Macie Duran Week 5
Template
User Page
Assignments
- Week 1 Assignment
- Week 2 Assignment
- Week 3 Assignment
- Week 4 Assignment
- Week 5 Assignment
- Week 6 Assignment
- Week 7 Assignment
- Week 8 Assignment
- Week 9 Assignment
- Week 10 Assignment
- Week 11 Assignment
- Week 12 Assignment
- Week 14 Assignment
Journals
- Macie Duran Week 1
- Macie Duran Week 2
- Macie Duran Week 3
- Macie Duran Week 4
- Macie Duran Week 5
- Macie Duran Week 6
- Macie Duran Week 7
- Therapeutic Target Database (TTD) Review
- Macie Duran Week 10
- Macie Duran Week 11
- The Mutants Week 12
- The Mutants Week 14
Class Journals
- Class Journal Week 1
- Class Journal Week 2
- Class Journal Week 3
- Class Journal Week 4
- Class Journal Week 5
- Class Journal Week 6
- Class Journal Week 7
- Class Journal Week 8
- Class Bibliography Week 10
- Class Journal Week 11
- Class Journal Week 14
Purpose
The purpose of this lab is to analyze the sequences and structures of proteins, as well as their interactions with one another. Having an understanding of how amino acid sequences interact with one another allows us to analyze the binding affinity between viral spike proteins and the host cell being infected.
Methods & Results
Exploring the Spike Protein Structure
I navigated to UniProt KB and searched the database using the keyword "SARS-CoV-2".
- 1,598 unreviewed and 18 reviewed results came up.
- Not all of the results are viral proteins. Some other proteins include mouse ACE2 proteins and human ACE2 proteins.
I selected the entry with the accession number P0DTC2, which corresponds with the spike glycoprotein precursor for SARS-CoV-2.
- This entry includes information about the spike protein such as its function, structure, and interaction with different ACE2 receptors. The function of spike protein S1 is to initiate infection by attaching the virion to the host cell receptor. Spike protein S2 acts as a class I viral fusion protein and mediates fusion of the virion and hot's cell membrane. Spike protein S2' is a viral fusion peptide that is unmasked following S2 cleavage, which occurs during the virus's endocytic entry into the host cell.
I used the NCBI Structure Database web-based iCn3D viewer to analyze the structure of the SARS-CoV RBD (2002) complexed with the human ACE2 receptor (accession number 2AJF), which was the structure used in Figure 1A of the Wan et al. paper.
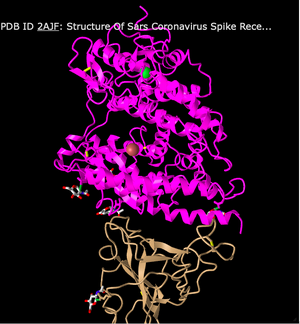
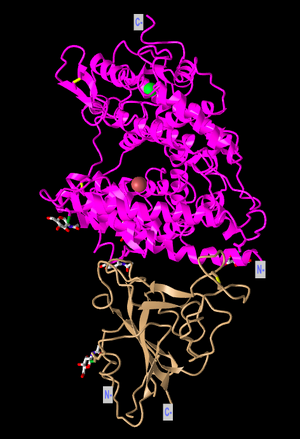
- I attempted to recreate the view of the structure that was used in Figure 1A of the Wan et al. paper by rotating the model. Here is a screenshot of my recreation:
- The structure above is a quaternary structure because it consists of two different polypeptide chains. There are two domains in the structure, as one is the ACE2 receptor and one is the SARS spike protein.
Next, I clicked on the Style > Proteins menu and changed the style multiple times.
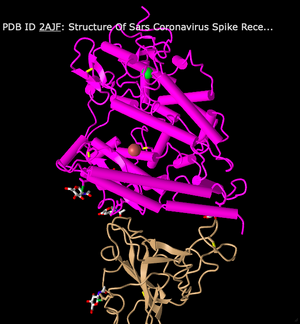
- First, I changed the style to Cylinder & Plate. This style is unique in that it changed the alpha-helix structure to a solid cylinder and the beta-sheets are changed to plates. The plates don't look too different from the sheets, but the cylinders make quite a difference in visualization.
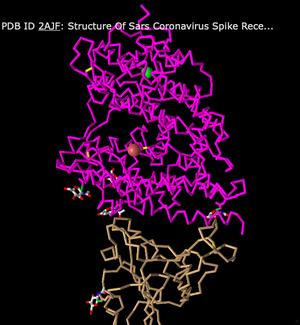
- Next, I switched to a C Alpha Trace style. This style shows less differentiation between alpha-helices and beta-sheets. Instead, the structure now looks more like a series of zig-zags.
