Desireegonzalez Week 2
From OpenWetWare
Jump to navigationJump to search
Electronic Lab Notebook
Purpose
- The purpose of this weekly assignment is to gain a greater understanding of the experimental process of using DNA microarrays. This increased understanding will be formed by viewing data samples in literature and reading articles that explain how to analyze and interpret the data collected using DNA microarray methodology.
Discovery Questions from Chapter 4
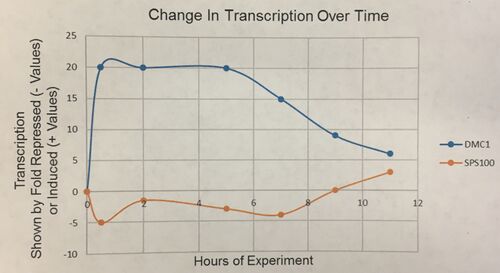
- (Question 5, p. 110) Choose two genes from Figure 4.6b (PDF of figures on Brightspace) and draw a graph to represent the change in transcription over time. You can either create your plot in Excel and put the image up on your wiki page or you can do it by hand and upload a picture or scan.
- (Question 6b, p. 110) Look at Figure 4.7, which depicts the loss of oxygen over time and the transcriptional response of three genes. These data are the ratios of transcription for genes X, Y, and Z during the depletion of oxygen. Using the color scale from Figure 4.6, determine the color for each ratio in Figure 4.7b. (Use the nomenclature "bright green", "medium green", "dim green", "black", "dim red", "medium red", or "bright red" for your answers.)
- gene X: At hour 1 (ratio of 1.0) the color is black. At hour 3 (ratio of 2.2) the color is dim red. At hour 5 (ratio of 1.0) the color is black. At hour 9 (ratio of 0.15) the color is dim green.
- gene Y: At hour 1 (ratio of 1.0) the color is black. At hour 3 (ratio of 4.5) the color is medium red. At hour 5 (ratio of 0.95) is black. At hour 9 (ratio of 0.05) is medium green.
- gene Z: At hour 1 (ratio of 1.0) the color is black. At hour 3 (ratio of 1.5) the color is dim red. At hour 5 (ratio of 2.0) is dim red. At hour 9 (ratio of 2.0) is dim red.
- (Question 7, p. 110) Were any of the genes in Figure 4.7b transcribed similarly? If so, which ones were transcribed similarly to which ones?
- There were genes in Figure 4.7b that were transcribed similarly. Genes X and Y were transcribed similarly to each other since their ratio data fell in similar ranges and colors on the fold repressed and fold induced color spectrum.
- (Question 9, p. 118) Why would most spots be yellow at the first time point? I.e., what is the technical reason that spots show up as yellow - where does the yellow color come from? And, what would be the biological reason that the experiment resulted in most spots being yellow?
- Most spots would initially be yellow at the first time point since the fluorescently labeled cDNAs would not have yet bound to the DNA microarray in the short amount of time. The yellow color appears when the gene expresses both conditions given in the labeled cDNAs. The biological reasoning that the experiment were to result in most spots being yellow is that all genes would have an equal expression of the given characteristic labeled by the fluorescent cDNAs.
- (Question 10, p. 118) Go to the Saccharomyces Genome Database and search for the gene TEF4; you will see it is involved in translation. Look at the time point labeled OD 3.7 in Figure 4.12, and find the TEF4 spot. Over the course of this experiment, was TEF4 induced or repressed? Hypothesize why TEF4’s change in expression was part of the cell’s response to a reduction in available glucose (i.e., the only available food).
- Over the course of this experiment, the TEF4 was repressed since its dot became more green in color the longer the experiment proceeded. TEF4's change in expression to repressed can be hypothesized to occur since it no longer has glucose as a source of energy to keep up transcription within the cell. A decrease in transcription would be revealed as a repressed gene; especially if the glucose was needed to run the transcription due to the presence of a repressor (I am thinking about the Lac Operon that I learned about in other Biology courses).
