Nathan R Beshai Week 6
Nathan R. Beshai User Page
Nathan R. Beshai Template Page
Course assignments
Individual journal assignments
- Nathan R Beshai Week 2
- Nathan R Beshai Week 3
- Nathan R Beshai Week 4
- Nathan R Beshai Week 5
- Nathan R Beshai Week 6
- Nathan R Beshai Week 7
- Nathan R Beshai Week 8
- Nathan R Beshai Week 9
- Nathan R Beshai Week 10
- Nathan R Beshai Week 11
- The D614G Research Group Week 12
- The D614G Research Group Week 14
Class Journals
- Class Journal 1
- Class Journal 2
- Class Journal 3
- Class Journal 4
- Class Journal 5
- Class Journal 6
- Class Journal 7
- Class Journal 8
- Class Journal 9
- Class Journal 10
- Class Journal 11
- Class Journal 12
- Class Journal 14
Link to Brightspace and LMU's Homepage
Purpose
To see how closely related species' ACE2 sequences that are more compatible with the 2019-nCoV spike protein are, to see how well conserved the necessary human ACE2 amino acid residues are across the species' sequence, and to see if any of the human 5 amino acid residues that bind to the 2019-nCoV spike protein are necessary for the ACE2 function and structure.
Methods and results
1. Searched and collected the ACE-2 protein sequences from UniProt and if they were not available collected the sequences from the NCBI database.
1.Went to UniProt and copied the ACE-2 sequence for the Pig (unreviewed).
>tr|A0A220QT48|A0A220QT48_PIG Angiotensin-converting enzyme OS=Sus scrofa domesticus OX=9825 GN=ACE2 PE=2 SV=1 MSGSFWLLLSLIPVTAAQSTTEELAKTFLEKFNLEAEDLAYQSSLASWNYNTNITDENIQ KMNDARAKWSAFYEEQSRIAKTYPLDEIQTLILKRQLQALQQSGTSGLSADKSKRLNTIL NTMSTIYSSGKVLDPNNPQECLVLEPGLDEIMENSKDYSRRLWAWESWRAEVGKQLRPLY EEYVVLENEMARANNYEDYGDYWRGDYEVTGTGDYDYSRNQLMEDVERTFAEIKPLYEHL HAYVRAKLMDAYPSRISPTGCLPAHLLGDMWGRFWTNLYPLTVPFGEKPSIDVTEAMVNQ SWDAIRIFEEAEKFFVSIGLPNMTQGFWNNSMLTEPGDGRKVVCHPTAWDLGKGDFRIKM CTKVTMDDFLTAHHEMGHIQYDMAYAIQPYLLRNGANEGFHEAVGEIMSLSAATPHYLKA LGLLPPDFYEDSETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKEQWMQKWWEM KREIVGVVEPLPHDETYCDPACLFHVAEDYSFIRYYTRTIYQFQFHEALCRTAKHEGPLY KCDISNSTEAGQKLLQMLSLGKSEPWTLALENIVGVKTMDVKPLLSYFEPLLTWLKAQNG NSSVGWNTDWTPYADQSIKVRISLKSALGKEAYEWNDNEMYLFRSSIAYAMRNYFSSAKN ETIPFGAEDVWVSDLKPRISFNFFVTSPANMSDIIPRSDVEKAISMSRSRINDAFRLDDN TLEFLGIQPTLGPPDEPPVTVWLIIFGVVMGLVVVGIVVLIFTGIRDRRKKKQASSEENP YGSMDLSKGESNSGFQNGDDIQTSF
2.Went to NCBI and copied the ACE-2 sequence for the Ferrat.
>BAE53380.1 angiotensin I converting enzyme 2 [Mustela putorius furo] MLGSSWLLLSLAALTAAQSTTEDLAKTFLEKFNYEAEELSYQNSLASWNYNTNITDENIQKMNIAGAKWS AFYEEESQHAKTYPLEEIQDPIIKRQLRALQQSGSSVLSADKRERLNTILNAMSTIYSTGKACNPNNPQE CLLLEPGLDDIMENSKDYNERLWAWEGWRSEVGKQLRPLYEEYVALKNEMARANNYEDYGDYWRGDYEEE WADGYSYSRNQLIEDVEHTFTQIKPLYEHLHAYVRAKLMDAYPSRISPTGCLPAHLLGDMWGRFWTNLYP LMVPFRQKPNIDVTDAMVNQSWDARRIFEEAETFFVSVGLPNMTEGFWQNSMLTEPGDNRKVVCHPTAWD LGKRDFRIKMCTKVTMDDFLTAHHEMGHIQYDMAYAEQPFLLRNGANEGFHEAVGEIMSLSAATPNHLKN IGLLPPDFSEDSETDINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKEQWMQKWWEMKRDIVGVVEP LPHDETYCDPAALFHVANDYSFIRYYTRTIYQFQFQEALCQIAKHEGPLYKCDISNSSEAGQKLHEMLSL GRSKPWTFALERVVGAKTMDVRPLLNYFEPLFTWLKEQNRNSFVGWNTDWSPYADQSIKVRISLKSALGE KAYEWNDNEMYFFQSSIAYAMREYFSKVKNQTIPFVGKDVRVSDLKPRISFNFIVTSPENMSDIIPRADV EEAIRKSRGRINDAFRLDDNSLEFLGIQPTLEPPYQPPVTIWLIVFGVVMGVVVVGIFLLIFSGIRNRRK NNQARSEENPYASVDLSKGENNPGFQNVDDVQTSF
3.Went to UniProt and copied the ACE-2 sequence for the Chimpanzee (unreviewed).
>tr|A0A2J8KU96|A0A2J8KU96_PANTR Angiotensin-converting enzyme OS=Pan troglodytes OX=9598 GN=ACE2 PE=3 SV=1 MSGSSWLLLSLVAVTAAQSTIEEQAKTFLDKFNHEAEDLFYQSSLASWNYNTNITEENVQ NMNNAGDKWSAFLKEQSTLAQMYPLQEIQNLTVKLQLQALQQNGSSVLSEDKSKRLNTIL NTMSAIYSTGKVCNPNNPQECLLLEPGLNEIMANSLDYNERLWAWESWRSEVGKQLRPLY EEYVVLKNEMARANHYEDYGDYWRGDYEVNGVDGYDYSRGQLIEDVEHTFEEIKPLYEHL HAYVRAKLMNAYPSYISPIGCLPAHLLGDMWGRFWTNLYSLTVPFGQKPNIDVTDAMVDQ AWDAQRIFKEAEKFFVSVGLPNMTQGFWENSMLTDPGNVQKAVCHPTAWDLGKGDFRILM CTKVTMDDFLTAHHEMGHIQYDMAYAAQPFLLRNGANEGFHEAVGEIMSLSAATPKHLKS IGLLSPDFQEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPEDQWMKKWWEM KREIVGVVEPVPHDETYCDPASLFHVSNDYSFIRYYTRTLYQFQFQEALCQAAKHEGPLH KCDISNSTEAGQKLFNMLRLGKSEPWTLALENVVGAKNMNVRPLLNYFEPLFTWLKDQNK NSFVGWSTDWSPYADQSIKVRISLKSALGDKAYEWNDNEMYLFRSSVAYAMRQYFLKVKN QMILFGEEDVRVANLKPRISFNFFVTAPKNVSDIIPRTEVEKAIRKSRSRINDAFRLNDN SLEFLGIQPTLGPPNQPPVSIWLIVFGVVMGVIVVGIVILIFTGIRDRKKKNKARSEENP YASVDTSKGENNPGFQNTDDVQTSF
