Macie Duran Week 11
Template
User Page
Assignments
- Week 1 Assignment
- Week 2 Assignment
- Week 3 Assignment
- Week 4 Assignment
- Week 5 Assignment
- Week 6 Assignment
- Week 7 Assignment
- Week 8 Assignment
- Week 9 Assignment
- Week 10 Assignment
- Week 11 Assignment
- Week 12 Assignment
- Week 14 Assignment
Journals
- Macie Duran Week 1
- Macie Duran Week 2
- Macie Duran Week 3
- Macie Duran Week 4
- Macie Duran Week 5
- Macie Duran Week 6
- Macie Duran Week 7
- Therapeutic Target Database (TTD) Review
- Macie Duran Week 10
- Macie Duran Week 11
- The Mutants Week 12
- The Mutants Week 14
Class Journals
- Class Journal Week 1
- Class Journal Week 2
- Class Journal Week 3
- Class Journal Week 4
- Class Journal Week 5
- Class Journal Week 6
- Class Journal Week 7
- Class Journal Week 8
- Class Bibliography Week 10
- Class Journal Week 11
- Class Journal Week 14
Purpose
The purpose of this assignment is to analyze and evaluate sources based on their reliability and validity. It is important to be able to be able to distinguish between reliable and non-reliable sources when conducting research. This week we will also be gaining a better understanding on SARS-CoV-2 in order to develop a final research project.
Methods & Results
Searching the Scientific Literature Part 2: Evaluating Scientific Sources
- I evaluated my group's assigned article in three areas: availability, the journal, and the article metadata.
- Article title: The Impact of Mutations in SARS-CoV-2 Spike on Viral Infectivity and Antigenicity
- Article citation: Li, Q., Wu, J., Nie, J., Zhang, L., Hao, H., Liu, S., ... & Qin, H. (2020). The impact of mutations in SARS-CoV-2 spike on viral infectivity and antigenicity. Cell, 182(5), 1284-1294. doi: 10.1016/j.cell.2020.07.012
- Link to article abstract on PubMed
- Link to the full text of the article in PubMed Central
- Link to the full text of the article (HTML format) from the publisher website
- Link to the full PDF version of the article from the publisher website
- Who owns the rights to the article?
- The publishing company -- Elsevier Incorporated
- How is the article available to you?
- The article is not open access, but LMU subscribes to Cell Press Journal. The Cell Press website also states that all of their peer-reviewed articles that are relevant to the COVID-19 outbreak have been made immediately open access or freely available for the duration of the outbreak.
- Is the article available online-only or both in print and online?
- The article is available in print and online.
- Evaluating the source -- the journal
- Who is the publisher of the journal?
- Elsevier Incorporated
- Is the publisher for-profit or non-profit?
- The publisher is for-profit.
- Is the publisher a scientific society?
- No, it is not a scientific society.
- Does the publisher belong to the Open Access Publishers Association?
- No, Elsevier Inc. does not belong to the Open Access Publishers Association.
- What country is the journal published in?
- Cell Press Journal is published in the United States.
- How long has the journal been in operation?
- Volume 1 of the article was published in 1974.
- Are articles in this journal peer-reviewed?
- Yes, the articles are peer-reviewed. Their peer-review process is explained here.
- Link to the scientific advisory board/editorial board of the journal
- What is the journal impact factor?
- The journal impact factor is 38.637, as of 2019. The journal metrics are found here.
- Who is the publisher of the journal?
- Evaluating the source -- the article
- Is the article a review or primary research article?
- The article is a primary research article.
- On what date was the article submitted?
- June 8, 2020
- On what date was the article accepted?
- July 10, 2020
- Did the article undergo any revisions before acceptance?
- Yes, the revised article was received on July 7, 2020.
- When was the article published?
- July 17, 2020
- What is the approximate elapsed time between submission and publication?
- A little over a month elapsed between the first submission and publication.
- What are the institutions with which the authors are affiliated?