- (Question, 11, p. 120) Why would TCA cycle genes be induced if the glucose supply is running out?
- TCA cycle genes would be induced if the glucose supply is running out because the TCA cycle would help in the formation of more NADH and FADH2 and ATP. The glucose supply that would be running out (due to its use in glycolysis) would also result in pyruvate that would then enter the TCA cycle to keep forming NADH, FADH2, and ATP.
- (Question 12, p. 120) What mechanism could the genome use to ensure genes for enzymes in a common pathway are induced or repressed simultaneously?
- The mechanism of guilt by association would help to ensure the genes for enzymes in a common pathway are induced or repressed simultaneously. Guilt by association states that genes with similar expressions have similar promoters and vice versa.
- (Question 13, p. 121) Consider a microarray experiment where cells deleted for the repressor TUP1 were subjected to the same experiment of a timecourse of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later timepoints (glucose depleted) are labeled red. What color would you expect the spots that represented glucose-repressed genes to be in the later time points of this experiment?
- The color that I would expect the spots that represented glucose-repressed genes to be in the later time points of this experiment, is red (signifying induction). The TUP1 gene is responsible for repressing glucose-repressed genes. Therefore, it is easy to identify that a deletion of the TUP1 gene would not allow repression of glucose-repressed cells to occur.
- (Question 14, p. 121) Consider a microarray experiment where cells that over express the transcription factor Yap1p were subjected to the same experiment of a time course of glucose depletion where cells at t0 (plenty of glucose available) are labeled green and cells at later timepoints (glucose depleted) are labeled red. What color would you expect the spots that represented Yap1p target genes to be in the later timepoints of this experiment?
- Yap1p is a transcription factor that gives resistance to environmental stressors. I believe that the color of the spots that represented Yap1p target genes would be is red (signifying induced) in later times in this experiment. I would guess that the color of the spots would be red since the over-expression characteristic of resistance to environmental factors in this gene would keep transcription occurring even as levels of glucose begin to deplete.
- (Question 16, p. 121) Using the microarray data, how could you verify that you had truly deleted TUP1 or over-expressed YAP1 in the experiments described in questions 8 and 9?
- Using microarray data of both the mutated (deleted TUP1 or over-expressed YAP1) genes and comparing it to microarray data of non-mutated (wild type) versions of both genes could assist you in verifying the presence of your mutation in the experiments described in questions 8 and 9.
Acknowledgements
- I sent an email to Dr. Dahlquist and Dr. Fitzpatrick to ask a question on how to create the graph for the first question and to gain a better understanding of how to be able to determine when genes are similar in transcription, by reading the color spectrum of fold repressed or induced genes. I also text with my homework partner Alison various times during the week to speak about how we would answer some of the questions. In addition to text messaging with my homework partner, I also text messaged Ava during the week, to speak about how to read the color scales that represented the fold of repressed or induced genes; we also spoke about how to answer some of the harder questions on this assignment.
Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
Desireegonzalez (talk) 18:02, 30 January 2019 (PST)
References
Campbell A.M. & Heyer, L.J. (2003), "Chapter 4: Basic Research with DNA Microarrays" in Discovering Genomics, Proteomics, and Bioinformatics, San Francisco, CA: Benjamin Cummings, pp. 107-124.
Important Links
Below are the links to all the Assignments and Journal Entries of the Spring 2019 Semester.
User Page: user:desireegonzalez
Template Page: template:desireegonzalez
Weekly Assignment Pages:
Individual Journal Entry Pages:
- desireegonzalez Week 1
- desireegonzalez Week 2
- desireegonzalez Week 3
- desireegonzalez Week 4/5
- desireegonzalez Week 6
- desireegonzalez Week 7
- desireegonzalez Week 9
- desireegonzalez Week 10
- desireegonzalez Week 11
- desireegonzalez Week 12
- desireegonzalez Week 15
Shared Journal Pages: