Avalekander Week 2
Week 2 Journal
- User Page: Ava Lekander
- Journal Entries: Journal Entry Week 1
- Assignment Page:Assignment Week 2
- Class Journal Page: Class Journal Week 1
- Biology 388 Home Page: BIOL 388 Class Page
Purpose
- The purpose of this assignment is to learn about the new technology behind DNA microarrays and it's applications to molecular biology, as well as human health. The reading and figures associated also display evidence of how biology and mathematics can be used together in order to make and transform complex data into something easier to comprehend.
Answers to Discovery Questions from Chapter 4
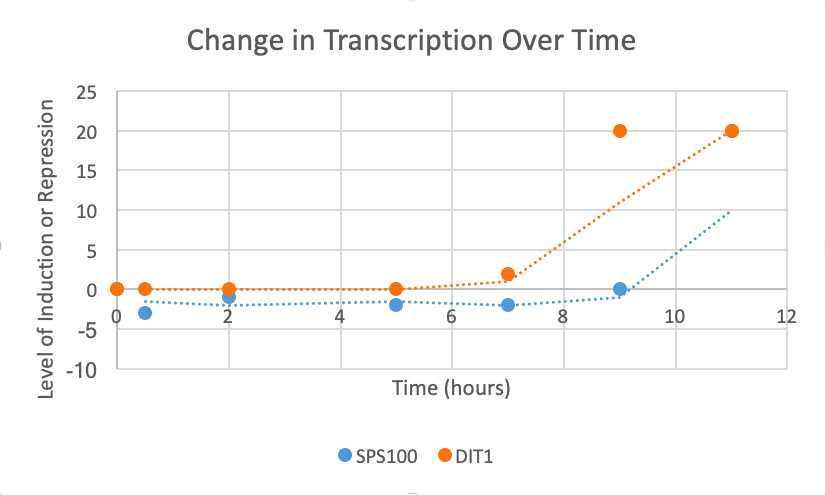
Choose two genes from Figure 4.6b (PDF of figures on Brightspace) and draw a graph to represent the change in transcription over time. You can either create your plot in Excel and put the image up on your wiki page or you can do it by hand and upload a picture or scan.
Look at Figure 4.7, which depicts the loss of oxygen over time and the transcriptional response of three genes. These data are the ratios of transcription for genes X, Y, and Z during the depletion of oxygen. Using the color scale from Figure 4.6, determine the color for each ratio in Figure 4.7b. (Use the nomenclature "bright green", "medium green", "dim green", "black", "dim red", "medium red", or "bright red" for your answers.)
- Gene x: 1 hr-black,3 hr- dim red, 5 hr- black, 9 hr- dim green
- Gene y: 1 hr-black, 3 hr- dim red, 5 hr- dim green, 9 hr- bright green
- Gene z: 1 hr-black, 3 hr- dim red, 5 hr- dim red, dim red
Were any of the genes in Figure 4.7b transcribed similarly? If so, which ones were transcribed similarly to which ones?
- After the nine hour mark none of the three genes had transcribed similarly as gene x was a dim green, y a bright green, and z a dim red. X and y did transcribe more similarly than z though as the red color in z signified increased transcription in the experimental lack of oxygen whereas both x and y had reduced transcription in the experimental.
Why would most spots be yellow at the first time point? I.e., what is the technical reason that spots show up as yellow - where does the yellow color come from? And, what would be the biological reason that the experiment resulted in most spots being yellow?
- The spots were yellow to begin with at the start of the experiment because not enough time had passed for a change to occur in transcription rates and thus the depletion of oxygen had not taken its toll yet. There has not been any change in the gene's transcription between control and experimental when the color is yellow. The yellow color signifies a perfect merging of green and red in a 1:1 ratio.
Go to the Saccharomyces Genome Database and search for the gene TEF4; you will see it is involved in translation. Look at the time point labeled OD 3.7 in Figure 4.12, and find the TEF4 spot. Over the course of this experiment, was TEF4 induced or repressed? Hypothesize why TEF4’s change in expression was part of the cell’s response to a reduction in available glucose (i.e., the only available food).
- TEF4 was repressed, which we know based on the green color that it turned shown in fig. 12. If this TEF4 is involved in translation but there is no food source available then it makes sense why this response occurred to conserve energy by not conducting translation.
Why would TCA cycle genes be induced if the glucose supply is running out?
- Energy metabolism genes as a group were induced while the protein production genes were repressed. This is the coordinated regulation of genes that occurs when environmental factors are sensed. The gene encoding pyruvate decarboxylase was repressed, whereas acetyl carboxylase was induced which shifts pyruvate away from acetaldehyde and toward oxaloacetate and the TCA cycle.
What mechanism could the genome use to ensure genes for enzymes in a common pathway are induced or repressed simultaneously?
- Combinatorial control (regulatory transcription factors) can be used by yeast as it is a eukaryote to regulate expression or the phenomenon of 'guilt by association.'
Consider a microarray experiment where cells deleted for the repressor TUP1 were subjected to the same experiment of a timecourse of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later timepoints (glucose depleted) are labeled red. What color would you expect the spots that represented glucose-repressed genes to be in the later time points of this experiment?
- I would expect the genes of later time points to be red because glucose levels would be low so glucose-repressed genes would be induced.
Consider a microarray experiment where cells that overexpress the transcription factor Yap1p were subjected to the same experiment of a timecourse of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later timepoints (glucose depleted) are labeled red. What color would you expect the spots that represented Yap1p target genes to be in the later time points of this experiment?
- Cells that overexpress Yap1p would be red later in the experiment because a lot of transcription had taken place and glucose levels low.
Using the microarray data, how could you verify that you had truly deleted TUP1 or overexpressed YAP1 in the experiments described in questions 8 and 9?
- To verify, one could compare the experimental results with the wild type controls and determine if they appeared similar or different. The experimental results should be mostly red colored rather than green.
Acknowledgements
- I would like to acknowledge my homework partner, Leanne with whom I discussed a few of the questions with via text message.
- Additionally, Desireee and I talked about the assignment and helped one another with the later questions.
- I would also like to acknowledge Dr. Dahlquist for answering a question about the second discovery question of this assignment, as well as responding to an email about the format of the citation needed.
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source. Avalekander (talk) 10:33, 27 January 2019 (PST)
References
- Campbell, A.M. and Heyer, L.J. (2003), “Chapter 4: Basic Research with DNA Microarrays”, in Discovering Genomics, Proteomics, and Bioinformatics, Cold Spring Harbor Laboratory Press, pp. 107-124.
- Dahlquist, K. & Fitzpatrick, B.G. (2019, January). BIOL388/S19:Week 2. BIOL 388-01: Biomathematical Modeling MATH 388-01: Survey of Biomathematics Loyola Marymount University. Retrieved from https://openwetware.org/wiki/BIOL388/S19:Week_2
Template
- User Page: Ava Lekander
- Journal Entries:
- Assignment Pages:
- Class Journal Pages:
- Biology 388 Home Page: BIOL 388 Class Page