Matthew K. Oki Individual Journal 10
From OpenWetWare
Jump to navigationJump to search
Week 10 Individual Journal
Purpose
- Our purpose was to continue using the programs from BIOL368/F16:Week 9 to further explore our HIV project question and hypothesis.
Methods & Results (Assignment)
- Protein Structure from Huang et al. (2005) was downloaded and opened in Cn3D software site.
- Locate the V3 region and figure out the location of the Markham et al. (1998) sequences in the structure.
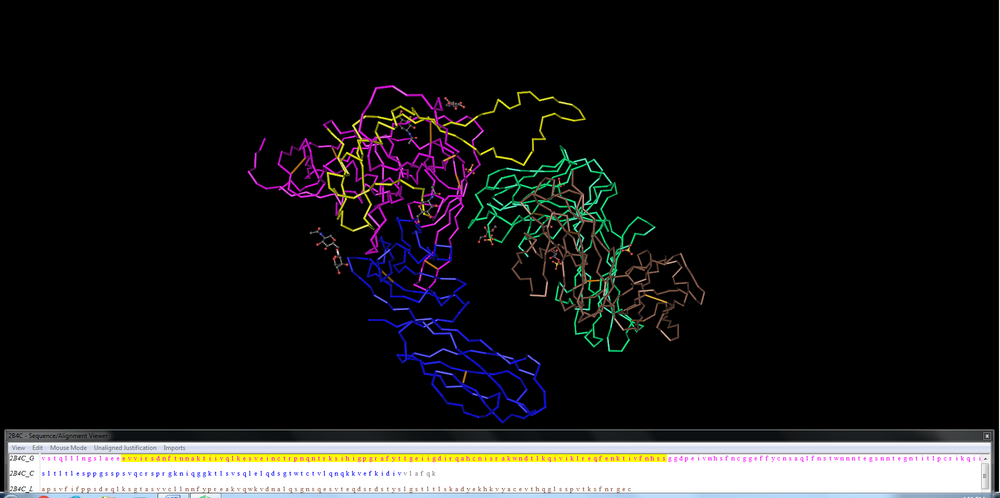
- In order to determine the V3 region, we found the general equivalent to the Markham et al. sequences in the Huang et al. structure. We then highlighted the sequence in the Cn3D software, which in turn highlighted the structure as shown below:
- Locate the V3 region and figure out the location of the Markham et al. (1998) sequences in the structure.
- Perform a multiple sequence alignment on the protein sequences that you and your partner are analyzing for your project using the Biology Workbench.
- Are there more or fewer differences between the sequences when you look at the DNA sequences versus the protein sequences? How do you account for this?
- There were significantly more difference in the DNA sequences, but this can be associated with the change from nucleotides to amino acids. This decreases the sample size from 285 to 95. The theta value accounts for this change by the difference in sample size.
- Are there more or fewer differences between the sequences when you look at the DNA sequences versus the protein sequences? How do you account for this?
Methods & Results (Project)
- Use Biology Workbench to upload Subject 10's clones and run a ClustalW multiple sequence alignment.
- This step was ran separately for amino acid sequences and nucleotide sequences.
- S and Theta values were calculated from the alignment.
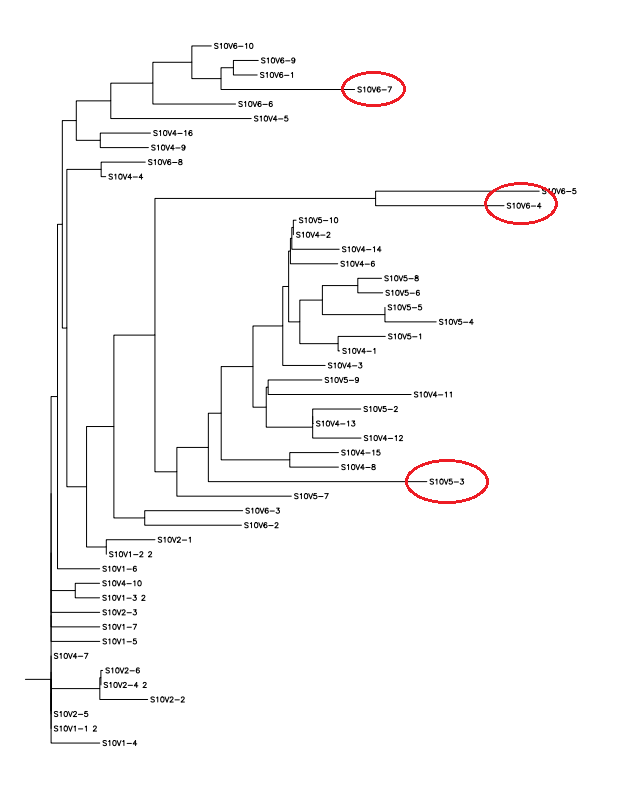
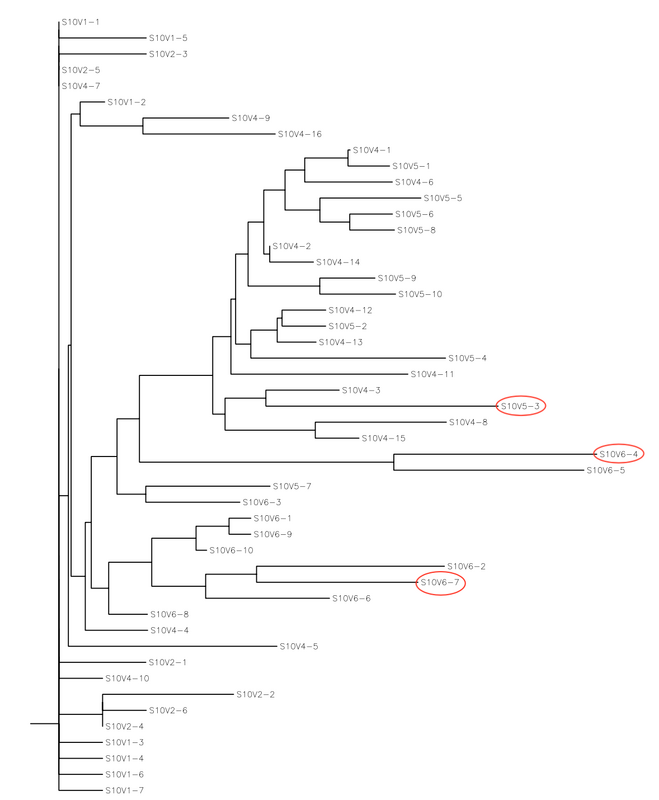
- This also gives an output of rooted trees for the amino acid and nucleotide sequence as shown below:
- This step was ran separately for amino acid sequences and nucleotide sequences.
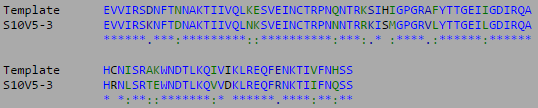
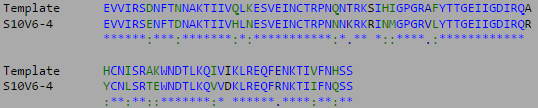
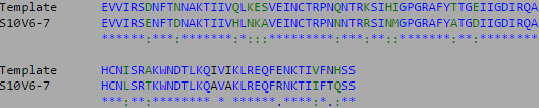
- The multiple sequence alignment for the three highest diversity clones is shown below:
- Using Biology Workbench again, the ClustalDist function was ran for amino acid and nucleotide sequences.
- This gave an output for min and max.
- The most divergent visit and clones were isolated and compared to the V3 target sequence provided by the Huang et al. (2005) data set.
- (to be continued in Week 11)
Data and Files
A pdf version of our work-in-progress can be found here.
Scientific Conclusion
- Using the data obtained from our methods, S10V6-4, S10V5-3, and S10V6-7 had the highest diversity compared to the original strain. Given that these clones are from the last visit, it numerically shows how much the amino acid sequences changed from the first visit to the last visit. In our next step, that has not been completed yet, we will look at which specific amino acid sequences were changed. This will give us an idea if there are certain amino acids that are more important in the physiological output of the change than others are.
Acknowledgements
- I would like to thank my partner, William P. Fuchs, for the assistance on this week's project, both in assistance on the structure program methods in class and completion of the our project idea outside of class.
- I would also like to thank Kam D. Dahlquist, Ph.D. for providing the instructions and information for this assignment both in class and on this document: BIOL368/F16:Week 10.
- Even though I worked with the people noted above, this individual journal entry was completed by me and not copied from another source.
- Matthew K. Oki 18:51, 7 November 2016 (EST):
References
- Biology Workbench
- BIOL368/F16:Week 10
- ExPASY Translate tool
- UniProt Knowledgebase (UniProt KB)
- PredictProtein server
- Cn3D software site
- Markham, R.B., Wang, W.C., Weisstein, A.E., Wang, Z., Munoz, A., Templeton, A., Margolick, J., Vlahov, D., Quinn, T., Farzadegan, H., & Yu, X.F. (1998). Patterns of HIV-1 evolution in individuals with differing rates of CD4 T cell decline. Proc Natl Acad Sci U S A. 95, 12568-12573. doi: 10.1073/pnas.95.21.12568
- Huang, C. C., Tang, M., Zhang, M. Y., Majeed, S., Montabana, E., Stanfield, R. L., ... & Wyatt, R. (2005). Structure of a V3-containing HIV-1 gp120 core. Science, 310(5750), 1025-1028. doi: 10.1126/science.1118398
Class Homework Links
Weekly Assignments
Individual Journals
Class Journals
Other Links
My Home Page: Matthew K. Oki
Class Home Page: Bioinformatics Home Page
My Paper Resume: Matthew K. Oki Resume