4. Went to UniProt and copied the ACE-2 sequence for the Homosapien (human).
>sp|Q9BYF1|ACE2_HUMAN Angiotensin-converting enzyme 2 OS=Homo sapiens OX=9606 GN=ACE2 PE=1 SV=2 MSSSSWLLLSLVAVTAAQSTIEEQAKTFLDKFNHEAEDLFYQSSLASWNYNTNITEENVQ NMNNAGDKWSAFLKEQSTLAQMYPLQEIQNLTVKLQLQALQQNGSSVLSEDKSKRLNTIL NTMSTIYSTGKVCNPDNPQECLLLEPGLNEIMANSLDYNERLWAWESWRSEVGKQLRPLY EEYVVLKNEMARANHYEDYGDYWRGDYEVNGVDGYDYSRGQLIEDVEHTFEEIKPLYEHL HAYVRAKLMNAYPSYISPIGCLPAHLLGDMWGRFWTNLYSLTVPFGQKPNIDVTDAMVDQ AWDAQRIFKEAEKFFVSVGLPNMTQGFWENSMLTDPGNVQKAVCHPTAWDLGKGDFRILM CTKVTMDDFLTAHHEMGHIQYDMAYAAQPFLLRNGANEGFHEAVGEIMSLSAATPKHLKS IGLLSPDFQEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKDQWMKKWWEM KREIVGVVEPVPHDETYCDPASLFHVSNDYSFIRYYTRTLYQFQFQEALCQAAKHEGPLH KCDISNSTEAGQKLFNMLRLGKSEPWTLALENVVGAKNMNVRPLLNYFEPLFTWLKDQNK NSFVGWSTDWSPYADQSIKVRISLKSALGDKAYEWNDNEMYLFRSSVAYAMRQYFLKVKN QMILFGEEDVRVANLKPRISFNFFVTAPKNVSDIIPRTEVEKAIRMSRSRINDAFRLNDN SLEFLGIQPTLGPPNQPPVSIWLIVFGVVMGVIVVGIVILIFTGIRDRKKKNKARSGENP YASIDISKGENNPGFQNTDDVQTSF
5.Went to UniProt and copied the ACE-2 sequence for the Masked Palm civet.
>sp|Q56NL1|ACE2_PAGLA Angiotensin-converting enzyme 2 OS=Paguma larvata OX=9675 GN=ACE2 PE=1 SV=1 MSGSFWLLLSFAALTAAQSTTEELAKTFLETFNYEAQELSYQSSVASWNYNTNITDENAK NMNEAGAKWSAYYEEQSKLAQTYPLAEIQDAKIKRQLQALQQSGSSVLSADKSQRLNTIL NAMSTIYSTGKACNPNNPQECLLLEPGLDNIMENSKDYNERLWAWEGWRAEVGKQLRPLY EEYVALKNEMARANNYEDYGDYWRGDYEEEWTGGYNYSRNQLIQDVEDTFEQIKPLYQHL HAYVRAKLMDTYPSRISRTGCLPAHLLGDMWGRFWTNLYPLTVPFGQKPNIDVTDAMVNQ NWDARRIFKEAEKFFVSVGLPNMTQGFWENSMLTEPGDGRKVVCHPTAWDLGKGDFRIKM CTKVTMDDFLTAHHEMGHIQYDMAYAAQPFLLRNGANEGFHEAVGEIMSLSAATPNHLKT IGLLSPAFSEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGAIPKEQWMQKWWEM KRNIVGVVEPVPHDETYCDPASLFHVANDYSFIRYYTRTIYQFQFQEALCQIAKHEGPLH KCDISNSTEAGKKLLEMLSLGRSEPWTLALERVVGAKNMNVTPLLNYFEPLFTWLKEQNR NSFVGWDTDWRPYSDQSIKVRISLKSALGEKAYEWNDNEMYLFRSSIAYAMREYFSKVKN QTIPFVEDNVWVSDLKPRISFNFFVTFSNNVSDVIPRSEVEDAIRMSRSRINDAFRLDDN SLEFLGIEPTLSPPYRPPVTIWLIVFGVVMGAIVVGIVLLIVSGIRNRRKNDQAGSEENP YASVDLNKGENNPGFQHADDVQTSF
6.Went to UniProt and copied the ACE-2 sequence for Mus musculus.
>sp|Q8R0I0|ACE2_MOUSE Angiotensin-converting enzyme 2 OS=Mus musculus OX=10090 GN=Ace2 PE=1 SV=1 MSSSSWLLLSLVAVTTAQSLTEENAKTFLNNFNQEAEDLSYQSSLASWNYNTNITEENAQ KMSEAAAKWSAFYEEQSKTAQSFSLQEIQTPIIKRQLQALQQSGSSALSADKNKQLNTIL NTMSTIYSTGKVCNPKNPQECLLLEPGLDEIMATSTDYNSRLWAWEGWRAEVGKQLRPLY EEYVVLKNEMARANNYNDYGDYWRGDYEAEGADGYNYNRNQLIEDVERTFAEIKPLYEHL HAYVRRKLMDTYPSYISPTGCLPAHLLGDMWGRFWTNLYPLTVPFAQKPNIDVTDAMMNQ GWDAERIFQEAEKFFVSVGLPHMTQGFWANSMLTEPADGRKVVCHPTAWDLGHGDFRIKM CTKVTMDNFLTAHHEMGHIQYDMAYARQPFLLRNGANEGFHEAVGEIMSLSAATPKHLKS IGLLPSDFQEDSETEINFLLKQALTIVGTLPFTYMLEKWRWMVFRGEIPKEQWMKKWWEM KREIVGVVEPLPHDETYCDPASLFHVSNDYSFIRYYTRTIYQFQFQEALCQAAKYNGSLH KCDISNSTEAGQKLLKMLSLGNSEPWTKALENVVGARNMDVKPLLNYFQPLFDWLKEQNR NSFVGWNTEWSPYADQSIKVRISLKSALGANAYEWTNNEMFLFRSSVAYAMRKYFSIIKN QTVPFLEEDVRVSDLKPRVSFYFFVTSPQNVSDVIPRSEVEDAIRMSRGRINDVFGLNDN SLEFLGIHPTLEPPYQPPVTIWLIIFGVVMALVVVGIIILIVTGIKGRKKKNETKREENP YDSMDIGKGESNAGFQNSDDAQTSF
