Avalekander Week 4/5
Purpose
Through using Microsoft excel to perform data analysis, ANOVA tests, and Bonferroni, and Benjamini & Hochbergthe p-value corrections, significant changes in gene expression can be underscored for further analysis of specific genes.
Materials & Methods
- Filename: BIOL388_S19_microarray-data_dHAP4AL
- Strain: Δhap4
- Timepoints and replicates:
- t15min: 4 replicates
- t30min: 4 replicates
- t60min: 4 replicates
- t90min: 3 replicates
- t120min: 3 replicates
- The timepoints t15, t30, & t60 experienced cold shock at 13°C
- Points t90 and t120 experienced cold shock at 13°C followed by 30 or 60 minutes of recovery at 30°C.
Statistical Analysis Part 1: Within-stain ANOVA
Purpose: The purpose is to determine if any genes had a gene expression change that was significantly different than zero.
- The worksheet dHAP4_ANOVA was created
- The first three columns containing the "MasterIndex", "ID", and "Standard Name" from the "Master_Sheet" worksheet created by Dr. Dahlquist and Dr. Fitzpatrick for the dHAP4 strain were copied and pasted into the new worksheet.
- At the top of the first columns to the right of the data, five column headers labeled dHAP4_AvgLogFC_(TIME) where (TIME) was 15, 30, etc.
- In the cell below the dHAP4_AvgLogFC_t15 header, formula =AVERAGE(D2:G2) was typed. This was done for each time point t15-120 minutes.
- Then, in the first empty column to the right of the dHAP4_AvgLogFC_t120 calculation, the header was created titled dHAP4_ss_HO.
- In the first cell below this header, type =SUMSQ(D2:W2)to square the sum of the genes. The rest of the column was then autofilled.
- In the next empty column to the right of dHAP4_ss_HO, the column header dHAP4_ss_(TIME) was input for each time point. The formula =SUMSQ(D2:G2)-COUNTA(D2:G2)*X2^2 was used for dHAP4 at each time point minutes. Remaining genes and time points were autofilled in Excel.
- Make a note of how many data points you have at each time point. The two points at t90 and t120 only had three data points.
- The
COUNTAfunction counts the number of cells in the specified range that have data in them (i.e., does not count cells with missing values).
- The
- In the first column to the right of dHAP4_ss_t120, the column header dHAP4_SS_full was created. The sum for the range of cells containing "ss" for dHAP4 was calculated using the formula =SUM(AD2:AH2)
- In the next two columns to the right, the headers dHAP4_Fstat and dHAP4_p-value were made.
- N should be 18 in the formula.
- Under the dHAP4_Fstat column, the formula =((18-5)/5)*(AC2-AI2)/AI2) was used for gene dHAP4. Remaining genes were autofilled in Excel.
- In the first cell below the (STRAIN)_p-value header, =FDIST(AJ2,5,20-5) was the equation used for gene dHAP4. Remaining genes were autofilled in Excel.
Sanity check
- Click on cell A1 and click on the Data tab. Select the Filter icon (looks like a funnel). Little drop-down arrows should appear at the top of each column. This will enable us to filter the data according to criteria we set.
- Click on the drop-down arrow on your dHAP4_p-value column. Select "Number Filters". In the window that appears, set a criterion that will filter your data so that the p value has to be less than or equal to 0.05.
- Excel will now only display the rows that correspond to data meeting that filtering criterion. A number will appear in the lower left hand corner of the window giving you the number of rows that meet that criterion.
Calculate the Bonferroni and p-value Correction
- Label the next two columns to the right with the label, dHAP4_Bonferroni_p-value
- Under the first column insert the formula =dHAP4_p-value>*6189
- Corrected p-values greater than 1 were replaced with the number 1 using the formula =IF(AL2>1,1,AL2) for dHAP4 in the second dHAP4_Bonferroni_p-value column. The remaining genes were autofilled in Excel.
Calculate the Benjamini & Hochberg p-value Correction
- A worksheet was added named dHAP4_ANOVA_B-H
- Copy and paste the "MasterIndex", "ID", and "Standard Name" columns from the previous worksheet into the first two columns of the new worksheet using the paste special feature. Also, copy the unadjusted p-values from the ANOVA worksheet and paste into Column D.
- Sort the columns A-D by ascending values on Column D. Click the sort button from A to Z on the toolbar, in the window that appears, sort by column D, in ascending order.
