Isai Lopez Individual Journal 6
Purpose
The purpose of our study was to examine extremes in baseline nucleotide sequence differences across multiple subjects from the Markham study and analyze the correlation to slope of change in nucleotides between visits per clone per year. We hypothesize that clones from subjects with high baseline nucleotide differences will display lower slopes for change in intravisit nucleotide differences per year, while clones from subjects with low nucleotide difference will display higher slopes of change in nucleotide difference per year.
Methods/ Results
- We took the nucleotide sequences from the clones of the first visits for subjects 1,15,9,4,13, and 14 from Visit 1 S 1-9 and Visit 1 S 10-15
- Subjects were chosen by their initial intravisit nucleotide sequence differences, taking the top three and bottom three for this category.
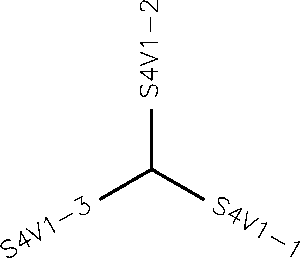
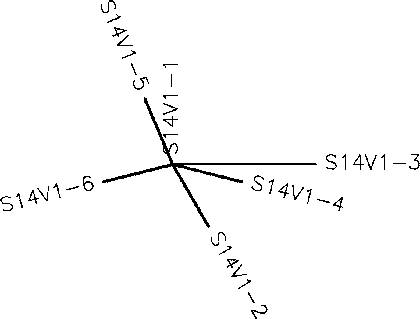
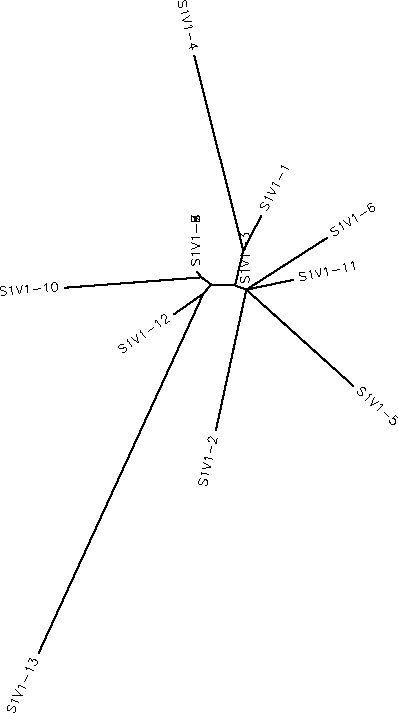
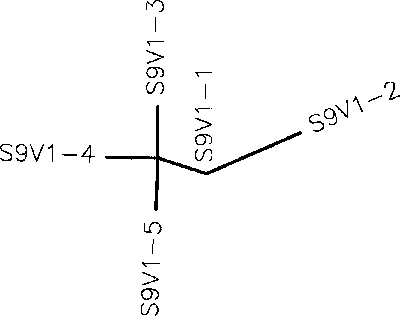
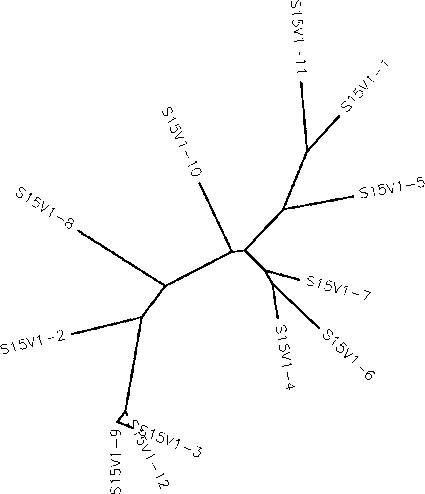
- Generate an unrooted tree for the sequences of each of the six subjects using the ClustalW tool on the Biology Workbench
- Use the ClustalDist tool to measure base pair difference across all the clones for each subject to generate minimum and maximum differences. To do this, multiply the smallest non-zero and largest values of the distance matrix by the number of base pairs in the sequence.
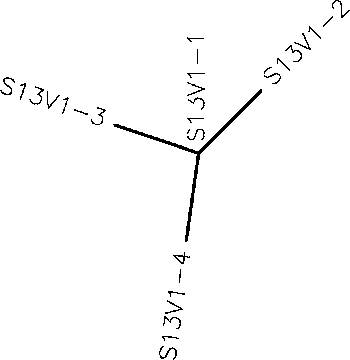
- Calculate the range of distances by subtracting the min and max distances for each subject.