7. Went to UniProt and copied the ACE-2 sequence for Rattus norvegicus.
>sp|Q5EGZ1|ACE2_RAT Angiotensin-converting enzyme 2 OS=Rattus norvegicus OX=10116 GN=Ace2 PE=1 SV=1 MSSSCWLLLSLVAVATAQSLIEEKAESFLNKFNQEAEDLSYQSSLASWNYNTNITEENAQ KMNEAAAKWSAFYEEQSKIAQNFSLQEIQNATIKRQLKALQQSGSSALSPDKNKQLNTIL NTMSTIYSTGKVCNSMNPQECFLLEPGLDEIMATSTDYNRRLWAWEGWRAEVGKQLRPLY EEYVVLKNEMARANNYEDYGDYWRGDYEAEGVEGYNYNRNQLIEDVENTFKEIKPLYEQL HAYVRTKLMEVYPSYISPTGCLPAHLLGDMWGRFWTNLYPLTTPFLQKPNIDVTDAMVNQ SWDAERIFKEAEKFFVSVGLPQMTPGFWTNSMLTEPGDDRKVVCHPTAWDLGHGDFRIKM CTKVTMDNFLTAHHEMGHIQYDMAYAKQPFLLRNGANEGFHEAVGEIMSLSAATPKHLKS IGLLPSNFQEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFQDKIPREQWTKKWWEM KREIVGVVEPLPHDETYCDPASLFHVSNDYSFIRYYTRTIYQFQFQEALCQAAKHDGPLH KCDISNSTEAGQKLLNMLSLGNSGPWTLALENVVGSRNMDVKPLLNYFQPLFVWLKEQNR NSTVGWSTDWSPYADQSIKVRISLKSALGKNAYEWTDNEMYLFRSSVAYAMREYFSREKN QTVPFGEADVWVSDLKPRVSFNFFVTSPKNVSDIIPRSEVEEAIRMSRGRINDIFGLNDN SLEFLGIYPTLKPPYEPPVTIWLIIFGVVMGTVVVGIVILIVTGIKGRKKKNETKREENP YDSMDIGKGESNAGFQNSDDAQTSF
8.Went to UniProt and copied the ACE-2 sequence for the Felis Cat.
>sp|Q56H28|ACE2_FELCA Angiotensin-converting enzyme 2 OS=Felis catus OX=9685 GN=ACE2 PE=2 SV=1 MSGSFWLLLSFAALTAAQSTTEELAKTFLEKFNHEAEELSYQSSLASWNYNTNITDENVQ KMNEAGAKWSAFYEEQSKLAKTYPLAEIHNTTVKRQLQALQQSGSSVLSADKSQRLNTIL NAMSTIYSTGKACNPNNPQECLLLEPGLDDIMENSKDYNERLWAWEGWRAEVGKQLRPLY EEYVALKNEMARANNYEDYGDYWRGDYEEEWTDGYNYSRSQLIKDVEHTFTQIKPLYQHL HAYVRAKLMDTYPSRISPTGCLPAHLLGDMWGRFWTNLYPLTVPFGQKPNIDVTDAMVNQ SWDARRIFKEAEKFFVSVGLPNMTQGFWENSMLTEPGDSRKVVCHPTAWDLGKGDFRIKM CTKVTMDDFLTAHHEMGHIQYDMAYAVQPFLLRNGANEGFHEAVGEIMSLSAATPNHLKT IGLLSPGFSEDSETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKEQWMQKWWEM KREIVGVVEPVPHDETYCDPASLFHVANDYSFIRYYTRTIYQFQFQEALCRIAKHEGPLH KCDISNSSEAGKKLLQMLTLGKSKPWTLALEHVVGEKKMNVTPLLKYFEPLFTWLKEQNR NSFVGWNTDWRPYADQSIKVRISLKSALGDEAYEWNDNEMYLFRSSVAYAMREYFSKVKN QTIPFVEDNVWVSNLKPRISFNFFVTASKNVSDVIPRSEVEEAIRMSRSRINDAFRLDDN SLEFLGIQPTLSPPYQPPVTIWLIVFGVVMGVVVVGIVLLIVSGIRNRRKNNQARSEENP YASVDLSKGENNPGFQHADDVQTSF
9.Went to UniProt and copied the ACE-2 sequence for the Horseshoe Bat (unreviewed).
>tr|E2DHI2|E2DHI2_RHIFE Angiotensin-converting enzyme OS=Rhinolophus ferrumequinum OX=59479 GN=ACE2 PE=2 SV=1 MSGSSWFLLSLVAVTAAQSTTEDLAKKFLDDFNSEAENLSHQSSLASWEYNTNISDENVQ KMDEAGAKWSDFYEKQSKLAKNFSLEEIHNDTVKLQLQILQQSGSPVLSEDKSKRLNSIL NAMSTIYSTGKVCKPNNPQECLLLEPGLDNIMGTSKDYNERLWAWEGWRAEVGKQLRPLY EEYVVLKNEMARGYHYEDYGDYWRRDYETEGSPDLEYSRDQLIKDVERIFAEIKPLYEQL HAYVRTKLMDTYPFHISPTGCLPAHLLGDMWGRFWTNLYPLTVPFGQKPNIDVTDAMLNQ NWDAKRIFKEAEKFLVSIGLPNMTEGFWNNSMLTDPGDGRKVVCHPTAWDLGKGDFRIKM CTKVTMEDFLTAHHEMGHIQYDMAYASQPYLLRNGANEGFHEAVGEVMSLSVATPEHLKT MGLLSSDFLEDNETEINFLFKQALNIVGTLPLTYMLEKWRWMVFKGEIPKEEWMKKWWEM KRKIVGVVEPVPHDETYCDPASLFHVANDYSFIRYYTRTIFEFQFHEALCRIAKHDGPLH KCGISNSTDAGEKLHQMLSVGKSQPWTSVLKDFVGSKNMDVGPLLRYFEPLYTWLTEQNR KSFVGWNTDWSPYADQSIKVWISLKSALGEKAYEWNNNEMYLFRSSVAYAMREYFLKTKN QTILFGEEDVWVSNLKPRISFNFYVTSPRNLSDIIPRPEVEGAIRMSRSRINDAFRLDDN SLEFLGIQPTLGPPYQPPVTIWLIVFGVVMAVVVVGIVVLIITGIRDRRKKDQARSEENP YSSVDLSKGENNPGFQNGNDVQTSF
10.Went to NCBI and copied the ACE-2 sequence for the Rabbit.