- Type the header "Rank" in cell E1. Create a series of numbers in ascending order from 1 to 6189 in this column by typing 1 in E2 and then 2 into E3 and double-clicking the plus sign in the corner to fill the column with values 1-6189. This will rank the p-values from smallest to largest.
- Now calculate the Benjamini and Hochberg p-value correction by typing dHAP4_B-H_p-value in cell F1.
- Type the following formula in cell F2: =(D2*6189)/E2 and copy that equation to the entire column.
- Type dHAP4_B-H_p-value into cell G1
- Type the following formula into cell G2 =IF(F2>1,1,F2) and copy that equation to the entire column
- Select columns A through G and sort them by your MasterIndex in Column A in ascending order.
- Copy column G and use the paste special option to paste it into the next column on the right of the ANOVA sheet
- Upload the excel sheet onto box
Clustering and GO Term Enrichment with stem
- Prepare the microarray data file for loading into STEM.
- A new worksheet was inserted into the Excel workbook, and named dHAP4_stem.
- All of the data from the dHAP4_ANOVA worksheet was pasted using paste special > paste values into your dHAP4_stem worksheet.
- Some columns were renamed such as Master_Index to Spot and Column B renamed Gene Symbol. The column named Standard_Name was removed.
- Data on the B-H corrected p value was filtered to be > 0.05.
- Once the data was been filtered, all of the rows (except for your header row) were selected and then deleted by choosing "Delete Row" from the context menu. Undo the filter.
- All of the data columns were removed except for the Average Log Fold change columns for each timepoint.
- Data columns were renamed with just the time and units (for example, 15m, 30m, etc.).
- Save work and then save this spreadsheet as Text (Tab-delimited) (*.txt).
- STEM software was downloaded and extracted. Click here to go to the STEM web site.
- Click on the download link and download the
stem.zipfile to your Desktop. - Unzip the file by right clicking on the file icon and selecting the menu item 7-zip > Extract Here.
- Download the Gene Ontology and yeast GO annotations and place them in the folder opened.
- Click here to download the file "gene_ontology.obo".
- Click here to download the file "gene_association.sgd.gz".
- Inside the folder, launch the STEM program.
- Click on the download link and download the
- Running STEM
- In section 1 (Expression Data Info) of the the main STEM interface window, click on the browse button to navigate to and select the file.
- Click on the button No normalization/add 0.
- Check the box next to Spot IDs included in the data file.
- In section 2 (Gene Info) of the main STEM interface window, leave the default selection for the three drop-down menu selections for Gene Annotation Source, Cross Reference Source, and Gene Location Source as "User provided".
- Click the "Browse" button to the right of the "Gene Annotation File" item. Browse to your "stem" folder and select the file "gene_association.sgd.gz" and click Open.
- In section 3 (Options) of the main STEM interface window, make sure that the Clustering Method says "STEM Clustering Method" and do not change the defaults for Maximum Number of Model Profiles or Maximum Unit Change in Model Profiles between Time Points.
- In section 4 (Execute) click on the yellow Execute button to run STEM.
- A new window will open called "All STEM Profiles (1)". The colored boxes represent the significant transcription factors.
- Click on the button that says "Interface Options...". At the bottom of the Interface Options window that appears below where it says "X-axis scale should be:", click on the button that says "Based on real time". Then close the Interface Options window.
- Take a screenshot to save.
- Click on each of the Significant profiles to open a window showing a more detailed plot containing all of the genes in that profile.
- Take a screenshot of each of the individual profile windows and save the images in your PowerPoint presentation.
- At the bottom of each profile window, there are two yellow buttons "Profile Gene Table" and "Profile GO Table". For each of the profiles, click on the "Profile Gene Table" button to see the list of genes belonging to the profile. In the window that appears, click on the "Save Table" button and save the file to your desktop.
- Upload these files to OpenWetWare and link to them on your individual journal page (zip them together).
- For each of the significant profiles, click on the "Profile GO Table" to see the list of Gene Ontology terms belonging to the profile. In the window that appears, click on the "Save Table" button and save the file to your desktop.
- Upload these files to OpenWetWare and link to them on your individual journal page. (Note that it will be easier to zip all the files together and upload them as one file).
- In section 1 (Expression Data Info) of the the main STEM interface window, click on the browse button to navigate to and select the file.
