AVM BIOL368 Week 10
Purpose
The purpose of this lab is to continue working with our partner on our HIV Structure research assignment and create a powerpoint to present to the class next week.
Methods and Results
- Continue where we left off last week on the HIV structure project
- Use NCBI Structure Database or Star Biochem program to answer the following questions:
- Find the N-terminus and C-terminus of each polypeptide tertiary structure.
- Locate all the secondary structure elements. Do these match the predictions made by the PredictProtein server?
- Yes, these do match the predictions
- Locate the V3 region and figure out the location of the Markham et al. (1998) sequences in the structure.
- The V3 region was found using the sequences that we have from the markham data. I located the region using the starting amino acids "env".
- Perform a multiple sequence alignment on the protein sequences that you and your partner are analyzing for your project using the Biology Workbench.
- Are there more or fewer differences between the sequences when you look at the DNA sequences versus the protein sequences?
- There are more differences between the DNA sequence than there are in the protein sequence.
- How do you account for this?
- There are major and minor changes in the amino acid sequence. Some of the changes observed may not make much difference.
- Are there more or fewer differences between the sequences when you look at the DNA sequences versus the protein sequences?
- Once you have oriented yourself, analyze whether the amino acid changes that you see in the multiple sequence alignment would affect the 3D structure and explain why you think this.
- The changes in the amino acid structure may or may not affect the 3D structure. Some of the major changes, say between an amino acid with a long chain to an amino acid with a benzene ring, would have a more dramatic effect on the structure.
HIV Structure Project:
- What is your question?
- Does diversity and divergence increase within the amino acid sequences over time more within the rapid progressors compared to non-progressors?