>QHX39726.1 angiotensin I converting enzyme 2 [Oryctolagus cuniculus] MSGSSWLLLSLVAVTAAQSTIEELAKTFLEKFNQEAEDLSYQSALASWDYNTNITEENVQKMNDAEAKWS AFYEEQSKLAKTYPSQEVQNLTVKRQLQALQQSGSSALSADKSKQLNTILSTMSTIYSTGKVCNQSNPQE CFLLEPGLDEIMAKSTDYNERLWAWEGWRSVVGKQLRPLYEEYVVLKNEMARANNYEDYGDYWRADYEAE GADGYDYSRSQLIDDVERTFSEIKPLYEQLHAYVRTKLMDAYPSRISPTGCLPAHLLGDMWGRFWTNLYS LTVPFGQKPNIDVTDTMVNQGWDAERIFKEAEKFFVSVGLPSMTHGFWENSMLPESGDGRKVVCHPTAWD LGKRDFRIKMCTKVTMDNFLTAHHEMGHIQYDMAYATQPFLLRNGANEGFHEAVGEIMSLSAATPEHLKS IGLLPYDFHEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKEQWMQKWWEMKREIVGVVEP MPHDETYCDPAALFHVANDYSFIRYYTRTIYQFQFQEALCQAAQHEGPLHKCDISNSTEAGQKLLNMLRL GRSEPWTLALENVVGAKNMDVRPLLNYFEPLFTWLKEQNRNSFVGWSTEWTPYADQSIKVRISLKTALGD QAYEWNDSEMYLFRSSVAYAMRKYFSEVKNQTILFGEEDVRVSDLKPRISFNFFVTAPNNVNDIIPRNEV EEAISMSRSRINDIFRLDDNSLEFVGIQPTLEPPYESPVPIWLVVFGVVMGMIVIGIVVLIFTGIKDRRK QKQAKREENPYGFVDMSKGENNSGFQNSDDIQTSF
11. Went to UniProt and copied the ACE-2 sequence for the opossum.
>tr|F6WXR7|F6WXR7_MONDO Angiotensin-converting enzyme OS=Monodelphis domestica OX=13616 GN=ACE2 PE=3 SV=2 MLDPLWLFFSLLAVTAAQNSIEEDAKTFLDDYNAKAEELSHQSALASWEYNTNITNENVE KMNEAAARWSSFYENQSSISRTYPLNEITNATVKLQLKSLQKKEGAVLSTEQSVRLNTIL NTMSTLYSTGSVCNSETPQQCFLLEPGLDKIMDESTDYDERLWAWEGWRSKVGKEMRPLY EEYVELKNELAKGNNYEDYGDYWRGDYEVEEPSEYVYSRPQLKKDVENTFKQIKSLYEHL HAYVRRKMRNTYGSLISETGGLPAHLLGDMWGRFWTNLYSLTMPYREKPNIDVTSAMKKQ NWSARRIFQEAEMFFASVGLPNMTEGFWKNSMLTEPNDGRKVVCHPTAWDLGKNDFRIKM CTKVTMDDFLTAHHEMGHIQYDMAYAKQPFTLRNGANEGFHEAVGEIMSLSAATPKHLQA LGLLPPTFQEDNETEINFLFKQALTIIGTMPFTYMLENWRWMVFEGKIPKEEWMKKWWEM KREIVGVVEPLPHDETYCDPAALFHVANDYSFIRYYTRTIYQFQFHKALCKIAQPSAALH KCDITNSTEAGTKLQNMLKMGKSEPWTKALESIVGNKMMDAGPLLEYFEPLFTWLKEQNK DAYVGWNTDWSPYNAYKIKVRISLKTLGENAYTWNENEMYLFQSSIVFAMRQYFLIKKKQ SIPFSNENVKMFDLKPRISFYFFVTFPPNGTSFVPREEVEAAISMSRDRINDAFRLNDNS LEFVGISPTLAPPYEPPVTVWMIVFGVVMGIVVIGIVYLIYTGVRDRKKRAKTSSSNDEN PYVDVDVAGGQHNPAFQSSEDAQTSF
2.Went to a phylogeny tree builder and sequence aligner.
- Copied the ACE-2 sequence and pasted them in the one-click.
- Copied the generated sequence alignment and pasted the results below.
Sequence alignment
CLUSTAL FORMAT: MUSCLE (3.8) multiple sequence alignments
tr|E2DHI2| MSGSSWFLLSLVAVTAAQSTTEDLAKKFLDDFNSEAENLSHQSSLASWEYNTNISDENVQ
sp|Q8R0I0| MSSSSWLLLSLVAVTTAQSLTEENAKTFLNNFNQEAEDLSYQSSLASWNYNTNITEENAQ
sp|Q5EGZ1| MSSSCWLLLSLVAVATAQSLIEEKAESFLNKFNQEAEDLSYQSSLASWNYNTNITEENAQ
tr|A0A220Q MSGSFWLLLSLIPVTAAQSTTEELAKTFLEKFNLEAEDLAYQSSLASWNYNTNITDENIQ
QHX39726.1 MSGSSWLLLSLVAVTAAQSTIEELAKTFLEKFNQEAEDLSYQSALASWDYNTNITEENVQ
tr|A0A2J8K MSGSSWLLLSLVAVTAAQSTIEEQAKTFLDKFNHEAEDLFYQSSLASWNYNTNITEENVQ
sp|Q9BYF1| MSSSSWLLLSLVAVTAAQSTIEEQAKTFLDKFNHEAEDLFYQSSLASWNYNTNITEENVQ
BAE53380.1 MLGSSWLLLSLAALTAAQSTTEDLAKTFLEKFNYEAEELSYQNSLASWNYNTNITDENIQ
sp|Q56NL1| MSGSFWLLLSFAALTAAQSTTEELAKTFLETFNYEAQELSYQSSVASWNYNTNITDENAK
sp|Q56H28| MSGSFWLLLSFAALTAAQSTTEELAKTFLEKFNHEAEELSYQSSLASWNYNTNITDENVQ
* .* *:***: .:::*** *: *:.**: ** **::* :*.::***:*****::** :
tr|E2DHI2| KMDEAGAKWSDFYEKQSKLAKNFSLEEIHNDTVKLQLQILQQSGSPVLSEDKSKRLNSIL
sp|Q8R0I0| KMSEAAAKWSAFYEEQSKTAQSFSLQEIQTPIIKRQLQALQQSGSSALSADKNKQLNTIL
sp|Q5EGZ1| KMNEAAAKWSAFYEEQSKIAQNFSLQEIQNATIKRQLKALQQSGSSALSPDKNKQLNTIL
tr|A0A220Q KMNDARAKWSAFYEEQSRIAKTYPLDEIQTLILKRQLQALQQSGTSGLSADKSKRLNTIL
QHX39726.1 KMNDAEAKWSAFYEEQSKLAKTYPSQEVQNLTVKRQLQALQQSGSSALSADKSKQLNTIL
tr|A0A2J8K NMNNAGDKWSAFLKEQSTLAQMYPLQEIQNLTVKLQLQALQQNGSSVLSEDKSKRLNTIL
sp|Q9BYF1| NMNNAGDKWSAFLKEQSTLAQMYPLQEIQNLTVKLQLQALQQNGSSVLSEDKSKRLNTIL
BAE53380.1 KMNIAGAKWSAFYEEESQHAKTYPLEEIQDPIIKRQLRALQQSGSSVLSADKRERLNTIL
