Nicolette S. Harmon Week 6
DNA Glycosylase Exercise
hOGG1's Protein Structure Basics
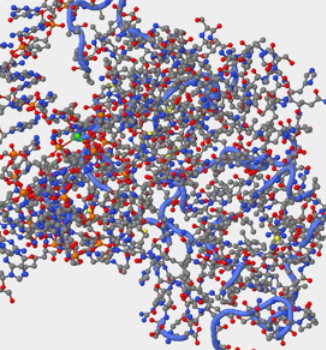
- Question 1:Identify the DNA and the hOGG1 protein in the structure. Where is the DNA segment?
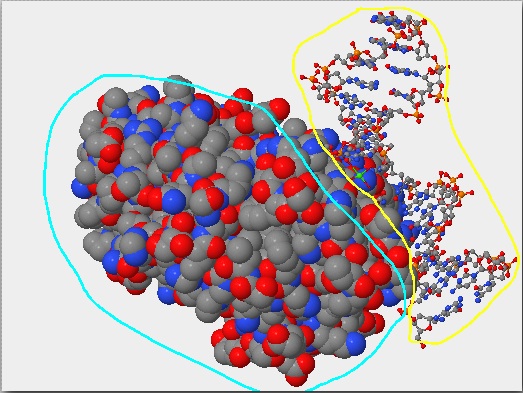
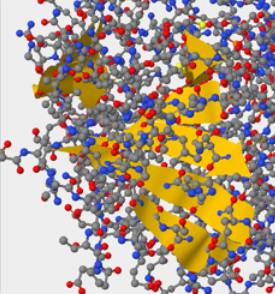
- Answer:When the atom size under the primary tab is increased to 100, you can clearly see the difference between the protein and the DNA. The hOOG1 protein is circled in the color Turquoise and the DNA double helix is circled in the color Yellow. The DNA interacts with an amino group on the protein through it's backbone which is negatively charged.
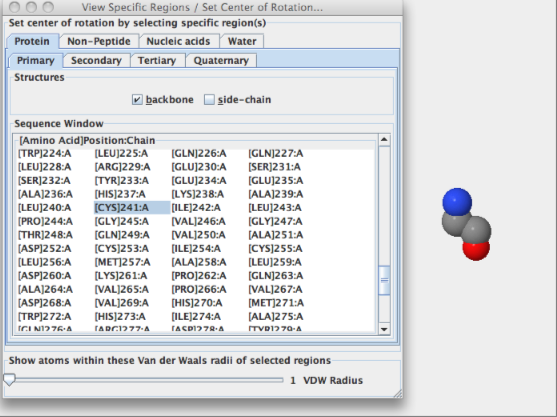
- Question 2: a)Identify the name and sequence number of on of the amino acids in the structure that contains a sulfur atom.
b)Is the sulfur atom located in the backbone or in the side chain of the amino acid?
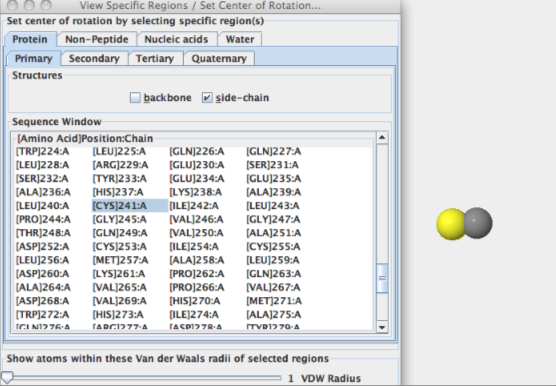
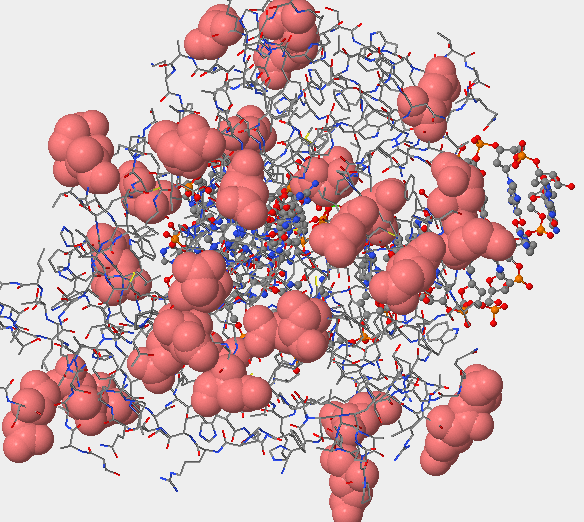
- Answers: a)[Cys]241:A SG #2406
b)These images above display that the sulfur atom is in the side chain of the amino acid, with Sulfur being the atom colored in yellow.