- Qianqian Li is associated with the Graduate School of Peking Union Medical College in Beijing, China. Li and several other authors are all associated with the Division of HIV/AIDS and Sex-Transmitted Virus Vaccines, Institute for Biological Product Control, National Institutes for Food and Drug Control (NIFDC) and WHO Collaborating Center for Standardization and Evaluation of Biologicals. Qi Zhang and Linqi Zhang are associated with the Center for Global Health and Infectious Diseases, Comprehensive AIDS Research Center, and Beijing Advanced Innovation Center for Structural Biology, School of Medicine, Tsinghua University, Beijing, China. Xuguang Li is associated with the Centre for Vaccine Evaluation, Biologics and Genetic Therapies Directorate, HPFB, and Health Canada.
- Have the authors published other articles on this subject?
- The authors have also published an article on the [establishment and validation of a pseudovirus neutralization assay for SARS-CoV-2 https://pubmed.ncbi.nlm.nih.gov/32207377/].
- Is there a conflict of interest for any of the authors?
- No, the authors declare no competing interests.
- Is this a good article to evaluate further? Why or why not?
- This does seem like a good article. The publisher and journal are legitimate, and the authors show no conflict of interest. The article has also been peer-reviewed and revised before being published. Overall, it seems to be up to standard.
- Is the article a review or primary research article?
Preparation for Journal Club 2
- Article Title: The Impact of Mutations in SARS-CoV-2 Spike on Viral Infectivity and Antigenicity
- Article Citation: Li, Q., Wu, J., Nie, J., Zhang, L., Hao, H., Liu, S., ... & Qin, H. (2020). The impact of mutations in SARS-CoV-2 spike on viral infectivity and antigenicity. Cell, 182(5), 1284-1294. doi: 10.1016/j.cell.2020.07.012
10 biological terms and definitions
- Glycosylation: a biochemical process where a glycan attaches to a protein, a lipid, or other organic molecule, especially through the catalytic action of certain enzymes (Biology Online Dictionary, 2020).
- Glycan: any polymer consisting of more than about 10 monosaccharide (i.e. glycose) residues joined by glycosidic linkages (Oxford Dictionary of Biochemistry and Molecule Biology, 2006).
- Convalescent serum: serum from patients recently recovered from a disease; useful in preventing or modifying by passive immunization the same disease in exposed susceptible individuals (Biology Online Dictionary, n.d.)
- Trimer: a molecular complex having three components or subunits (Oxford Dictionary of Biochemistry and Molecular Biology, 2006).
- Hemagglutinin: any agglutinin of red blood cells (Oxford Dictionary of Biochemistry and Molecular Biology, 2006).
- Pseudotyping: the process of producing viruses or viral vectors in combination with foreign viral envelope proteins (Definitions.net, n.d.)
- mAbs: abbreviation for monoclonal antibody (Oxford Dictionary of Biochemistry and Molecular Biology, 2006).
- Epitope: any immunological determinant group of an antigen (Oxford Dictionary of Biochemistry and Molecular Biology, 2006).
- Neutralization: The process of neutralizing a pathogen by antibody acting on the receptors or specific antigen (Biology Online Dictionary, 2019).
- Antigenicity: The capacity of a molecule or an antigen to induce an immune response, that is to be recognized by and interact with an immunologically specific antibody or T-cell receptor (Biology Online Dictionary, n.d.).
Outline of article
Abstract & Introduction
- COVID-19 is caused by SARS-CoV-2, which infects the lower respiratory tract and can cause severe acute respiratory syndrome.
- SARS-CoV-2 is a single-stranded positive-strand RNA virus with four structural proteins: the spike protein (S), small protein (E), matrix (M), and nucleocapsid (N).
- Because SARS-CoV-2 enters target cells via the ACE2 receptor, the spike protein determines viral infectivity and transmissibility. Therefore, it is important to closely monitor any changes in the spike protein.