sp|Q56NL1| NMNEAGAKWSAYYEEQSKLAQTYPLAEIQDAKIKRQLQALQQSGSSVLSADKSQRLNTIL
sp|Q56H28| KMNEAGAKWSAFYEEQSKLAKTYPLAEIHNTTVKRQLQALQQSGSSVLSADKSQRLNTIL
:*. * *** : :::* *: :. *:: :* **. ***.*:. ** ** :.**:**
tr|E2DHI2| NAMSTIYSTGKVCKPNNPQECLLLEPGLDNIMGTSKDYNERLWAWEGWRAEVGKQLRPLY
sp|Q8R0I0| NTMSTIYSTGKVCNPKNPQECLLLEPGLDEIMATSTDYNSRLWAWEGWRAEVGKQLRPLY
sp|Q5EGZ1| NTMSTIYSTGKVCNSMNPQECFLLEPGLDEIMATSTDYNRRLWAWEGWRAEVGKQLRPLY
tr|A0A220Q NTMSTIYSSGKVLDPNNPQECLVLEPGLDEIMENSKDYSRRLWAWESWRAEVGKQLRPLY
QHX39726.1 STMSTIYSTGKVCNQSNPQECFLLEPGLDEIMAKSTDYNERLWAWEGWRSVVGKQLRPLY
tr|A0A2J8K NTMSAIYSTGKVCNPNNPQECLLLEPGLNEIMANSLDYNERLWAWESWRSEVGKQLRPLY
sp|Q9BYF1| NTMSTIYSTGKVCNPDNPQECLLLEPGLNEIMANSLDYNERLWAWESWRSEVGKQLRPLY
BAE53380.1 NAMSTIYSTGKACNPNNPQECLLLEPGLDDIMENSKDYNERLWAWEGWRSEVGKQLRPLY
sp|Q56NL1| NAMSTIYSTGKACNPNNPQECLLLEPGLDNIMENSKDYNERLWAWEGWRAEVGKQLRPLY
sp|Q56H28| NAMSTIYSTGKACNPNNPQECLLLEPGLDDIMENSKDYNERLWAWEGWRAEVGKQLRPLY
.:**:***:**. . *****::*****::** .* **. ******.**: *********
tr|E2DHI2| EEYVVLKNEMARGYHYEDYGDYWRRDYETEGSPDLEYSRDQLIKDVERIFAEIKPLYEQL
sp|Q8R0I0| EEYVVLKNEMARANNYNDYGDYWRGDYEAEGADGYNYNRNQLIEDVERTFAEIKPLYEHL
sp|Q5EGZ1| EEYVVLKNEMARANNYEDYGDYWRGDYEAEGVEGYNYNRNQLIEDVENTFKEIKPLYEQL
tr|A0A220Q EEYVVLENEMARANNYEDYGDYWRGDYEVTGTGDYDYSRNQLMEDVERTFAEIKPLYEHL
QHX39726.1 EEYVVLKNEMARANNYEDYGDYWRADYEAEGADGYDYSRSQLIDDVERTFSEIKPLYEQL
tr|A0A2J8K EEYVVLKNEMARANHYEDYGDYWRGDYEVNGVDGYDYSRGQLIEDVEHTFEEIKPLYEHL
sp|Q9BYF1| EEYVVLKNEMARANHYEDYGDYWRGDYEVNGVDGYDYSRGQLIEDVEHTFEEIKPLYEHL
BAE53380.1 EEYVALKNEMARANNYEDYGDYWRGDYEEEWADGYSYSRNQLIEDVEHTFTQIKPLYEHL
sp|Q56NL1| EEYVALKNEMARANNYEDYGDYWRGDYEEEWTGGYNYSRNQLIQDVEDTFEQIKPLYQHL
sp|Q56H28| EEYVALKNEMARANNYEDYGDYWRGDYEEEWTDGYNYSRSQLIKDVEHTFTQIKPLYQHL
****.*:*****. :*:******* *** . .*.*.**:.*** * :*****::*
tr|E2DHI2| HAYVRTKLMDTYPFHISPTGCLPAHLLGDMWGRFWTNLYPLTVPFGQKPNIDVTDAMLNQ
sp|Q8R0I0| HAYVRRKLMDTYPSYISPTGCLPAHLLGDMWGRFWTNLYPLTVPFAQKPNIDVTDAMMNQ
sp|Q5EGZ1| HAYVRTKLMEVYPSYISPTGCLPAHLLGDMWGRFWTNLYPLTTPFLQKPNIDVTDAMVNQ
tr|A0A220Q HAYVRAKLMDAYPSRISPTGCLPAHLLGDMWGRFWTNLYPLTVPFGEKPSIDVTEAMVNQ
QHX39726.1 HAYVRTKLMDAYPSRISPTGCLPAHLLGDMWGRFWTNLYSLTVPFGQKPNIDVTDTMVNQ
tr|A0A2J8K HAYVRAKLMNAYPSYISPIGCLPAHLLGDMWGRFWTNLYSLTVPFGQKPNIDVTDAMVDQ
sp|Q9BYF1| HAYVRAKLMNAYPSYISPIGCLPAHLLGDMWGRFWTNLYSLTVPFGQKPNIDVTDAMVDQ
BAE53380.1 HAYVRAKLMDAYPSRISPTGCLPAHLLGDMWGRFWTNLYPLMVPFRQKPNIDVTDAMVNQ
sp|Q56NL1| HAYVRAKLMDTYPSRISRTGCLPAHLLGDMWGRFWTNLYPLTVPFGQKPNIDVTDAMVNQ
sp|Q56H28| HAYVRAKLMDTYPSRISPTGCLPAHLLGDMWGRFWTNLYPLTVPFGQKPNIDVTDAMVNQ
***** ***:.** ** ********************.* .** :**.****::*::*
tr|E2DHI2| NWDAKRIFKEAEKFLVSIGLPNMTEGFWNNSMLTDPGDGRKVVCHPTAWDLGKGDFRIKM
sp|Q8R0I0| GWDAERIFQEAEKFFVSVGLPHMTQGFWANSMLTEPADGRKVVCHPTAWDLGHGDFRIKM
sp|Q5EGZ1| SWDAERIFKEAEKFFVSVGLPQMTPGFWTNSMLTEPGDDRKVVCHPTAWDLGHGDFRIKM
tr|A0A220Q SWDAIRIFEEAEKFFVSIGLPNMTQGFWNNSMLTEPGDGRKVVCHPTAWDLGKGDFRIKM
QHX39726.1 GWDAERIFKEAEKFFVSVGLPSMTHGFWENSMLPESGDGRKVVCHPTAWDLGKRDFRIKM
tr|A0A2J8K AWDAQRIFKEAEKFFVSVGLPNMTQGFWENSMLTDPGNVQKAVCHPTAWDLGKGDFRILM
sp|Q9BYF1| AWDAQRIFKEAEKFFVSVGLPNMTQGFWENSMLTDPGNVQKAVCHPTAWDLGKGDFRILM
BAE53380.1 SWDARRIFEEAETFFVSVGLPNMTEGFWQNSMLTEPGDNRKVVCHPTAWDLGKRDFRIKM
sp|Q56NL1| NWDARRIFKEAEKFFVSVGLPNMTQGFWENSMLTEPGDGRKVVCHPTAWDLGKGDFRIKM
sp|Q56H28| SWDARRIFKEAEKFFVSVGLPNMTQGFWENSMLTEPGDSRKVVCHPTAWDLGKGDFRIKM
*** ***:***.*:**:*** ** *** ****.:..: .*.**********: **** *
tr|E2DHI2| CTKVTMEDFLTAHHEMGHIQYDMAYASQPYLLRNGANEGFHEAVGEVMSLSVATPEHLKT
sp|Q8R0I0| CTKVTMDNFLTAHHEMGHIQYDMAYARQPFLLRNGANEGFHEAVGEIMSLSAATPKHLKS
sp|Q5EGZ1| CTKVTMDNFLTAHHEMGHIQYDMAYAKQPFLLRNGANEGFHEAVGEIMSLSAATPKHLKS
