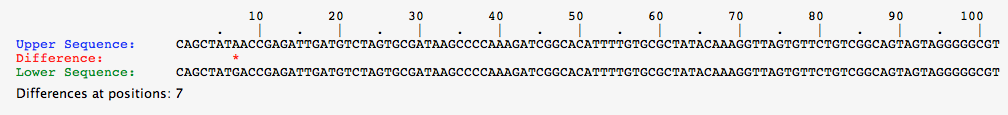Matthew K. Oki Individual Journal 2
From OpenWetWare
Jump to navigationJump to search
Matthew K. Oki I.J. Week 2
Purpose
- The purpose in this investigation was to find the amino acid sequence that yielded a pure-breeding purple plant. This amino acid sequence was reverse engineered from proteins.
Methods & Results
- The formal instructions for the methods of this investigation were taken from the following link: Part 3: Molecular Biology
- Open Aipotu and go to the "Molecular Biology" Tab
- Task 1: Are all the white alleles the same?
- All white alleles are not the same
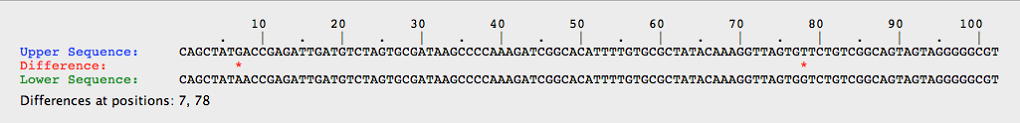
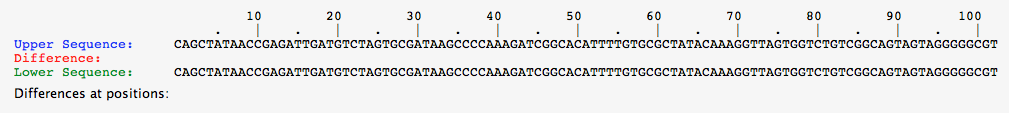
- This image below shows the comparison of two white alleles, and clearly they have two differences.
- The top white allele was found by taking the white allele from a red organism, while the bottom was from a white organism.
- All white alleles are not the same
- Task 2: DNA Sequences of the Four Starting Organisms
- Double click your desired organism in the "Greenhouse"
- This will postulate the two different genes of the organism in the upper and lower windows
- Clicking on the compare tab "Upper vs. Lower" will compare each individual nucleotide of the two genes
- Compare all "Greenhouse" organisms
- Double click your desired organism in the "Greenhouse"
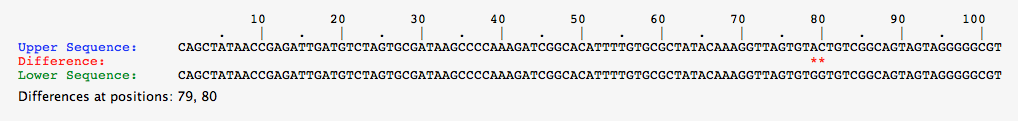
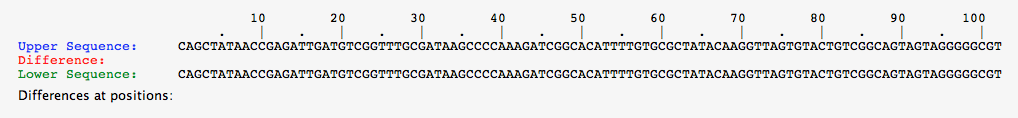
- Comparison of Green-1 Organism Above
- Comparison of Green-2 Organism Above
- Comparison of Red Organism Above
- Comparison of White Organism Above
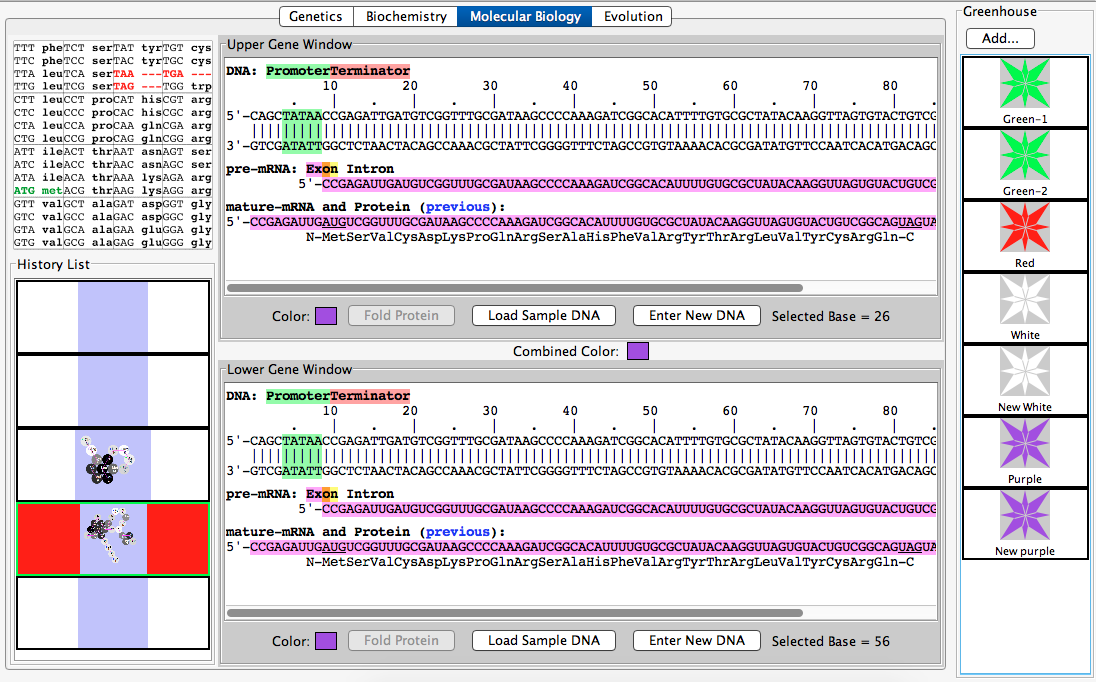
- Task 3: Pure-Breeding Purple Organism
- Enter the protein: MLVKEIAMYRFATHER