- RNA viruses have exhibited higher mutation rates than DNA viruses, and mutations in the surface protein amino acids can impact the virus's function and its interaction with the host's neutralizing antibodies.
- By May 6, 2020, 329 naturally occurring variants in the spike protein were reported.
- A previous study indicated that the D614G mutation could increase the virus's fatality rate by altering the conformation of the S protein and increasing infectivity.
- Mutations that affect glycosylation of proteins may also affect the viral life cycle, as well as its interaction with the host. It is unknown how glycosylation of the SARS-CoV-2 S protein could impact viral infectivity and interaction with neutralizing antibodies.
- This study was done to better understand the significance of natural variants with amino acid changes. 106 S protein mutants were analyzed for infectivity and reactivity to neutralizing antibodies.
Materials & Methods
- The GISAID database was used to obtain natural variance S protein sequences. Incomplete, redundant, and ambiguous sequences were removed, leaving a total of 13,406 sequences for alignment.
- Wuhan-1 strain (GenBank: MN_908947) was used as a template to select three groups of variants and mutants to construct 106 pseudotyped viruses.
- 26 cell lines were infected with pseudotyped viruses to analyze infectivity. Three human cell lines (293T-hACE2, 293T, and Huh-7) and three non-human primate cell lines (Vero, VeroE6, and LLC-MK2) were selected for further experiments.
- The 106 pseudotyped viruses were tested for infectivity and reactivity to neutralization using chemiluminescence detection.
Results
- 13,406 S protein sequences were selected to be aligned.
- Three groups of variants (naturally occurring sequences reported in GISAID) and mutants (investigative or experimental mutants) were used to construct pseudo types viruses.
- Group A: High-frequency variants and combined variants with D614G across entire S gene, excluding RBD region (29 strains).
- Group B: Includes variants in RBD region (51 strains).
- 8 single mutations (Q239K, V341I, A435S, K458R, I472V, H519P, A831V, and S943T) in groups A and B do not exist on their own, but they do occur in combination with D614G. Pseudotyped viruses of the single mutations were constructed to compare with the double mutants that include D614G.
- Group C: Mutants at 22 putative glycosylation sites (26 strains).
- Group C includes both variants (4) and investigative mutants (22) in order to study the effects of glycosylation.
- A total of 106 pseudotyped viruses (80 variants and 26 glycosylation mutants) were constructed for the study.
- 26 cell lines were infected with pseudotyped viruses with either the SARS-CoV-2 S protein or Vesicular stomatitis virus G (VSV G) protein. The researchers found that the two viruses showed different infectivity.
- Nearly all 26 cell lines were generally susceptible to the VSV G pseudotyped virus. The SARS-CoV-2 pseudotyped virus was able to efficiently infect certain cell lines including three human cell lines (293T-hACE2, 293T, Huh-7) and three non-human primate cell lines (Vero, VeroE6, LLC-MK2).
- 4 cell lines (293T-hACE2, Huh-7, Vero, and LLC-MK2) were selected for further testing.
- The 106 pseudotyped viruses were tested for infectivity in 293T-hACE2 cells.
- 24 were low-infectivity -- 17 natural variants and 7 glycosylation mutants. 13 were located in the RBD region.
- 2 were no-infectivity -- 1 natural variant and 1 glycosylation mutant. Both were located in the RBD region.
- Double glycosylation deletions at N331 and N343 results in a dramatic reduction in infectivity. Single deletions at each site caused a less drastic reduction in infectivity.
- The results suggest that N331 and N343 may be associated with binding to the receptor or maintaining RBD region conformation.
- The remaining 63 natural variants were tested with the other three cell lines.
- In all four cell lines, single and combined D614G variants showed increased infectivity compared to the reference strain. No difference was noted between single or combined variants, further indicating that D614G is responsible for the increase.
- Next, researchers tested the infectious mutants for antigenicity using 13 neutralizing monoclonal antibodies (mAbs).
- Sensitivity to neutralizing mAbs was altered by some changes in the RBD region.