tr|A0A220Q CTKVTMDDFLTAHHEMGHIQYDMAYAIQPYLLRNGANEGFHEAVGEIMSLSAATPHYLKA
QHX39726.1 CTKVTMDNFLTAHHEMGHIQYDMAYATQPFLLRNGANEGFHEAVGEIMSLSAATPEHLKS
tr|A0A2J8K CTKVTMDDFLTAHHEMGHIQYDMAYAAQPFLLRNGANEGFHEAVGEIMSLSAATPKHLKS
sp|Q9BYF1| CTKVTMDDFLTAHHEMGHIQYDMAYAAQPFLLRNGANEGFHEAVGEIMSLSAATPKHLKS
BAE53380.1 CTKVTMDDFLTAHHEMGHIQYDMAYAEQPFLLRNGANEGFHEAVGEIMSLSAATPNHLKN
sp|Q56NL1| CTKVTMDDFLTAHHEMGHIQYDMAYAAQPFLLRNGANEGFHEAVGEIMSLSAATPNHLKT
sp|Q56H28| CTKVTMDDFLTAHHEMGHIQYDMAYAVQPFLLRNGANEGFHEAVGEIMSLSAATPNHLKT
******::****************** **:****************:****.*** :**
tr|E2DHI2| MGLLSSDFLEDNETEINFLFKQALNIVGTLPLTYMLEKWRWMVFKGEIPKEEWMKKWWEM
sp|Q8R0I0| IGLLPSDFQEDSETEINFLLKQALTIVGTLPFTYMLEKWRWMVFRGEIPKEQWMKKWWEM
sp|Q5EGZ1| IGLLPSNFQEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFQDKIPREQWTKKWWEM
tr|A0A220Q LGLLPPDFYEDSETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKEQWMQKWWEM
QHX39726.1 IGLLPYDFHEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKEQWMQKWWEM
tr|A0A2J8K IGLLSPDFQEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPEDQWMKKWWEM
sp|Q9BYF1| IGLLSPDFQEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKDQWMKKWWEM
BAE53380.1 IGLLPPDFSEDSETDINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKEQWMQKWWEM
sp|Q56NL1| IGLLSPAFSEDNETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGAIPKEQWMQKWWEM
sp|Q56H28| IGLLSPGFSEDSETEINFLLKQALTIVGTLPFTYMLEKWRWMVFKGEIPKEQWMQKWWEM
:***. * **.**:****:****.******:************.. ** ::* :*****
tr|E2DHI2| KRKIVGVVEPVPHDETYCDPASLFHVANDYSFIRYYTRTIFEFQFHEALCRIAKHDGPLH
sp|Q8R0I0| KREIVGVVEPLPHDETYCDPASLFHVSNDYSFIRYYTRTIYQFQFQEALCQAAKYNGSLH
sp|Q5EGZ1| KREIVGVVEPLPHDETYCDPASLFHVSNDYSFIRYYTRTIYQFQFQEALCQAAKHDGPLH
tr|A0A220Q KREIVGVVEPLPHDETYCDPACLFHVAEDYSFIRYYTRTIYQFQFHEALCRTAKHEGPLY
QHX39726.1 KREIVGVVEPMPHDETYCDPAALFHVANDYSFIRYYTRTIYQFQFQEALCQAAQHEGPLH
tr|A0A2J8K KREIVGVVEPVPHDETYCDPASLFHVSNDYSFIRYYTRTLYQFQFQEALCQAAKHEGPLH
sp|Q9BYF1| KREIVGVVEPVPHDETYCDPASLFHVSNDYSFIRYYTRTLYQFQFQEALCQAAKHEGPLH
BAE53380.1 KRDIVGVVEPLPHDETYCDPAALFHVANDYSFIRYYTRTIYQFQFQEALCQIAKHEGPLY
sp|Q56NL1| KRNIVGVVEPVPHDETYCDPASLFHVANDYSFIRYYTRTIYQFQFQEALCQIAKHEGPLH
sp|Q56H28| KREIVGVVEPVPHDETYCDPASLFHVANDYSFIRYYTRTIYQFQFQEALCRIAKHEGPLH
**.*******:**********.****::***********:::***:****. *:::*.*:
tr|E2DHI2| KCGISNSTDAGEKLHQMLSVGKSQPWTSVLKDFVGSKNMDVGPLLRYFEPLYTWLTEQNR
sp|Q8R0I0| KCDISNSTEAGQKLLKMLSLGNSEPWTKALENVVGARNMDVKPLLNYFQPLFDWLKEQNR
sp|Q5EGZ1| KCDISNSTEAGQKLLNMLSLGNSGPWTLALENVVGSRNMDVKPLLNYFQPLFVWLKEQNR
tr|A0A220Q KCDISNSTEAGQKLLQMLSLGKSEPWTLALENIVGVKTMDVKPLLSYFEPLLTWLKAQNG
QHX39726.1 KCDISNSTEAGQKLLNMLRLGRSEPWTLALENVVGAKNMDVRPLLNYFEPLFTWLKEQNR
tr|A0A2J8K KCDISNSTEAGQKLFNMLRLGKSEPWTLALENVVGAKNMNVRPLLNYFEPLFTWLKDQNK
sp|Q9BYF1| KCDISNSTEAGQKLFNMLRLGKSEPWTLALENVVGAKNMNVRPLLNYFEPLFTWLKDQNK
BAE53380.1 KCDISNSSEAGQKLHEMLSLGRSKPWTFALERVVGAKTMDVRPLLNYFEPLFTWLKEQNR
sp|Q56NL1| KCDISNSTEAGKKLLEMLSLGRSEPWTLALERVVGAKNMNVTPLLNYFEPLFTWLKEQNR
sp|Q56H28| KCDISNSSEAGKKLLQMLTLGKSKPWTLALEHVVGEKKMNVTPLLKYFEPLFTWLKEQNR
**.****::**:** :** :*.* *** .*: .** ..*:* *** **:** **. **
tr|E2DHI2| KSFVGWNTDWSPYADQSIKVWISLKSALGEKAYEWNNNEMYLFRSSVAYAMREYFLKTKN
sp|Q8R0I0| NSFVGWNTEWSPYADQSIKVRISLKSALGANAYEWTNNEMFLFRSSVAYAMRKYFSIIKN
sp|Q5EGZ1| NSTVGWSTDWSPYADQSIKVRISLKSALGKNAYEWTDNEMYLFRSSVAYAMREYFSREKN
tr|A0A220Q NSSVGWNTDWTPYADQSIKVRISLKSALGKEAYEWNDNEMYLFRSSIAYAMRNYFSSAKN
QHX39726.1 NSFVGWSTEWTPYADQSIKVRISLKTALGDQAYEWNDSEMYLFRSSVAYAMRKYFSEVKN
tr|A0A2J8K NSFVGWSTDWSPYADQSIKVRISLKSALGDKAYEWNDNEMYLFRSSVAYAMRQYFLKVKN