- The third style I looked at was Lines. This style shows all of the individual peptide chains, rather than displaying a higher structure level like a helix or sheet. This can make the structure look a bit messier overall.
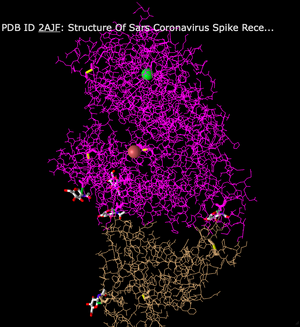
- Then I looked at the Ball and Stick style. The Ball and Stick style is similar to the lines style, but there are balls on the end of each chain. In my opinion, this added to the messiness of the lines style and made it even more difficult to visualize the structure.
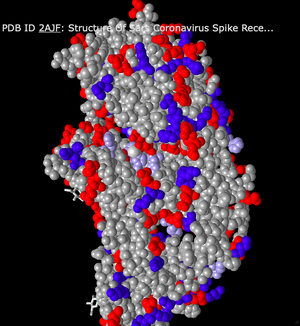
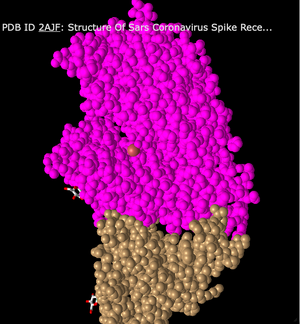
- Lastly, I changed the style to Sphere. This style shows all of the peptide chains as spheres clumped together. There is no differentiation between sheets and helices. Overall, the model just looks like a big mound of spheres.
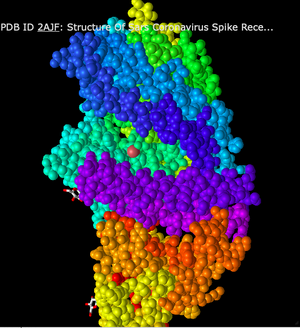
Staying in the Sphere style, I clicked on the Color menu and changed the color scheme multiple times.
- First, I selected the Spectrum color scheme. This altered the model so each secondary structure was color coded. Because the sphere style is a bit clustered and difficult to differentiate each chain, this was helpful in getting a better idea of the overall structure.
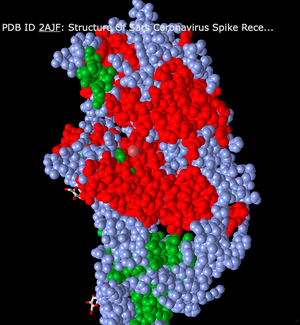
- Next, I changed the color scheme to Secondary. This made the alpha-helix structures all red and the beta-sheets are all coded green. The rest of the structure is a grey-blue color.
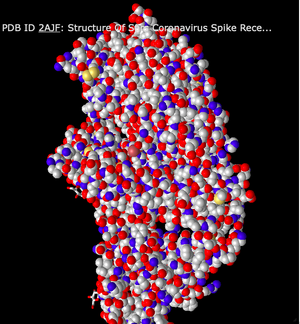
- The third color scheme I looked at was Charge. This color scheme displays the different charges of the protein chains.
- The last color scheme I looked at was Atom. This shows the charged ends of the atoms in the protein.
- Next, I clicked on the Analysis > Label menu and selected N- & C- termini to view the termini locations. I also restored the color scheme and style to the original settings.
- The human ACE2 receptor has alpha-helix structures and the human SARS-CoV RBD has beta-sheets. This can be seen in the Ribbon style, where the top portion (ACE2) highlighted in pink is made up of coils and the bottom portion (SARS-CoV RBD) in the tan color has sheets.
3SCK civet ACE2 spike protein
Next, I looked at the civet ACE2-spike protein structure used in Figure 4B of the Wan et al. paper. Again, I used iCn3D to observe the structure.
- I clicked on the Analysis menu to View Sequences & Annotations.
- In the new window that appeared, I clicked on the Details tab to display the amino acid sequences. There are two sets of spike proteins.
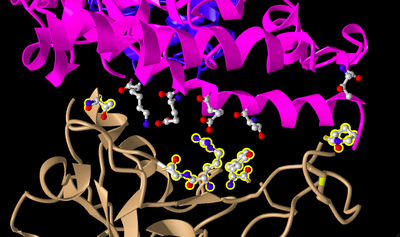
- I focused on the pink and tan chains, and oriented them to mimic Figure 4B in the Wan et al. paper.