- A475V reduced the sensitivity to mAbs 157, 247, CB6, P2C-1F11, B38, and CA1.
- F490L reduced the sensitivity to mAbs X593, 261-262, H4, and P2B-2F6.
- L452R reduced the sensitivity to mAbs X593 and P2B-2F6.
- Y508H reduced the sensitivity to mAb H014.
- N439K reduced the sensitivity to mAb H00S022.
- A831V reduced the sensitivity to mAb B38.
- D614G+I472V reduced sensitivity to mAb X593.
- D614G+A435S reduced sensitivity to mAb H014.
- Some changes in the RBD region were observed to be more susceptible to neutralization mediated by mAbs.
- V367F, Q409E, Q414E, I468F, I468T, Y508H, and A522V all increased sensitivity to neutralization by mAbs.
- Sensitivity to neutralizing mAbs was altered by some changes in the RBD region.
- Glycosylation mutants were also tested for antigenicity using the same panel of mAbs.
- N165Q increased sensitivity to mAb P2B-2F6.
- N234Q reduced neutralization sensitivity to mAbs 157, 247, CB6, P2C-1F11, H00S022, B38, AB35, and H014.
- These results confirm that glycosylation sites 165 and 234 are important in receptor binding.
- Overall, five of the mAbs (157, 247, CB6, P2C-1F11, and B38) were unable to neutralize variant A475V or mutant N234Q. mAbs X593 and P2B-2F6 were unable to effectively neutralize L452R, V483A, and F490L. P2B-2F6 was effective in neutralizing N165Q. mAb H014 was not able to neutralize N234Q, Y508H, and D614G+A435S. mAbs H4 and 261-262 were not able to neutralize F490L. H00S022 was unable to neutralize N439K or N234Q.
- Lastly, the strains were tested for their sensitivity to ten COVID-19 convalescent sera.
- None of the variants or mutations showed significant changes in sensitivity to any of the ten convalescent sera.
- F490L and H519P showed more than a 4-fold decrease in sensitivity to 3 of the 10 sera.
- Six variants and mutants (N149H, N149Q, N165Q, N354D, N709Q, and N1173Q) showed a 4-fold increase in sensitivity to one or two of the sera. Five of these were glycan deletion mutants.
- When the data of individual convalescent sera were compiled to analyze sensitivity of all variants, no marked difference was observed.
- Modest differences between some of the variants in comparison to the reference strain were observed in their sensitivity to grouped convalescent sera.
- Variants F338L, V367F, I468F, I468T, and V615L showed increased sensitivity to the convalescent sera when compared to the reference strain.
- Single amino acid changes Y145del, Q414E, N439K, G446V, K458N, I472V, A475V, T478I, V483I, F490L and double amino acid changes D614G+Q321L, D614G+I472V, D614G+A831V, D614G+A879S and D614G+M1237I all showed resistance to the tested convalescent sera.
- The differences between glycosylation mutants and the reference strain were found to be statistically significant, although the magnitude in sensitivity is less than 4-fold.
- N331Q and N709Q significantly increased sensitivity to convalescent sera.
Figures
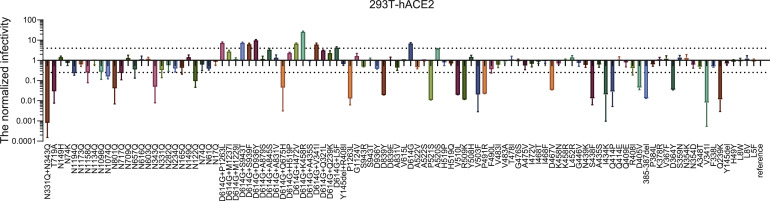
Figure S1
- Figure S1 shows the infectivity analysis of both variants and mutants using the 293T-hACE2 cell line.
- 106 pseudotyped viruses were tested for infectivity. The RLU values resulting from viral infection were quantified by luminescence meter and normalized to the reference strain (Wuhan-1). A 4-fold difference was considered significant. The horizontal dashed lines indicate a 4-fold difference.