- Make a prediction about the answer to your question before you begin your analysis.
- Since divergence and diversity increases over time within the DNA sequences for all progressor types, and more so within the rapid progress ors, we would expect to see more diversity within the amino acid sequences over time within the rapid progressors.
- Which subjects, visits, and clones will you use to answer your question?
- For rapid progressors we are looking at subject 3 and 10's first and last visits and for non-progressors we are looking at subject 12 and 13's first and last visits. These were chosen based on similar number of visits.
- To start our research project we created multiple sequence alignments for subjects 3, 10, 12 and 13
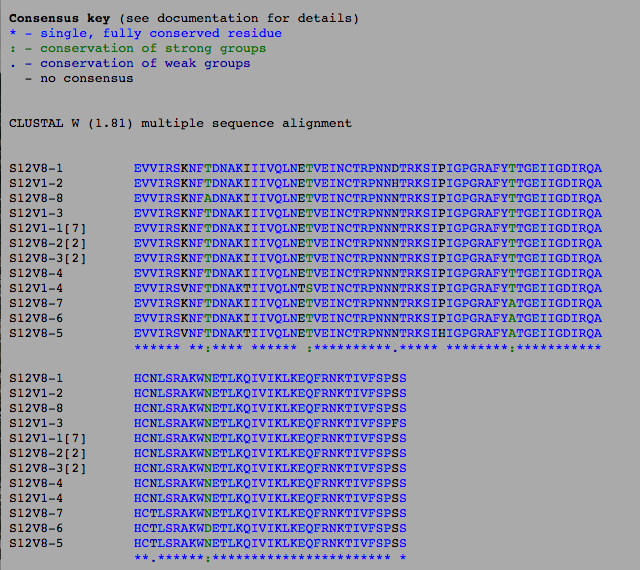
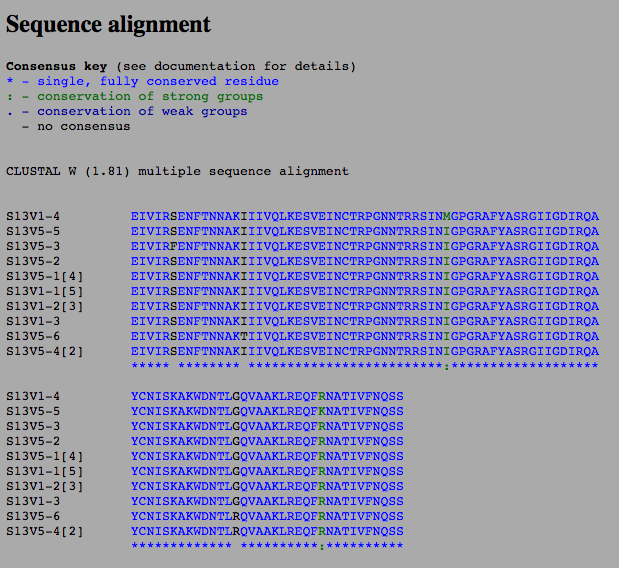
- Photos below show the sequence analysis of subjects 12 and 13
- The sequences we analyzed within the box shade application
- Boxshade photos shown below for subjects 12 and 13
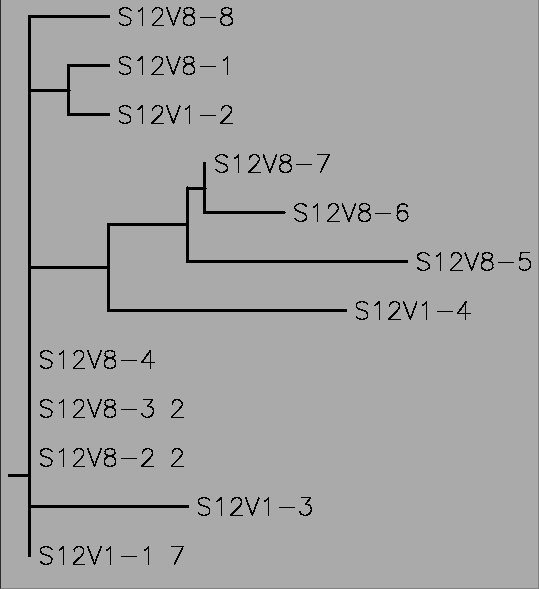
- A rooted tree was created for each subject using clustalW
- Rooted tree shown below for subject 12
- A rooted tree showing all the subjects with their first and last visits may help us determine if the categorization as non-progressors and rapid progressors is the best was to classify the subjects
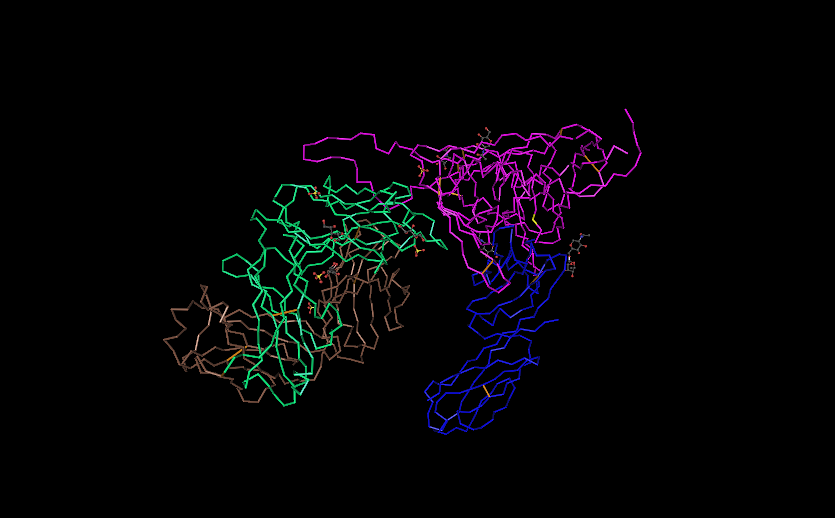
- The number of differences among the amino acids were collected for each subject and entered into the protein structure so that the differences could be visualized and see where they occurred within the structure, below are a few of the differences observed.
- Subject 13 last visit, mutation 5 shows a mutation within the V3 loop highlighted in yellow.
- Finally we performed a Clustaldist to find the distance between each subjects first and last visits to compare the amount of diversity each subject had. We can compare the rapid progressors to the nonprogressors and see if one group was more or less diverse than the other.