- Question 3:List the 13 amino acids numbered 105-117 in order.
- Answer:
105-Threonine 106-Leucine 107-Alanine 108-Glutamine 109-Leucine 110-Tyrosine 111-Histidine 112-Histidine 113-Tryptophan 114-Glycine 115-Serine 116-Valine 117-Aspartic Acid
- Question 4: a)Are helices, sheets or coils present in hOGG1? Describe the color that represents each secondary structure you observe.
b)Which secondary structure do amino acids 105-117 fold into?
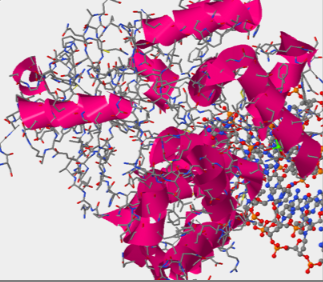
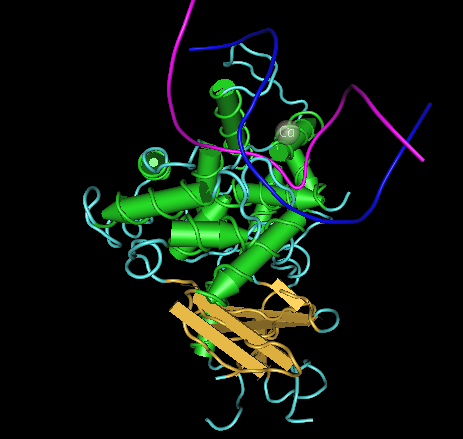
- Answers: a)Helices, sheets, and coils are all present within thehOGG1 protein. Helices are pink, sheets are yellow, and coils are blue.
b)Amino acids 105-117 fold into the helical structure.
- Question 5: Are the negatively charged amino acids located on the inside(buried) or outside(exposed) of this protein? What does that suggest about the cellular environment surrounding this protein, is it hydrophobic or hydrophilic?
- Answer: The negatively charged amino acids are located on the outside of this protein suggesting the cellular environment surrounding this protein is hydrophilic.
hOGG's Interaction with DNA
- I was unable to obtain screenshots for the following questions due to program malfunctions.
- Question 6: a)How many DNA base pairs can you count within this double helix?
b)How many DNA bases are unpaired (not paired to its partner on the other strand)?
c)Is the oxidized guanine base paired or unpaired? Describe the position of the oxidized guanine with respect to the hOGG1 protein and the double helix. What does this suggest about the mechanism that hOGG1 uses to identify damaged DNA bases?
- Answers: a) There were 14 base pairs.
b) Only 1 unpaired bases was seen. c)The oxidized guanine was unpaired.
- Question 7: Where are you more likely to find the amino acids that recognize damaged guanine bases within hOGG1, in helix 1 of helix 16?
- Answer: Helix 16 is the more likely place due to it's close proximity to the guanine bases.
Cn3D
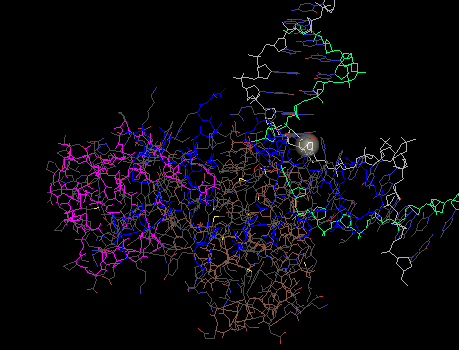
- This structure is a quarternary structure, this is because there is more than one polypeptide chain in the image shown below.
- DNA glycosylase has three domains. This can be seen on this program by selecting "Domain" in the coloring tool, this tool colors the domains in different colors.
- It is possible to view the same things in the structure using both programs, however it's a little more challenging to figure out how to use Cn3D. Star Biochem has everything you need clearly visible, while you have to search around on Cn3D to figure out how to get the same results as Star Biochem.
The image above is similar to the first screen shot on this page, it shows where the DNA is and where the protein is.
- I definitely prefer to use Star Biochem bascially for the reason i said in the answer above. Everything you need is clearly labeled and visible and you don't have to spend a lot of time figuring out how to use the program.
Links
BIOL368/F11:Class Journal Week 1
BIOL368/F11:Class Journal Week 2
BIOL368/F11:Class Journal Week 3
BIOL368/F11:Class Journal Week 4
BIOL368/F11:Class Journal Week 5
BIOL368/F11:Class Journal Week 6
BIOL368/F11:Class Journal Week 7
BIOL368/F11:Class Journal Week 8
BIOL368/F11:Class Journal Week 9
BIOL368/F11:Class Journal Week 10
BIOL368/F11:Class Journal Week 11
BIOL368/F11:Class Journal Week 12