- 24 strains were deemed low-infectivity, with the RLU reading decreasing by 4- to 100-fold in comparison with the reference strain.
- Variant V341I and the investigative N331Q+N343Q glycosylation mutant showed no-infectivity, with over 100-fold decreases compared to the reference strain.
- Several of the variants and mutants with a D614G amino acid change, either by itself or in combination with another amino acid change, showed significant increases in infectivity, in comparison with the reference strain.
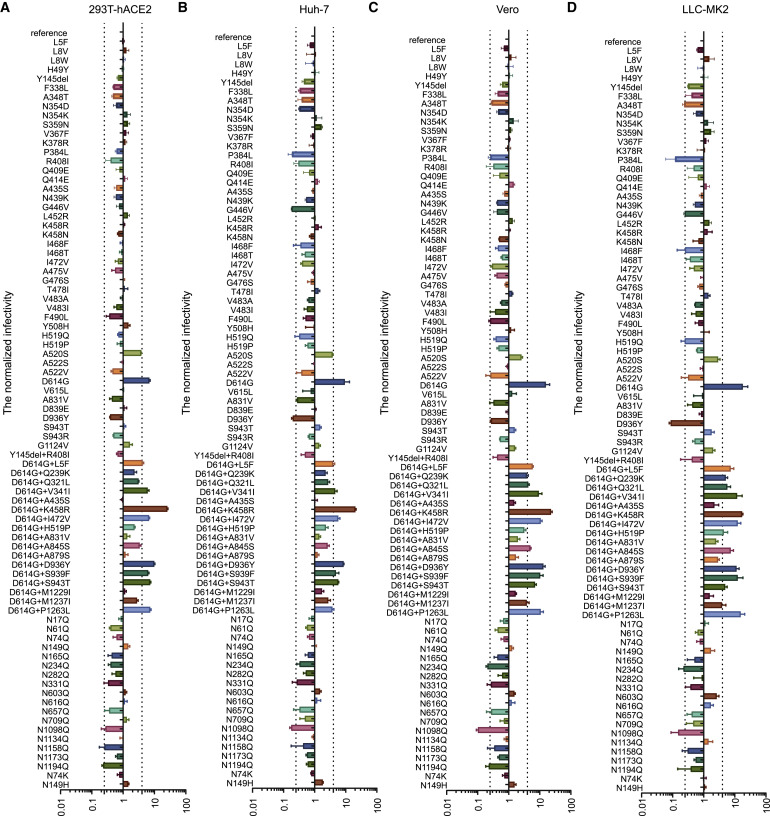
Figure 3
- Figure 3 shows the infectivity analysis of both variants and mutants conducted in all four of the selected cell lines (293T-hACE2, Huh-7, Vero, and LLC-MK2). 293T-hACE2 and Huh-7 are human cell lines, whereas Vero and LLC-MK2 are non-human primate cell lines.
- Again, RLU values resulting from viral infection were quantified by luminescence meter and normalized to the reference strain (Wuhan-1). A difference by 4-fold is considered significant. The vertical dashed lines indicate a 4-fold difference.
- In all four cell lines, D614G variants showed increased infectivity compared to the reference strain. No difference was noted between single or combined variants, further indicating that D614G is responsible for the observed increase.
Discussion
- SARS-CoV-2 is expected to continue to mutate in human populations, so it is important to monitor these changes.
- Through analyzing the 106 pseudotyped viruses, researchers found that variant D614G is a particularly significant change. This variant has been shown to be rapidly accumulating since its emergence. At the beginning of the study, 62.8% of circulating strains had this amino acid change. By the end of the study, 75.7% of strains had the D614G variant. This variant has the potential to increase viral infectivity by 10-fold.
- Most of the natural variants that affected sensitivity to neutralizing mAbs were located in the RBD region because the antibodies were targeting the RBD.