- In the sequence window, I looked at sequence Protein 3SCK_A and selected the following amino acids:
- T31
- E35
- E38
- T82
- K353
- The selected amino acids showed up on the model, highlighted in yellow.
- I clicked on the Styles menu and selected Ball and Stick.
- Then, I clicked on the Color menu and selected Atom. This caused the side chains to show in the figure.
- I then selected the following amino acids for the tan spike protein sequence 3SCK_E:
- T487
- R479
- G480
- Y442
- P472
- Again, I changed the style to Ball and Stick and the color to Atom:
- E35 on ACE2 forms an ionic bond with R479 on the spike protein. E35 is a glutamic acid and R479 is arginine. Glutamic acid is acidic and arginine is basic. In the figure, E35 is shown in red (nitrogen) and R479 is blue (oxygen). Nitrogen is typically negatively charged and oxygen is positively charged, which aligns with the glutamic acid being acidic/negatively charged and the arginine being basic/positively charged.
- T31 on ACE2 forms a hydrogen bond with Y442 on the spike protein. T31 is a threonine and Y442 is a tyrosine. Both threonine and tyrosine have uncharged polar side chains. In the figure, Y442 has an aromatic ring and an -NH and -OH group. T31 is also polar because it has an -OH group.
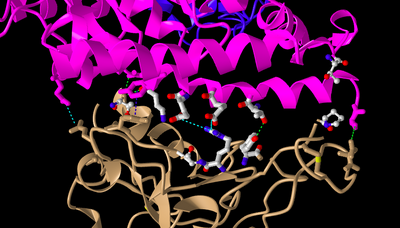
- I clicked on the Analysis menu and selected H bonds & Interactions.
- In Part 1 of the window that appeared, I unchecked Contacts/Interactions.
- In Part 2 of the window, I selected "3SCK_A".
- In Part 3, I selected "3SCK_E".
- In Part 4, I clicked "3D Display".
- This caused the dashed lines representing ionic bonds and hydrogen bonds to appear between the ACE2 receptor and the spike protein.
Beginning your research project
- What question will you answer about sequence-->structure-->function relationships in the spike and/or ACE2 protein?
- How well conserved are ACE2 amino acid sequences in different mammals, and how does this indicate the sequence's importance in function?
- What sequences will you use?
- We will be analyzing the ACE2 receptors obtained from rabbit, cat, ferret, pig, mouse, rat, chimp, bat, civet, and human.
Scientific Conclusion
Analyzing and visualizing the residue interactions between viral spike proteins and ACE2 receptors allows us to gain a better understanding of how viruses are able to enter and infect host cells. I was able to improve my ability to use and manipulate models in order to enhance visualization of the structures.
Acknowledgements
- I spoke with my homework partner, Nathan R. Beshai, over Zoom to discuss our research project.
- I copied and modified the protocol outlined on the Week 5 Assignment page.
- I used the Wan et al. article to reference the figures.
- I used UniProt Knowledgebase to find entries for protein sequences.
- I used the NCBI iCn3D viewer to visualize and analyze protein structures.
- I looked at the Foldit website and read the blog post to learn about how Foldit is contributing to coronavirus research.
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
(Macie Duran (talk) 21:22, 7 October 2020 (PDT))
References
- Foldit - Blogs. (2020). Retrieved 7 October 2020, from https://fold.it/portal/blog
- Foldit - Solve Puzzles for Science. (2020). Retrieved 7 October 2020, from https://fold.it/portal/
- iCn3D: Web-based 3D Structure Viewer 2AJF. (2020). Retrieved 1 October 2020, from https://www.ncbi.nlm.nih.gov/Structure/icn3d/full.html?&mmdbid=35213&bu=1&showanno=1
- iCn3D: Web-based 3D Structure Viewer 3SCK. (2020). Retrieved 7 October 2020, from https://www.ncbi.nlm.nih.gov/Structure/icn3d/full.html?pdbid=%203SCK
- OpenWetWare. (2020). BIOL368/F20:Week 5. Retrieved 1 October 2020, from https://openwetware.org/wiki/BIOL368/F20:Week_5
- Uniprot. (2020). S - Spike glycoprotein precursor - Severe acute respiratory syndrome coronavirus 2 (2019-nCoV) - S gene & protein. Retrieved 1 October 2020, from https://www.uniprot.org/uniprot/P0DTC2
- Wan, Y., Shang, J., Graham, R., Baric, R., & Li, F. (2020). Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. Journal Of Virology, 94(7). doi: 10.1128/jvi.00127-20