- In the "Molecular Biology" Tab, enter the corresponding nucleotides of the protein shown above into the "Enter New DNA" Tab.
- Enter the protein: MLVKEIAMYRFATHER
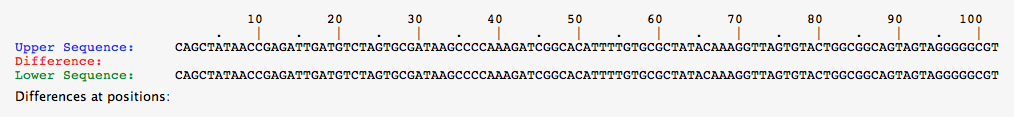
- The two sequences were compared to confirm a pure-breeding purple organism.
Scientific Conclusion
- We set out with goal to create a DNA sequence for a pure-breeding purple organisms.
- We discovered the pure-breeding purple organism DNA sequence to be the following: N-MetLeuValLysGluIleAlaMetTyrArgPheAlaThrHisGluArg-C
- Our purpose was fulfilled because we were able to get generations of pure-breeding purple plants with the above DNA sequence.
Acknowledgments
- I would like to thank Avery Vernon-Moore and Isai Lopez for assistance on this project.
- I would also like to thank Kam D. Dahlquist, Ph.D. for providing the instructions and information for this assignment both in class and on this document: BIOL368/F16:Week 2.
- The syntax for the "A "Rule of Five" Framework for Models and Modeling to Unify Mathematicians and Biologists and Improve Student Learning" article originally written by Kam D. Dahlquist, Ph.D. was used in the Reference section.
- Even though I worked with the people noted above, this second individual journal entry was completed by me and not copied from another source.
- Matthew K. Oki 01:54, 13 September 2016 (EDT):
References
- The following websites were used as references for this individual journal assignment:
- The following article was used as a reference for the shared journal assignment:
Class Homework Links
Weekly Assignments
Individual Journals
Class Journals
Other Links
My Home Page: Matthew K. Oki
Class Home Page: Bioinformatics Home Page
My Paper Resume: Matthew K. Oki Resume