- Analyzing and Interpreting STEM Results
- Select one of the profiles you saved in the previous step for further intepretation of the data.
- Open the GO list file saved for this profile in Excel. This list shows all of the Gene Ontology terms that are associated with genes that fit this profile. Select the third row and then choose from the menu Data > Filter > Autofilter. Filter on the "p-value" column to show only GO terms that have a p value of < 0.05.
- 6 Gene Ontology terms were selected from the filtered list (either p < 0.05 or corrected p < 0.05).
- Definitions for each of the terms were found at http://geneontology.org.
- Select one of the profiles you saved in the previous step for further intepretation of the data.
Using YEASTRACT to Infer which Transcription Factors Regulate a Cluster of Genes
Purpose: In the previous analysis using STEM, a number of gene expression profiles were found that grouped genes based on the similarity of gene expression changes over time. The YEASTRACT data based will be used to explore whether the genes share the same expression pattern because they are regulated by the same of transcription factors.
- Open the gene list in Excel for the one of the significant profiles from your stem analysis. Choose a cluster with a clear cold shock/recovery up/down or down/up pattern. You should also choose one of the largest clusters.
- Copy the list of gene IDs onto your clipboard.
- Launch a web browser and go to the YEASTRACT database.
- On the left panel of the window, click on the link to Rank by TF.
- Paste the list of genes from the cluster into the box labeled ORFs/Genes.
- Check the box for Check for all TFs.
- Accept the defaults for the Regulations Filter (Documented, DNA binding plus expression evidence)
- Do not apply a filter for "Filter Documented Regulations by environmental condition".
- Rank genes by TF using: The % of genes in the list and in YEASTRACT regulated by each TF.
- Click the Search button.
- Upload the Excel file to Box.
- Select from the list of significant transcription factors, which ones will be used to run the model. Add GLN3, HAP4, and ZAP1 if they are not in the list.
- Go back to the YEASTRACT database and follow the link to Generate Regulation Matrix.
- Copy and paste the list of transcription factors identified (plus HAP4, GLN3, and ZAP1) into both the "Transcription factors" field and the "Target ORF/Genes" field.
- Use the "Regulations Filter" options of "Documented", "Only DNA binding evidence"
- Generate.
- In the results window that appears, click on the link to the "Regulation matrix (Semicolon Separated Values (CSV) file)" that appears and save it.
Visualizing Gene Regulatory Networks with GRNsight
Purpose: Analyze the regulatory matrix files generated above in Excel and visualize them using GRNsight to determine which one will be appropriate to pursue further in the modeling.
- First, properly format the output files from YEASTRACT.
- Open the file in Excel. To fix the formatting, Select the entire Column A. Then go to the "Data" tab and select "Text to columns". In the Wizard that appears, select "Delimited" and click "Next". In the next window, select "Semicolon", and click "Next". In the next window, leave the data format at "General", and click "Finish".
- Save this file in Microsoft Excel workbook format (.xlsx).
- Select the menu item File > Open and select the regulation matrix .xlsx file that has the "network" worksheet in it that you formatted above.
- Delete any nodes from the network that have no connections to other genes. To do so, go back to the Excel workbook and network sheet and delete both the row and column with the floating gene's name. Then re-upload the edited file to GRNsight to visualize it.
Results
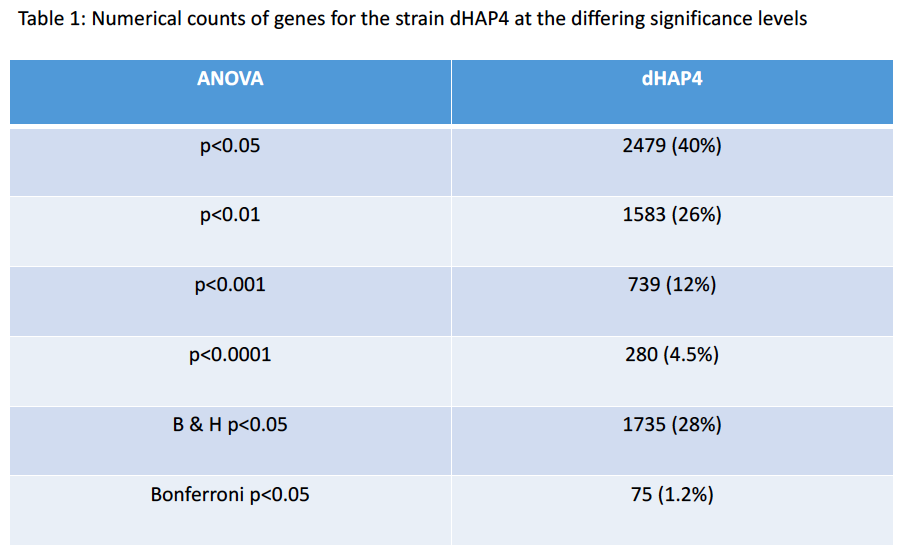
- How many genes have p < 0.05? and what is the percentage (out of 6189)?