- P2B-2F6 and P2C-1F11 were obtained through using RBD as a bait to isolate RBD-specific B cells in peripheral blood mononuclear cells of SARS-CoV-2 patients.
- P2C- 1F11 and P2B-2F6 bind to overlapping epitope. P2B-2F6 is involved in three hydrophobic interaction sites on RBD.
- Natural variants L452R and F490L showed decreased sensitivity to neutralization by P2B-2F6 mAb.
- Multiple variants with decreased sensitivity to neutralizing mAbs were identified, but it is important to note their frequency in the field.
- V483A in RBD is one of two variants with a frequency over 0.1%. It showed decreased sensitivity to mAbs P2B-2F6 and X593.
- Variant A475V in RBD is significantly resistant to several neutralizing mAbs (P2C-1F11, CA1, 247, and CB6).
- D614G+I472V was the only variant to show increased infectivity, as well as decreased sensitivity to neutralizing mAbs. However, this combination only appeared in one sequence reported in GISAID.
- N439K, L452R, A475V, V483A, F490L, and Y508H all showed decreased sensitivity to neutralizing mAbs, but only V483A exceeded a 0.1% frequency. N439K showed a significant increase in frequency from May to July 2020. Four sequences containing Y508H were reported in GISAID as of July 2020. Each of the other variants were reported in only one or two isolates.
- Continued analyses are necessary to keep up with new mutations and variant frequency.
- The 13 mAbs used in the study appear to have different inhibitory effects on the variants. Using these observations, it would be interesting to study the effects of a therapeutic regimen comprised of at least two mAbs.
- Although the glycosylation mutants are not found in circulation, it is important to note their sensitivity to mAbs.
- N165Q increased sensitivity to mAb P2b-2F6, but N234Q decreased sensitivity to mAbs CA1, CB6, 157, and others. N165 and N234 are both located near the RBD region, so they may impact the epitopes that are targeted by neutralizing mAbs. The N165 glycosylation site is involved in the binding of mAb to the RBD region, and it is likely that the sugar chain can mask the epitope from the antibody. This type of masking has been observed in other viruses such as HIV-1. Therefore, it is possible that the sugar chains are the targets of the neutralizing mAbs. When developing vaccines in bacteria, mammalian cells, or plant expression systems, it is important to consider that different glycosylation sites may impact their effectiveness.
- Testing reactivity to 10 convalescent sera further confirmed the results obtained in the mAbs analyses.
- The magnitude of altered reactivity to convalescent sera is lesser than that of reactivity to mAbs. However, differences in reactivity to human antibodies were still found to be statistically significant.
- RBD variants A475V and F490L are confirmed to have decreased sensitivity to human sera and neutralizing mAbs. A475V showed decreased sensitivity to 6 of the 13 mAbs and F490L had decreased sensitivity to 3 mAbs.
- Analyzing amino acid changes to the entire SARS-CoV-2 spike protein revealed that a variant (D614G) outside the RBD region was the most infectious. However, no evidence of resistance to neutralizing antibodies has been observed. It is important to continue to monitor the circulation of variant D614G. RBD natural variants that demonstrated resistance to neutralizing antibodies should also be closely monitored. Lastly, it was observed that glycosylation sites affect both infectivity and sensitivity to neutralizing antibodies, indicating that glycan could have significant effects on SARS-CoV-2 viral replication and vaccine effectiveness.
Future Directions
- It would be interesting to use this study's findings to further explore the development of a vaccine and therapeutic antibodies. The article mentioned using its findings on the 13 mAbs used in the study to develop a therapeutic regimen. Glycosylation sites were also found to affect both infectivity and antigenicity, which indicates that glycan may have an impact on SARS-CoV-2 viral replication and vaccine-induced immune response.
Critical Evaluation
- Overall, I found the study to be extremely interesting. I enjoyed learning more about the body's immune response to SARS-CoV-2 infection. All of their claims were backed up by solid evidence.