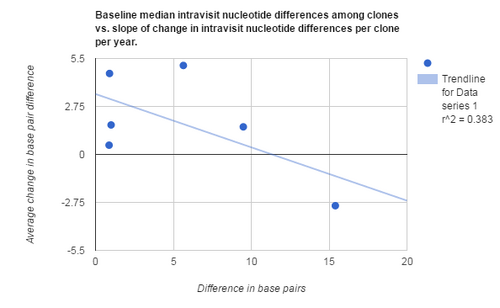
- Generate a scatterplot for the data comparing baseline median intravisit nucleotide differences among clones vs slope of change in intravisit nucleotide differences per clone per year
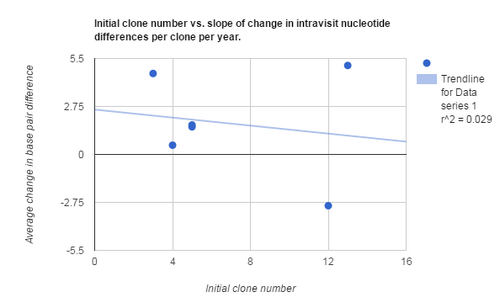
- Initial clone number vs slope of change in intravisit nucleotide differences per clone per year
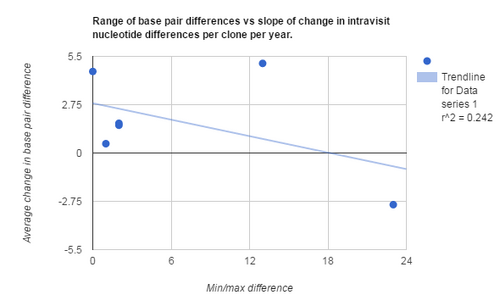
- Base pair difference range vs slope of change in intravisit nucleotide differences per clone per year
- All three graphs had p values that were greater than .05 when analyzed using a T-test
Data and Files
- Subject 1 unrooted
- Subject 4 Rooted
- Subject 9 unrooted
- Subject 13 Rooted
- Subject 14 Rooted
- Subject 15 unrooted
- Graph 1
- Graph 2
- Graph 3
Conclusion
After analyzing the data of all 6 subjects using correlation analysis and minimum/ maximum distances between sequences, we did not find any data to support our initial hypothesis. This suggests that, at least using the data from the Markham et al. paper, initial differences between baseline nucleotide sequences are not enough to explain the final values for the slopes of change in intravisit nucleotide differences per clone per year. This lack of correlation is also seen when comparing initial clone number and range in nucleotide differences compared to final slope. Later studies suggest that the results may be explained by a lack of information on the subjects, such as outside factors that could have contributed to the differences in immune response and the subsequent change in diversity seen across HIV strains.
Acknowledgements
- I worked with Matthew R Allegretti both electronically through text and in lab in collaborating on the powerpoint. I also used some of the same syntax in posting the pictures of our graphs and unrooted trees for our subjects, as well as reference information used on our powerpoint.
- While I worked with the people noted above, this individual journal entry was completed by me and not copied from another source.
- Isai Lopez 02:13, 11 October 2016 (EDT):
References
- Markham, R. B., Wang, W. C., Weisstein, A. E., Wang, Z., Munoz, A. Templeton, A., Margolick, J., Vlahov, D., Quinn, T., Farzadegan, H., Yu, X. F. (1998) Patterns of HIV-1 Evolution in individuals with differing rates of CD4 T cell decline. Proceedings of the National Academy of Sciences USA, 95(21), 12568-12573. DOI: 10.1073/pnas.95.21.12568
- Tully, D. C., Ogilvie, C. B., Batorsky, R. E., Bean, D. J., Power, K. A., Ghebremichael, M., et al. (2016) Differences in the Selection Bottleneck between Modes of Sexual Transmission Influence the Genetic Composition of the HIV-1 Founder Virus. PLoS Pathog 12(5): e1005619. doi:10.1371/journal.ppat.1005619
- Mehandru, S., Deren, S., Kang, S. Y., Banfield, A., Garg, A., Garmon, D., et al. (2015) Behavioural, Mucosal and Systemic Immune Parameters in HIV-infected and Uninfected Injection Drug Users. Journal of addiction research & therapy, 6(4), 1. doi:10.4172/2155-6105.1000257