- 2479 genes, 40%
- How many genes have p < 0.01? and what is the percentage (out of 6189)?
- 1583 genes, 26%
- How many genes have p < 0.001? and what is the percentage (out of 6189)?
- 739 genes, 12%
- How many genes have p < 0.0001? and what is the percentage (out of 6189)?
- 280 genes, 4.5%
- How many genes are p < 0.05 for the Bonferroni-corrected p value? and what is the percentage (out of 6189)?
- 75 genes, 1.2%
- How many genes are p < 0.05 for the Benjamini and Hochberg-corrected p value? and what is the percentage (out of 6189)?
- 1735 genes, 28%
P-values Table:
Files
The following files were generated through this assignment and are accessible through the LMU Box folder for this course (BIOL 388). Week 4: Raw Data Filename: BIOL388_S19_microarray-data_dHap4.zip Week 4 Workbook Filename: BIOL388_S19_microarray-data_dHAP4AL 2019.xlsx
Week 5:
- Powerpoint Presentation
- Gene List and GoLists
- Gene Expression Profiles
- Box: https://lmu.app.box.com/folder/67674514569
Comparing results with known data:
- The expression of the gene NSR1 (ID: YGR159C)is known to be induced by cold shock. Find NSR1 in your dataset. What is its unadjusted, Bonferroni-corrected, and B-H-corrected p values? What is its average Log fold change at each of the timepoints in the experiment?
- P-value: 0.0164
- B-H:0.055
- Bonferoni: 101.3
- Average Log Fold:
- t15: 2.7
- t30: 3.25
- t60: 3.5
- t90: -1.1
- t120: -1.8
Analyzing STEM Results
Why did you select this profile? In other words, why was it interesting to you?
- I selected profile 48 because
How many genes belong to this profile?
- 256 genes were assigned under this profile,
How many genes were expected to belong to this profile?
- 32.6 genes were expected to belong under this profile.
What is the p-value for the enrichment of genes in this profile?
- The p-value was 1.8*10^(-141)
How many GO terms are associated with this profile at p < 0.05?
- 167/578 genes fit the significance profile.
How many GO terms are associated with this profile with a corrected p value < 0.05?
- 6 GO terms are associated with this profile of corrected p<0.05.
Gene Ontology Terms
Six Gene Ontology terms from both the significant filtered corrected p-value (1 term) and non-corrected list (5 terms) were defined below.
SWI/SNF superfamily-type complex
- A protein complex that contains an ortholog of the Saccharomyces ATPase Swi2/Snf2 as one of the catalytic subunit components (ATPase) and mediates assembly of nucleosomes, changes to the spacing or structure of nucleosomes, or some combination of those activities in a manner that requires ATP.
Chromatin Organization
- Any process that results in the specification, formation or maintenance of the physical structure of eukaryotic chromatin.
Organelle lumen
- The internal volume enclosed by the membranes of a particular organelle; includes the volume enclosed by a single organelle membrane, e.g. endoplasmic reticulum lumen, or the volume enclosed by the innermost of the two lipid bilayers of an organelle envelope.
- Term appeared twice with slight variation.
INO80-type complex
- A chromatin remodeling protein complex initially purified from S. cerevisiae and containing more than 10 subunits, including the SWR1-related complexes. INO80 (inositol requiring 80)-type complexes have diverse functions, including promoting transcriptional activation and DNA repair.
snoRNA metabolic process
- The chemical reactions and pathways involving snoRNA, small nucleolar RNA, any of a class of small RNAs that are associated with the eukaryotic nucleus as components of small nucleolar ribonucleoproteins. They participate in the processing or modifications of many RNAs, mostly ribosomal RNAs (rRNAs) though snoRNAs are also known to target other classes of RNA, including spliceosomal RNAs, tRNAs, and mRNAs via a stretch of sequence that is complementary to a sequence in the targeted RNA.