sp|Q9BYF1| NSFVGWSTDWSPYADQSIKVRISLKSALGDKAYEWNDNEMYLFRSSVAYAMRQYFLKVKN
BAE53380.1 NSFVGWNTDWSPYADQSIKVRISLKSALGEKAYEWNDNEMYFFQSSIAYAMREYFSKVKN
sp|Q56NL1| NSFVGWDTDWRPYSDQSIKVRISLKSALGEKAYEWNDNEMYLFRSSIAYAMREYFSKVKN
sp|Q56H28| NSFVGWNTDWRPYADQSIKVRISLKSALGDEAYEWNDNEMYLFRSSVAYAMREYFSKVKN
:* ***.*:* **:******.****:*** :****.:.**::*.**:*****:** **
tr|E2DHI2| QTILFGEEDVWVSNLKPRISFNFYVTSPRNLSDIIPRPEVEGAIRMSRSRINDAFRLDDN
sp|Q8R0I0| QTVPFLEEDVRVSDLKPRVSFYFFVTSPQNVSDVIPRSEVEDAIRMSRGRINDVFGLNDN
sp|Q5EGZ1| QTVPFGEADVWVSDLKPRVSFNFFVTSPKNVSDIIPRSEVEEAIRMSRGRINDIFGLNDN
tr|A0A220Q ETIPFGAEDVWVSDLKPRISFNFFVTSPANMSDIIPRSDVEKAISMSRSRINDAFRLDDN
QHX39726.1 QTILFGEEDVRVSDLKPRISFNFFVTAPNNVNDIIPRNEVEEAISMSRSRINDIFRLDDN
tr|A0A2J8K QMILFGEEDVRVANLKPRISFNFFVTAPKNVSDIIPRTEVEKAIRKSRSRINDAFRLNDN
sp|Q9BYF1| QMILFGEEDVRVANLKPRISFNFFVTAPKNVSDIIPRTEVEKAIRMSRSRINDAFRLNDN
BAE53380.1 QTIPFVGKDVRVSDLKPRISFNFIVTSPENMSDIIPRADVEEAIRKSRGRINDAFRLDDN
sp|Q56NL1| QTIPFVEDNVWVSDLKPRISFNFFVTFSNNVSDVIPRSEVEDAIRMSRSRINDAFRLDDN
sp|Q56H28| QTIPFVEDNVWVSNLKPRISFNFFVTASKNVSDVIPRSEVEEAIRMSRSRINDAFRLDDN
: : * :*.*::****:** * ** . *:.*:*** :** ** **.**** * *:**
tr|E2DHI2| SLEFLGIQPTLGPPYQPPVTIWLIVFGVVMAVVVVGIVVLIITGIRDRRKKDQARSEENP
sp|Q8R0I0| SLEFLGIHPTLEPPYQPPVTIWLIIFGVVMALVVVGIIILIVTGIKGRKKKNETKREENP
sp|Q5EGZ1| SLEFLGIYPTLKPPYEPPVTIWLIIFGVVMGTVVVGIVILIVTGIKGRKKKNETKREENP
tr|A0A220Q TLEFLGIQPTLGPPDEPPVTVWLIIFGVVMGLVVVGIVVLIFTGIRDRRKKKQASSEENP
QHX39726.1 SLEFVGIQPTLEPPYESPVPIWLVVFGVVMGMIVIGIVVLIFTGIKDRRKQKQAKREENP
tr|A0A2J8K SLEFLGIQPTLGPPNQPPVSIWLIVFGVVMGVIVVGIVILIFTGIRDRKKKNKARSEENP
sp|Q9BYF1| SLEFLGIQPTLGPPNQPPVSIWLIVFGVVMGVIVVGIVILIFTGIRDRKKKNKARSGENP
BAE53380.1 SLEFLGIQPTLEPPYQPPVTIWLIVFGVVMGVVVVGIFLLIFSGIRNRRKNNQARSEENP
sp|Q56NL1| SLEFLGIEPTLSPPYRPPVTIWLIVFGVVMGAIVVGIVLLIVSGIRNRRKNDQAGSEENP
sp|Q56H28| SLEFLGIQPTLSPPYQPPVTIWLIVFGVVMGVVVVGIVLLIVSGIRNRRKNNQARSEENP
:***:** *** ** .**.:**::*****. :*:**.:**.:**..*.*:.:: ***
tr|E2DHI2| YSSVDLSKGENNPGFQNGNDVQTSF
sp|Q8R0I0| YDSMDIGKGESNAGFQNSDDAQTSF
sp|Q5EGZ1| YDSMDIGKGESNAGFQNSDDAQTSF
tr|A0A220Q YGSMDLSKGESNSGFQNGDDIQTSF
QHX39726.1 YGFVDMSKGENNSGFQNSDDIQTSF
tr|A0A2J8K YASVDTSKGENNPGFQNTDDVQTSF
sp|Q9BYF1| YASIDISKGENNPGFQNTDDVQTSF
BAE53380.1 YASVDLSKGENNPGFQNVDDVQTSF
sp|Q56NL1| YASVDLNKGENNPGFQHADDVQTSF
sp|Q56H28| YASVDLSKGENNPGFQHADDVQTSF
* :* .***.*.***: :* ****
3.Went to a phylogeny tree builder and sequence aligner.
- Copied the ACE-2 sequence and pasted them in the one-click.
- Copied the generated Phylogenetic tree and pasted the results below.

Figure 1: Phylogenetic tree results for the copied ACE-2 Sequences.
4. Went to the article titled Investigation of the genetic variation in ACE2 on the structural recognition by the novel coronavirus (SARS-CoV-2) and copied the 8 necessary human amino acids and pasted them below.
- HIS378
- SER19
- GLY211
- ASP 206
- ARG 219
- LYS 341
- LLE 468
- SER 547
5. Located the 8 necessary human ACE-2 amino acids for structure and function in the sequence alignment and highlighted them across sequences. Created a table making the sequence alignments clearer. Posted sequences and interpretations below.



Figure 2: Sequence alignment for 8 necessary human amino acids in all the sequences.
- Fully conserved residues: His378, Asp 206, Arg 219, Lys 341, LLE 468, and Ser 547.
- Strongly Conserved residue: Ser 19
- No conservation: Gly 211
6. Went to NCBI 3d-protein shapes iCn3D and created two structures.
- The first is highlighting the necessary human ACE2 amino residues.
- The second is the 5 human amino acid residues that the 2019-nCoV binds to.
