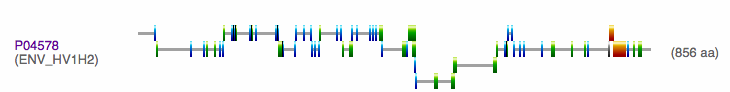Nicolette S. Harmon Week 8
Bioinformatics for Dummies Exercises
Chapter 2: Retrieving Protein Sequences
- To start I went to the The Swiss-Prot Data homepage.
- Since HIV is of interest, HIV gp120 was typed into the search.
- Clicking on one of the search results gives you a background about the protein selected.
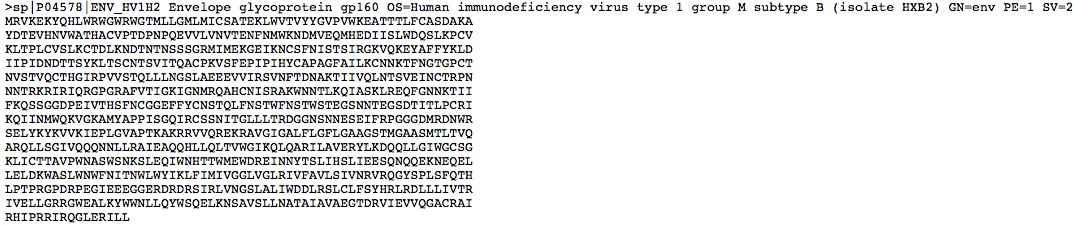
- Then the FASTA link on the top right of the page was clicked in order to get the actual protein sequence.
Chapter 4: Reading a Swiss-Prot Entry
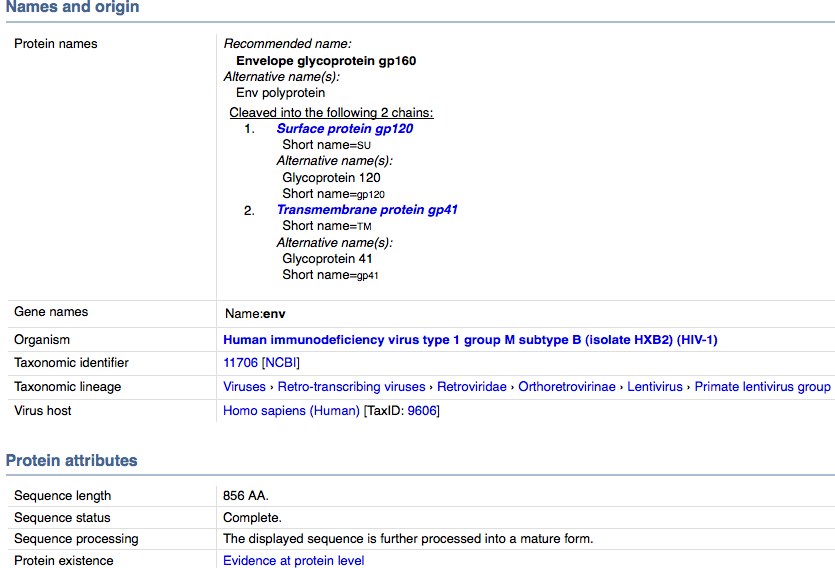
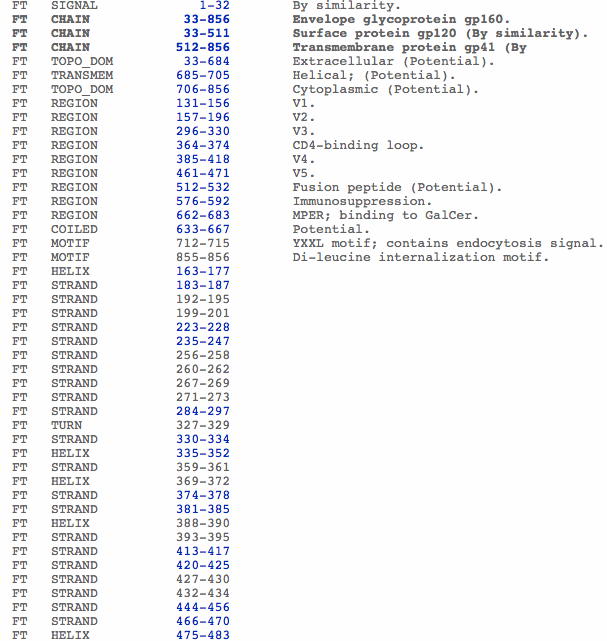
- When viewing the HIV gp120 Page that was searched in the above section, you can view the entry information all the way at the bottom of the page.
- This page gives information such as taxonomy, gene name, accession numbers for the sequence, function and other information displayed below.
Chapter 5: ORFing your DNA Sequence
- I went to the ORF Finder website.
- I copied and pasted my FASTA text into the text box.
