Isai Lopez Individual Journal 9
From OpenWetWare
Jump to navigationJump to search
Purpose
- The purpose of this assignment was to familiarize ourselves with the tools we will be using to analyze sequence and protein structure of the gp120 protein of HIV.
Methods/ Results
- Use the Biology Workbench website to view one of the Markham HIV sequences and post that sequence into one of the two DNA to protein converting websites given. We used ExPASY Translate tool but NCBI Open Reading Frame Finder is also appropriate.
- After pasting the sequence and translating the sequence, you will get a result like the one that follows.
- Visit the UniProt Knowledgebase (UniProt KB) site to guide research on the different proteins you will be working with.
- Typing "HIV" and "gp120" yields 206,278 results
- The listing for "P04578" provides a plethora of information about the protein ranging from processes and pathways the protein participates in, taxonomic listings, location within a cell, topology, structure, and important domains of the protein.
- Using the PredictProtein server, copy and paste once of the Markham sequences and enter the information to predict the protein based off the sequence.
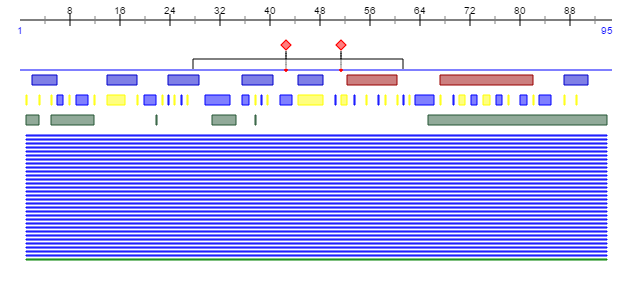
- The figure above shows the result for the protein prediction of Subject 9 Visit 1-5 from the Markham study
- The output information from the website shows a sort of road map for the protein derived from the amino acid sequence coded for by the sequence from the Markham study. Notably, the information from this program displays the locations of important secondary structures of the greater protein, such as helices, loops, and sheets. The same can be found in the UniProt database entry for this envelope protein, in addition to background information about the functions of the protein in interaction with other molecules.
- Use the following link to reach a file containing structural information from the paper read in class for the journal club. Kwong et al. (1998) structure 1GC1
- This is as far as I was able to get in class, and there is one unfinished section from the in-class activity
Data and Files
http://openwetware.org/wiki/Image:IsaiLopezTranslateToolResults.png http://openwetware.org/wiki/Image:IsaiLopezPredictProtein.png The above files were generated for use in the methods and results section.
Scientific Conclusion
- Throughout today's work, we worked to find out the relevant information on the gp120 protein using the UniProt Website. Using this as a jumping off point, we were able to use tools that would help us turn sequence information from the Markham study into amino acid sequences using http://web.expasy.org/translate/. We also learned how to identify which of the six reading frames from the output is the one to use in analysis. From here, we used PredictProtein server to turn the amino acid sequence into a visual guide for analyzing the locations of solvent accessibility, disorder, secondary structures, disulphide brides, as well as the degrees to which mutations in different residues cause a change in the type of amino acid. While we weren't able to use the last tool to analyze the 3D structure of the complex of the antibody/ CD4 portions/ gp120, this will be useful in visualizing how mutations in the DNA sequence change interactions within the greater complex.
Defining the Research Project
- What is your question?
- How does degree of amino acid change due to mutations in the gp120 sequence from the Markham et al. (1998) study relate to rate of HIV-1 progression?
- Make a prediction about the answer to your question before you begin your analysis.
- We predict that a greater degree of amino acid change will be associated with a greater level of HIV-1 progression.
- Which subjects, visits, and clones will you use to answer your question?
- We will use two clonal sequences from each of the 15 subjects at the beginning of the study and compare them to the same sequence in two clones for each of the 15 subjects at their last visit for the study. We chose to do this because it will ideally give a better overall view of how amino acid changes affect the seroconversion rate, while accounting for subjects in all three progression groups. This will give us a total of 60 sequences to work with, and while that is more than the ~50 recommended, we have successfully used the alignment tool with around this many sequences in the past with success.
Acknowledgments
- I worked with Colin Wikholm both in class formulating our research question and out side of class in finalizing the responses.
- I copied wiki syntax referencing the websites used in the in-class activity from the Week 9 Instruction Page posted and written by Dr. Dahlquist
- While I worked with the people noted above, this individual journal entry was completed by me and not copied from another source.
- Isai Lopez 22:54, 31 October 2016 (EDT):