Maturation of 5.8S rRNA
- Any process involved in the maturation of a precursor 5.8S ribosomal RNA (rRNA) molecule into a mature 5.8S rRNA molecule.
- Term appeared twice in slight variations.
All definitions were found at the following: http://geneontology.org
Yeastract
In the results window that appears, the p values colored green are considered "significant", the ones colored yellow are considered "borderline significant" and the ones colored pink are considered "not significant". How many transcription factors are green or "significant"?
- Five transcription factors are green colored and thus significant. One transcription factor is also yellow colored or borderline significant.
Are GLN3, HAP4, and/or ZAP1 on the list? If so, what is their "% in user set", "% in YEASTRACT", and "p value".
- Yes, HAP4, ZAP1, and GLN3 are all on the list but do not show significance.
- HAP4: 14.90% user set, 3.81% in YEASTRACT, and 0.296787375987255 p-value.
- GLN3:18.04% user set, 3.87% YEASTRACT, and 0.242276356702778 p-value.
- ZAP1:23.53% user set, 3.90% YEASTRACT, and 0.196629632608810 p-value.
You need to select from this list of "significant" transcription factors, which ones you will use to run the model. You will use these transcription factors and add GLN3, HAP4, and ZAP1 if they are not in your list. Explain in your electronic notebook how you decided on which transcription factors to include. Record the list and your justification in your electronic lab notebook. Each group member will select a different network (they can have some overlapping transcription factors, but some should also be different).
- I used the following transcription factors for the model:
- Sfp1p (significant)
- Ace2p (significant)
- Rim101p (significant)
- Arg81p (significant)
- Ash1p (significant)
- Yox1p (borderline significant)
- Met31p
- YLR278C
- Mth1p
- Gcn4p
- Stb5p
- Ume6p
- Asg1p
- Msa1p
- Ndt80p
- Hap4p
- Zap1p
- Gln3p
- I used these transcription factors because they were the ones that were either significant/boreder-line significant and the rest were chosen based upon the order they appeared. Zap1p, Gln3p, and Hap4p were also included based on the interest of this project.
Conclusion
The P-value cut-off being p<0.05 often amounts to numbers seeming significant by chance. This means that 5% of our data would be seeming significant by chance or 309 times. Since we have more than 309 genes that pass the cutoff, some of our values are significant not simply by chance. This showed that the p-value is a moveable confidence level. Through applying the additional tests- Bonferroni, as well as Benjamini and Hochberg corrections to the unadjusted p-values we applied more strict criteria to the p-values than simply performing anova. In week 5, through various software such as STEM, GRNsight, and YEASTRACT, various tests were conducted to identify the relationships between transcription factors in the dHAP4 strain of yeast. The STEM program produced seven significant clusters, of which profiles 16, 40 45, and 48 were induced during cold shock and then repressed during recovery, while profiles 22, 2, and 9 showed an opposite trend. I chose profile 48 to further analyze due to its upward trend during cold-shock. Profile 48 was then further analyzed through YEASTRACT software to determine the transcription factors responsible for regulation of these genes. Five were determined to be significant and one border-line significant. These genes were then made into a matrix and uploaded to GRNsight where a network was formulated of all the interactions.
Acknowledgements
I would like to acknowledge my homework partners who also worked with the dHAP4 strain, Desiree, and Brianna. Desiree, Brianna and I texted a few times and met in person. Additionally, Dr. Dahlquist helped me multiple times via email and in office hours with errors I was making on excel. For the week 5 assignment Dr. Dahlquist also helped me to understand all of the software and some minor issues I was having with what information needed to be input in which of the various locations.
Except for what is noted above, this individual journal entry was completed by me and not copied from another source. Avalekander (talk) 19:43, 13 February 2019 (PST)
References
The Week 4 and Week 5 Assignments were referenced to create this journal entry.
The data was provided by Dr. Dahlquist's lab. She also provided the Excel file "BIOL388_S19_microarray-data_wt" with the raw data.
I would like to reference Gene Ontology where I defined the gene ontology terms I chose at http://geneontology.org
Assignment Pages
- Template: Ava Lekander
- User Page: Ava Lekander
- Journal Entries:
- Assignment Pages:
- Class Journal Pages:
- Biology 388 Home Page: BIOL 388 Class Page