Presentation Slides
Scientific Conclusion
This week's assignment was very helpful in gaining a better understanding of how to sift through a large number of articles and determine which sources are reliable and accurate. This is a very important skill to have when conducting research, and even in everyday life. This process upholds the criteria that scientific sources must meet in order to be trusted.
Acknowledgments
- I spoke with my partners, Yaniv Maddahi, JT Correy, and Fatimah Alghanem over Zoom and text to discuss our journal article and plan our presentation.
- I copied and modified the procedures shown on the Week 11 assignment page.
- I used the Li et al. article for my journal club outline and presentation.
- I defined biological terms using the following sources: Biology Online Dictionary, Oxford Dictionary of Biochemistry and Molecular Biology, and Definitions.net.
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
(Macie Duran (talk) 17:35, 18 November 2020 (PST))
References
- (2006). epitope. In Cammack, R., Atwood, T., Campbell, P., Parish, H., Smith, A., Vella, F., & Stirling, J. (Eds.), Oxford Dictionary of Biochemistry and Molecular Biology. : Oxford University Press. Retrieved 19 Nov. 2020, from https://www.oxfordreference.com/view/10.1093/acref/9780198529170.001.0001/acref-9780198529170-e-6383.
- (2006). glycan. In Cammack, R., Atwood, T., Campbell, P., Parish, H., Smith, A., Vella, F., & Stirling, J. (Eds.), Oxford Dictionary of Biochemistry and Molecular Biology. : Oxford University Press. Retrieved 19 Nov. 2020, from https://www.oxfordreference.com/view/10.1093/acref/9780198529170.001.0001/acref-9780198529170-e-8120.
- (2006). hemagglutinin. In Cammack, R., Atwood, T., Campbell, P., Parish, H., Smith, A., Vella, F., & Stirling, J. (Eds.), Oxford Dictionary of Biochemistry and Molecular Biology. : Oxford University Press. Retrieved 19 Nov. 2020, from https://www.oxfordreference.com/view/10.1093/acref/9780198529170.001.0001/acref-9780198529170-e-8746.
- (2006). Mab. In Cammack, R., Atwood, T., Campbell, P., Parish, H., Smith, A., Vella, F., & Stirling, J. (Eds.), Oxford Dictionary of Biochemistry and Molecular Biology. : Oxford University Press. Retrieved 19 Nov. 2020, from https://www.oxfordreference.com/view/10.1093/acref/9780198529170.001.0001/acref-9780198529170-e-11693.
- (2006). trimer. In Cammack, R., Atwood, T., Campbell, P., Parish, H., Smith, A., Vella, F., & Stirling, J. (Eds.), Oxford Dictionary of Biochemistry and Molecular Biology. : Oxford University Press. Retrieved 19 Nov. 2020, from https://www.oxfordreference.com/view/10.1093/acref/9780198529170.001.0001/acref-9780198529170-e-19978.
- Antigenicity - Biology Online Dictionary. (n.d.). Retrieved November 18, 2020, from https://www.biologyonline.com/dictionary/antigenicity
- Convalescent serum - Biology Online Dictionary. (n.d.). Retrieved November 18, 2020, from https://www.biologyonline.com/dictionary/convalescent-serum
- Glycosylation - Biology Online Dictionary. (2020, August 05). Retrieved November 18, 2020, from https://www.biologyonline.com/dictionary/glycosylation
- Li, Q., Wu, J., Nie, J., Zhang, L., Hao, H., Liu, S., . . . Wang, Y. (2020). The Impact of Mutations in SARS-CoV-2 Spike on Viral Infectivity and Antigenicity. Cell, 182(5). doi:10.1016/j.cell.2020.07.012
- Neutralization - Biology Online Dictionary. (2019, October 07). Retrieved November 18, 2020, from https://www.biologyonline.com/dictionary/neutralization
- Pseudotyping. (n.d.). Retrieved November 18, 2020, from https://www.definitions.net/definition/Pseudotyping