_parameters mean:
- alpha: Penalty term weighting (from the L-curve analysis)
- kk_max: Number of times to re-run the optimization loop. In some cases re-starting the optimization loop can improve performance of the estimation.
- MaxIter: Number of times MATLAB iterates through the optimization scheme. If this is set too low, MATLAB will stop before the parameters are optimized.
- TolFun: How different two least squares evaluations should be before the program determines that it is not making any improvement
- MaxFunEval: maximum number of times the program will evaluate the least squares cost
- TolX: How close successive least squares cost evaluations should be before the program determines that it is not making any improvement.
- production_function: = Sigmoid (case-insensitive) if sigmoidal model, =MM (case-insensitive) if Michaelis-Menten model
- L_curve: =0 if an L-curve analysis should NOT be run or =1 if an L-curve analysis SHOULD be run. The L-curve analysis will automatically run sequential rounds of estimation for an array of fixed alpha values (0.8, 0.5, 0.2, 0.1,0.08, 0.05,0.02,0.01, 0.008, 0.005, 0.002, 0.001, 0.0008, 0.0005, 0.0002, and 0.0001). GRNmap makes a copy of the user's selected input workbook and changes alpha to the first alpha in the list. The estimation runs and the resulting parameter values are used as the initial guesses for the next round of estimation with the next alpha value. This process repeats until all alpha values have been run. New input and output workbooks are generated for each alpha value, although currently, the graphs are only saved for the last run.
- estimate_params =1 if want to estimate parameters and =0 if the user wants to do just one forward run
- make_graphs =1 to output graphs; =0 to not output graphs
- fix_P =1 if the user does not want to estimate the production rate, P, parameter, just use the initial guess and never change; =0 to estimate
- fix_b =1 if the user does not want to estimate the b parameter, just use the initial guess and never change; =0 to estimate
- expression_timepoints: A row containing a list of the time points when the data was collected experimentally. Should correspond to the timepoint column headers in the STRAIN_log2_expression sheets.
- Strain: A row containing a list of all of the strains for which there is expression data in the workbook. Should correspond to the "STRAIN" portion of the names of the STRAIN_log2_expression sheets for each strain. Note that GRNmap will run the model for the wild type network (all genes present in the network) and for networks where the gene deleted from the designated STRAIN has been deleted from the network.
- simulation_timepoints: A row containing a list of the time points at which to evaluate the differential equations to generate the simulated data. This does not need to correspond to the actual measurement times, but should be in the same units (e.g. minutes).
threshold_b sheet
- These are the initial guesses for the estimation of the threshold_b parameters.
- There should be two columns.
- The left-most column should contain the header "id" and list the standard names for the genes in the model in the same order as in the other sheets.
- The second column should have the header "threshold_b" and should contain the initial guesses, we are going to use all 0.
- Upload the excel sheets onto box.
Dynamical Systems Modeling of your Gene Regulatory Network
The software used to run the model is called GRNmap, which stands for Gene Regulatory Network Modeling and Parameter Estimation. It is written in MATLAB and can be run from code or run as a stand-alone executable if you don't have MATLAB installed.
- To run GRNmap from code, you must have MATLAB R2014b installed on your computer.
- Download the GRNmap v1.10 code from the GRNmap Downloads page.
- Unzip the file. (Right-click, 7-zip > Extract here)
- Launch MATLAB R2014b.
- Open GRNmodel.m, which will be in the directory that you unzipped GRNmap-1.10 > matlab
- Click the Run button (green "play" arrow).
- You will be prompted to select your input workbook.
- Download the GRNmap v1.10 executable from the GRNmap Downloads page.
- This is a direct link to start downloading (688 MB)-->
- An optimization diagnostics graphic will open that shows the progress of the estimation.
- When the run is over, expression plots will display.
- Output .xlsx and .mat files will be saved in the same folder as your input folder, along with .jpg files containing the optimization diagnostic and individual expression plots. Save these files.
- Upload the output .xlsx file into GRNsight to visualize the results.