- Subject 12: 0.010
- Subject 13: 0.021
- Subject 10: 0.137
- Subject 3: 0.010 (Only four clones)
Data and Files
Conclusion
In this research project we chose two subjects from Markham's rapid progressors (3 and 10) and two subjects from the non-progressors (12 and 13) and analyzed whether more diversity would be observed within one groups amino acid sequences. To do this we took the first and last visits of each subject and analyzed the number of mutations within the amino acid sequences using alignment and protein tools in biology workbench. The non-progressors had a slightly lower number of mutations than the rapid progressors. Subject 12 had 11 mutations, 13 had 5 mutations. Subject 3 had 9 mutations and subject 10 had 24 mutations. Although these numbers suggest higher diversity among rapid progressors, we did not run any statistical tests to prove this, since there would most likely be no significant difference. We compared the distances between the number of mutations in each subjects last and first visit using Clustldist. These numbers slightly proved that the rapid progressors had a higher number of mutations than the non-progerssors, but was hard to confirm since Subject 3 only had four clones in the last visit and a lower difference in number of mutations. A number of the mutations were analyzed visually (using Huang's structure) by highlighting the sequence on gp120 and observing where they occurred within the V3 loop. Most of the mutations don't seem to be in a specific spot that could affect the binding of gp120. Lastly, rooted phylogenetic trees were constructed for each of the subjects, but don't but don't serve our research project specifically. They did not show any outstanding patterns.
Acknowledgments
- Worked in collaboration with research partner Mia Huddleston in class.
- We collaborated on our presentation both in class and at home throughout the week.
- Also spoke with Shivum in class, as his project was similar to ours.
- Template followed from weeks 9 and 10 found at BIOL368/F16, BIOL368/F16
- Help from our professor, Dr. Dahlquist in class.
- Note: While I worked with the people noted above, this individual journal entry was completed by me and not copied from another source.
Avery Vernon-Moore 14:07, 7 November 2016 (EST)
References
- Huang, C. C., Tang, M., Zhang, M. Y., Majeed, S., Montabana, E., Stanfield, R. L., ... & Wyatt, R. (2005). Structure of a V3-containing HIV-1 gp120 core. Science, 310(5750), 1025-1028.
- Kirchherr, J. L., Hamilton, J., Lu, X., Gnanakaran, S., Muldoon, M., Daniels, M., Kasongo, W., Chalwe, V., Mulenga, C., Mwananyanda, L., Musonda, R.M., Yuan, X., Montefiori, D.C., Korber, M.T., Haynes, B.F., & Musonda, R. M. (2011). Identification of amino acid substitutions associated with neutralization phenotype in the human immunodeficiency virus type-1 subtype C gp120. Virology, 409(2), 163-174. DOI: 10.1016/j.virol.2010.09.031
- Markham, R.B., Wang, W.C., Weinstein, A.E., Wang, Z., Munoz, A., Templeton, A., Margolick, J., Vlahov, D., Quinn, T., Farzadegan, H., & Yu, X.F. (1998). Patterns of HIV-1 evolution in individuals with differing rates of CD4 T cell decline. Proc Natl Acad Sci U S A. 95, 12568-12573. dos: 10.1073/pnas.95.21.12568
Weekly Assignments:
Assignment 1 Assignment 2 Assignment 3 Assignment 4 Assignment 5 Assignment 6 Assignment 7 Assignment 8 Assignment 9 Assignment 10 Assignment 11 Assignment 12 Assignment 14 Assignment 15
Individual Journals:
"Week 1: Create Wiki Page" Individual Journal 2 Individual Journal 3 Individual Journal 4 Individual Journal 5 Individual Journal 6 Individual Journal 7 Individual Journal 8 Individual Journal 9 Individual Journal 10 Individual Journal 11 Individual Journal 12 No Week 13 Assignment Individual Journal 14 Individual Journal 15
Shared Journals:
Class Journal 1 Class Journal 2 Class Journal 3 Class Journal 4 Class Journal 5 Class Journal 6 Class Journal 7 Class Journal 8 Class Journal 9 Class Journal 10 Class Journal 11 Class Journal 12 Class Journal 14 Class Journal 15