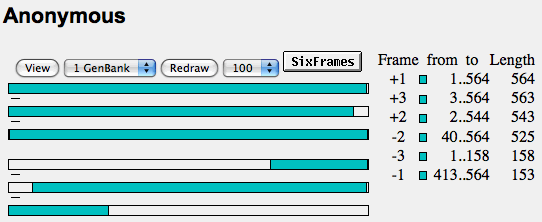
- Below are the results when I clicked the OrfFind button.
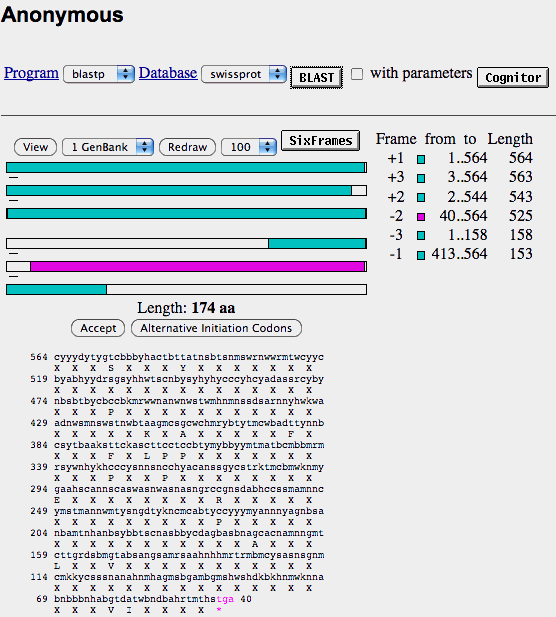
- When you click on on of the colored rectangles on the right-hand side of the screen, the program gives you the predicted amino-acid sequence as well as access to other databases that may have similar sequences to the one that ORF gives you.
Chapter 6: Working with A Single Protein Sequence
- For this chapter, I started by going to the ProtParam link.
- The accession number taken from SwissProt was entered into this database and below is a picture of the intermediate page, when you specify the segment of protein you want to look at you type the numbers in the N and C- terminus boxes.
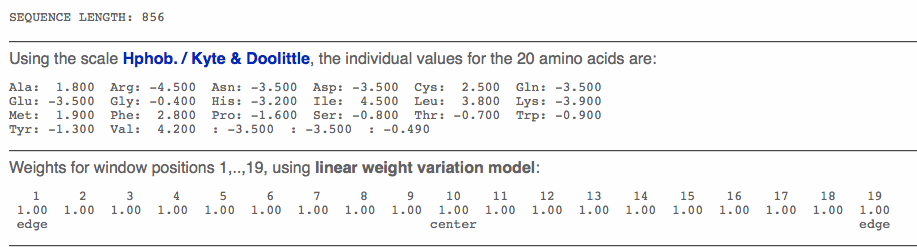
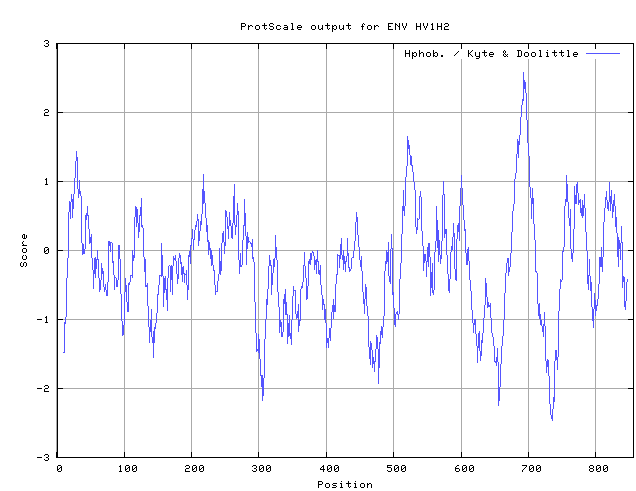
- The next program used was Protscale.
- Again the accession number from above was entered into the appropriate box.
- Below is the results page that shows the Kyte and Doolittle Hydrophobicity Scale.
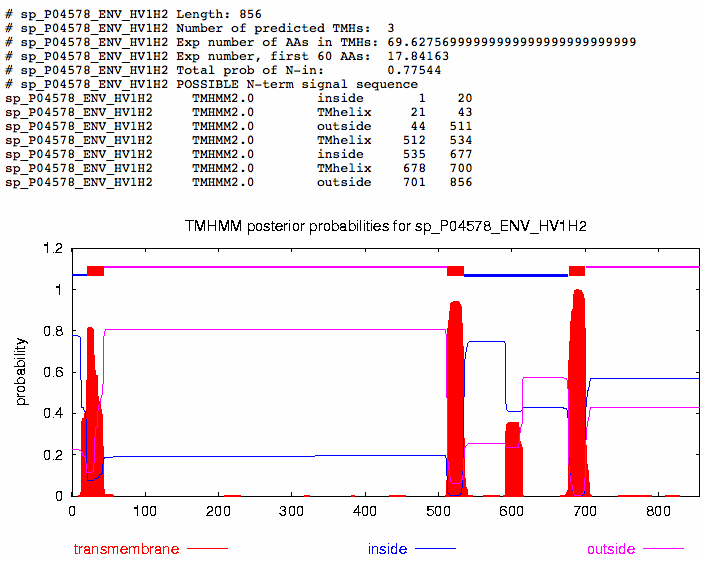
- The next program I used was TMHMM
- The FASTA version of the HIVgp120 sequence is required for this website, so that was copied and pasted into the text box.