- Pasted images below:
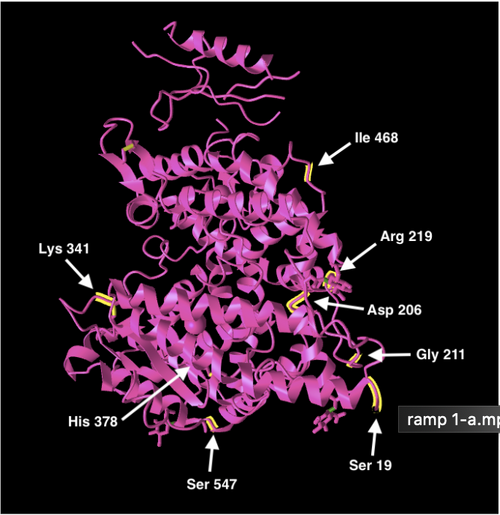
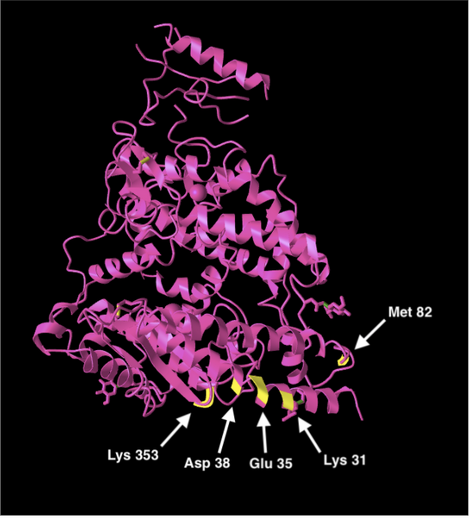
Figure 3: Protein structure showing 8 necessary amino acids for function and structure in human ACE2s (left). Protein structure showing 5 ACE2 amino acid residues that the novel 2019-nCoV binds to.
- None of the necessary amino acid residues for the human ACE2 structure and function is a residue where the 2019-nCoV spike protein binds to.
Powerpoint presentation
- Link to the PowerPoint presentation that will be presented in class.
Scientific Conclusion
- The goal of this lab is to compare the ACE2 sequences of different species, note fully conserved amino acids necessary for structure and function, and compare them to the 5 amino acid sequences that the novel 2019-nCoV spike protein binds to. This work finds through a phylogenetic tree and sequence alignment that the ACE-2 amino acid residues that are more compatible with the novel 2019-nCoV do not have to be more closely related in the tree. The 8 amino acid residues that are necessary for human ACE2 structure and function are not the same as the 5 residues that the 2019-nCoV spike protein binds to. The sequence alignment shows that across the 11 species tested only 6 of the 8 necessary amino acids were fully conserved, 1 were strongly conserved, and 1 was not conserved. Mutations, therefore, to the 5 amino acid sequences will not influence the ACE-2 structure or function, in humans. Future experiments will focus on mutations to the 5 key amino acid sequences for the 2019-nCoV spike protein attempt to lower the binding efficiency for the spike protein.
Acknowledgments
- Referenced and copied OpenWebWare syntax from the BIOL368/F20 week 1 page.
- Referenced and copied methods from the BIOL368/F20 week 6 page.
- Referenced MediaWiki for image formatting syntax.
- Worked with my partner Macie Duran on the presentation and research.
- Worked with Dr. Dahlquist, in class, on our research and protein databases.
- Referenced the article "Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus" for information regarding 3-dimensional proteins.
- Referenced the article Investigation of the genetic variation in ACE2 on the structural recognition by the novel coronavirus (SARS-CoV-2) for the 8 necessary human ACE2 amino acid residues.
- Used UniProtto find ACE-2 protein sequences.
- Used NCBI to find ACE-2 Protein sequences and study protein structures.
- Used Phylogeny.fr to make a phylogenetic tree and sequence alignment.
References
- OpenWetWare. (2020). BIOL368/F20:Week 1. Retrieved September 22, 2020, from https://openwetware.org/wiki/BIOL368/F20:Week_1
- OpenWetWare. (2020). BIOL368/F20:Week 6. Retrieved October 14, 2020, from https://openwetware.org/wiki/BIOL368/F20:Week_6
- Wan, Y., et al. (2020). Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. Journal of Virology, 54 (7), retrieved from https://doi.org/10.1128/JVI.00127-20.
- Phylogeny.fr (2020). Methodes et Algorithmes pour la Bio-informatique LIRM, retrieved from http://www.phylogeny.fr/simple_phylogeny.cgi?workflow_id=f6c1c87f6ac1d379903543866e4da087&tab_index=3%7C.
- NCBI Structure (2020). PDB ID 1R42: Native Human Angiotensin-Converting Enzyme-2, Retrieved from https://www.ncbi.nlm.nih.gov/Structure/icn3d/full.html?&mmdbid=26160&bu=1&showanno=1&source=full-feature.
- NCBI Database (2020). Angiotensin I converting enzyme 2 [Mustela putorius furo], https://www.ncbi.nlm.nih.gov/protein/BAE53380.1?report=fasta%7C.
- NCBI Database (2020). Angiotensin-converting enzyme 2 [Oryctolagus cuniculus], https://www.ncbi.nlm.nih.gov/protein/XP_002719891.1.
- UniProt (2020). UniProtKB - K7GLM4 (K7GLM4_PIG), retrieved from https://www.uniprot.org/uniprot/K7GLM4.
- UniProt (2020). UniProtKB - Q9BYF1 (ACE2_HUMAN), retrieved from https://www.uniprot.org/uniprot/Q56NL1.
- UniProt (2020). UniProtKB - Q5EGZ1 (ACE2_RAT), retrieved from https://www.uniprot.org/uniprot/Q5EGZ1.
- UniProt (2020). UniProtKB - Q56NL1 (ACE2_PAGLA), retrieved from https://www.uniprot.org/uniprot/Q56NL1.
- UniProt (2020). UniProtKB - A0A2J8KU96 (A0A2J8KU96_PANTR), retrieved fromhttps://www.uniprot.org/uniprot/A0A2J8KU96.
- UniProt (2020). UniProtKB - B6ZGN7 (B6ZGN7_RHIFE), retrieved from https://www.uniprot.org/uniprot/B6ZGN7.
- UniProt (2020). UniProtKB - Q8R0I0 (ACE2_MOUSE), retrieved from hhttps://www.uniprot.org/uniprot/Q8R0I0.
- UniProt (2020). UniProtKB - Q56H28 (ACE2_FELCA), retrieved from https://www.uniprot.org/uniprot/Q56H28.
- UniProt (2020). UniProtKB - F6WXR7 (F6WXR7_MONDO), retrieved from https://www.uniprot.org/uniprot/F6WXR7.
- Guo, X., Chen, Z., Xia, Y. et al. Investigation of the genetic variation in ACE2 on the structural recognition by the novel coronavirus (SARS-CoV-2). J Transl Med 18, 321 (2020). https://doi.org/10.1186/s12967-020-02486-7
"Except for what is noted above, this individual journal entry was completed by me and not copied from another source" Nathan R. Beshai (talk) 20:36, 14 October 2020 (PDT)