Analyzing the Modeling Results
In class on February 28/March 5, the modeling results will be looked at and we will discuss how to analyze them. We will discuss:
- LSE/minLSE ratio
- MSE's and expression plots for individual genes in relation to their ANOVA p values
- Visualization of the weighted graph with GRNsight
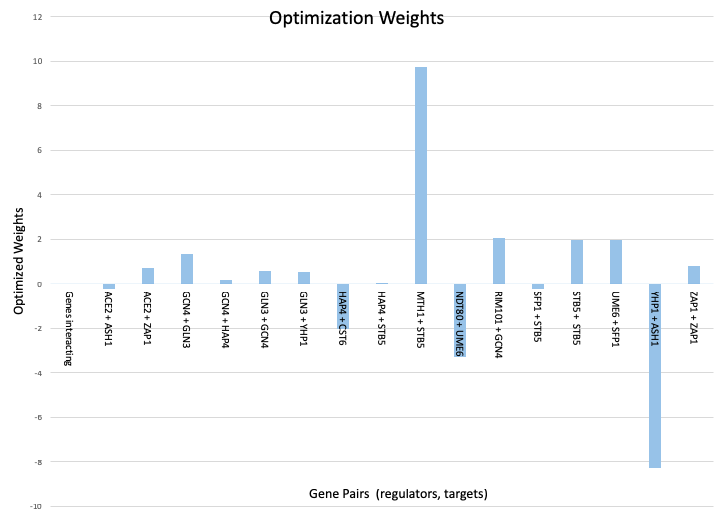
- Making bar charts to give a graphical representation of the parameter values.
Based on these analyses, a in silico experiment will be proposed that can be done with the model. A presentation discussing the results in a research presentation will be done in Week 9. Some ideas are:
- For our initial runs, we estimated all three parameters w, P, and b.
- How do the modeling results change if P is instead fixed and w and b are estimated?
- How do the modeling results change if b is fixed and w and P are estimated?
- How do the modeling results change if P and b are fixed, and only w is estimated?
- For our initial runs, we included all three microarray datasets, wt, Δgln3, and Δhap4.
- What happens to the results if we base the estimation on just two strains (wt + one deletion strain)?
- What happens to the results if we base the estimation on just the wt strain data?
- When viewing the modeling results in GRNsight, you may determine that one or more genes in the network does not appear to be doing much.
- What happens to the modeling results if you delete this gene from the network and re-run the model (remember you will have to delete references to this gene in all worksheets of the input file).
- You also might think that a particular edge (regulatory relationship) is not needed. What happens if you delete that edge?
- What happens if you include the t90 and t120 expression data?
Results
An Excel sheet containing parameter estimates (Pi, w, b), production rates, degradation rates, expression estimates, and threshold values was created and run in GRNmap. The output Excel sheet was then run in GRNsight to create the weighted regulatory matrices for further analysis in week 7.
GRNmap output folder: https://lmu.box.com/s/3jnc2ydauqvr6l3b31kgnuw1fc4gwys4
Excel sheet with bar graph: https://lmu.box.com/s/jrv4bmilxnigkoubimoa6wetmzf7pfpu
Conclusion
The networks were successfully run in GRNmap and all of the outputs were saved for further analysis in week 7.
Acknowledgements
I would like to acknowledge my homework partners who also worked with the dHAP4 strain, Desiree, and Brianna. Additionally, Dr. Dahlquist helped me multiple times via email and also helped me correct any errors so that GRN map could successfully recognize and run my excel sheet. Additionally, she reran the GRNmap software when my computer froze and was unable to save any of the results.
Except for what is noted above, this individual journal entry was completed by me and not copied from another source. Avalekander (talk) 19:29, 28 February 2019 (PST)
References
- Dahlquist, K. & Fitpatrick, B. (2019). "BIOL388/S19: Week 6" Biomathematical Modeling, Loyola Marymount University. Accessed from:Week 6 Assignment Page
Template
- Template: Ava Lekander
- User Page: Ava Lekander
- Journal Entries:
- Assignment Pages:
- Class Journal Pages:
- Biology 388 Home Page: BIOL 388 Class Page