- When I pressed submit, this is the page that appeared on my screen.
- Next, I used the ScanProsite page.
- When I typed in the accession number in the left-hand textbox, I unchecked the "Exclude Motifs with a High Probability of Occurrence" box and checked the "Do Not Scan Profiles" box.
- Below is the image generated by this page, but the full results can be seen here.
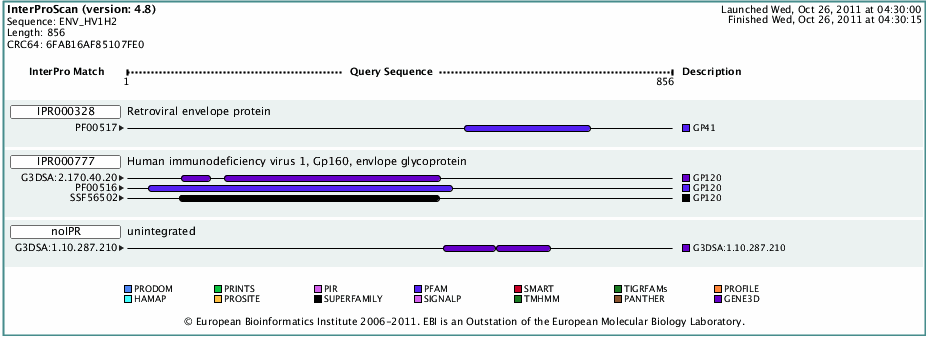
- Next I found the domains of HIV gp120 using InterProScan.
- Again I entered the FASTA formatted version of the HIV gp120 sequence into the textbox, below is the results.
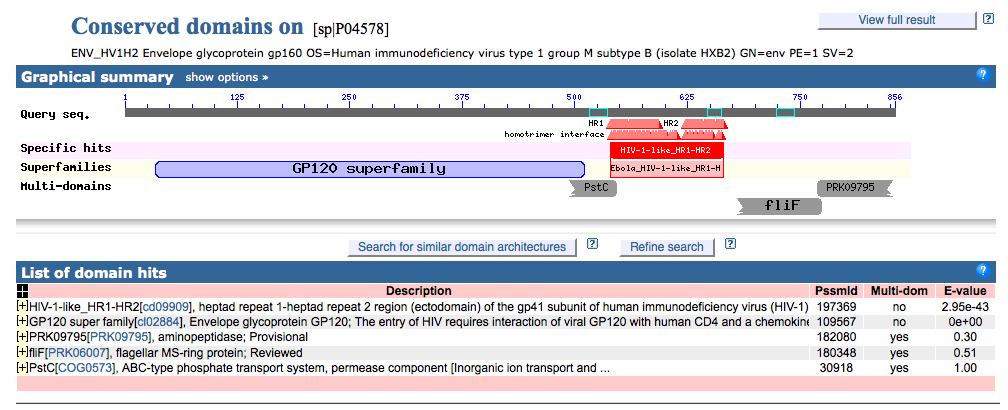
- To find the domains with the CD server, you can go to this page.
- Before clicking "Submit Query" uncheck the "Apply Low Complexity Filter" box and change the "Expect Value Threshold" to 1 instead of 0.1.
- Lastly, you can find domains using Motif Scans.
HIV Research Project
My partner for this research project is Samantha Hurndon. We will be examining the structures of the nonprogressor types and some of the moderate progressors from the markham study we used during the HIV Evolution project. The specific subjects we will be examining are 2,8,12,13 as they are subjects who were either classified as moderate progressors or nonprogressors. We will use the Star Biochem tool to analyze the protein structures.
Links
BIOL368/F11:Class Journal Week 1
BIOL368/F11:Class Journal Week 2
BIOL368/F11:Class Journal Week 3
BIOL368/F11:Class Journal Week 4
BIOL368/F11:Class Journal Week 5
BIOL368/F11:Class Journal Week 6
BIOL368/F11:Class Journal Week 7
BIOL368/F11:Class Journal Week 8
BIOL368/F11:Class Journal Week 9
BIOL368/F11:Class Journal Week 10
BIOL368/F11:Class Journal Week 11
BIOL368/F11:Class Journal Week 12