EdwardRyanTalatala Week 14/15
Purpose
The purpose of this assignment was use MATLAB models to improve our understanding of the experiment and analysis of the Tai et al. (2007) paper. We also investigated the glucose efficiency/waste constant for glucose-limited and ammonium-limited conditions.
Methods
Use the MATLAB files from Dr. Fitzpatrick (zipped package).
Determining temperature dependence using Arrhenius equation
Arrhenius equation: rate = A*e^(-B/(R*T))
- Figure out the constants A and B from the rate data in the TaiParamsRevised.m file.
- Simulate the chemostat for T = 15,20, 25 °C conditions and graph the time courses of the biomass and nutrients.
Investigating glucose efficiency/waste constant
Investigate the glucose efficiency/waste constant (that is not really a constant?) for the glucose-limited and ammonium-limited conditions.
- Note the values of E for glucose-limited and ammonium-limited conditions.
- For each temperature (12, 30), find a function E(y) that matches the two points of (y,E) data.
- Modify the chemostat_2nutrient_dynamics.m file to use the functions you've created.
- Compare the resulting simulation to the previous one.
Results
Determining A & B Constants
Variables calculated using Arrhenius equation:
B = 69,840.59
- found using B = (Rln(k1/k2))/(1/T1-1/T2)
A = 4.979 * 10^11
- found by plugging in all the variables in Arrhenius equation
r(15°C) = (4.979 x 10^11)e^(-69840.59)/((8.314)(288.15)) = 0.1087
r(20°C) = (4.979 x 10^11)e^(-69840.59)/((8.314)(293.15)) = 0.1787
r(25°C) = (4.979 x 10^11)e^(-69840.59)/((8.314)(298.15)) = 0.289
MATLAB files for temperature dependence
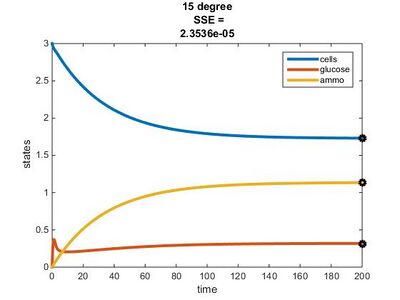
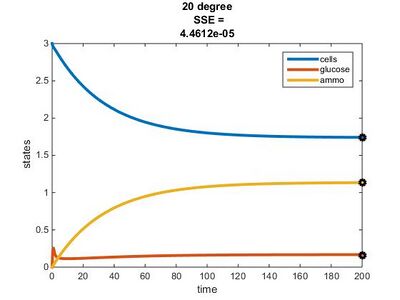
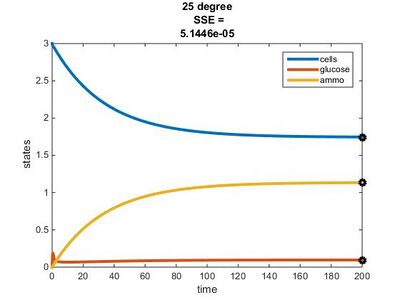
Temperature Dependence Graphs
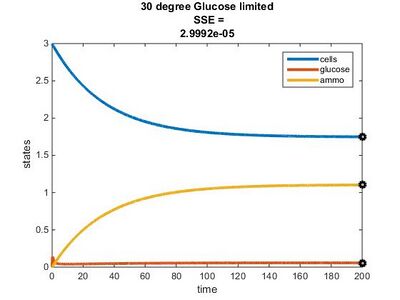
We used the glucose-limited conditions for the 15, 20, & 25°C graphs. We changed the values for the residual concentrations in the Params file until the SSE value was close to 0.
Efficiency Constant Investigation
Original: E = 1/Y
New equation: where E = my + b
- y = residual glucose
- use point intercept m = (y-y)/(x-x) to determine m
- plug new value into E = my + b to determine b
12°C: E = 0.363y + 14.11
- m = (14.3-20)/(0.5045-16.22) = 0.363
- b=14.11
30°C: E = 0.7y + 14.25
- m = (14.3-25)/(.0541-15.33) = 0.7
- b=14.25
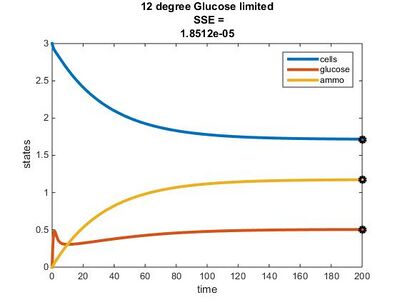
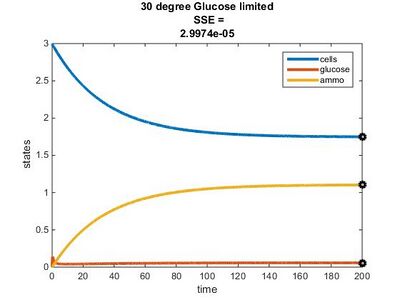
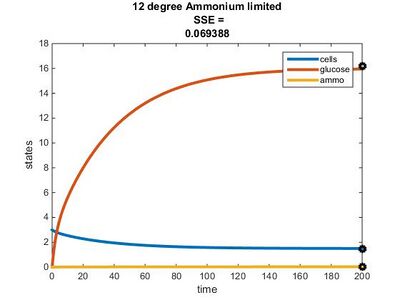
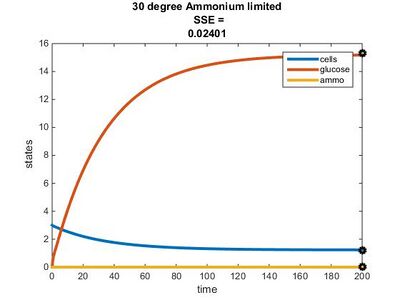
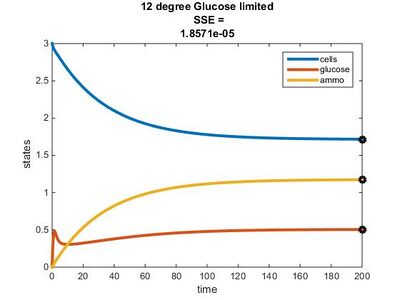
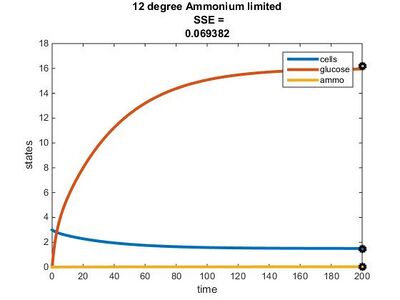
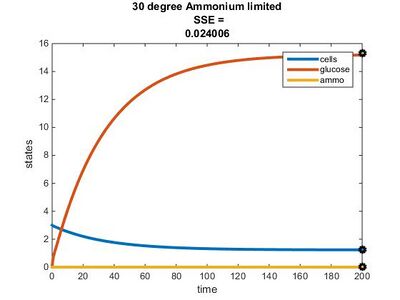
Efficiency Constant Graphs
Original Model
New Model
MATLAB files for efficiency investigation
Acknowledgments
I would like to acknowledge my homework partner, Fatimah with whom I worked with to complete the assignment.
I worked with Austin to work on the efficiency constant investigation.
Except for what is noted above, this individual journal entry was completed by me and not copied from another source. EdwardRyanTalatala (talk) 00:44, 9 May 2019 (PDT)
References
Dahlquist, K. and Fitzpatrick, B. (2019). BIOL388/S19:Week 14/15. [online] openwetware.org. Available at:Week 14/15 Assignment Page [Accessed May 8 2019].
Assignments
- Assignment-Week 1
- Assignment-Week 2
- Assignment-Week 3
- Assignment-Week 4
- Assignment-Week 5
- Assignment-Week 6
- Assignment-Week 7
- Assignment-Week 9
- Assignment-Week 10
- Assignment-Week 11
- Assignment-Week 12
- Assignment-Week 14/15
Journal Entries
- Journal-Week 2
- Journal-Week 3
- Journal-Week 4/5
- Journal-Week 6
- Journal-Week 7
- Journal-Week 9
- Journal-Week 10
- Journal-Week 11
- Journal-Week 12
- Journal-Week 14/15
Class Journal Entries