Moneil5 Week 7 Re-visited
Helpful Links
Assignment Pages:
- Week 1 Assignment
- Week 2 Assignment
- Week 3 Assignment
- Week 4 Assignment
- Week 5 Assignment
- Week 6 Assignment
- Week 7 Assignment
- Week 9 Assignment
- Week 10 Assignment
- Week 11 Assignment
- Week 12 Assignment
- Week 14/15 Assignment
Personal Journal Entries:
- Week 1
- Week 2
- Week 3
- Week 4
- Week 5
- Week 6
- Week 7
- Week 7 Re-visited
- Week 9
- Week 10
- Week 11
- Week 12
- Week 14/15
Shared Journal Entries:
Purpose
The purpose of this assignment is to re-visit the adapted nitrogen consumption system of differential equations and see if the model can be improved to better model the chemostat from the ter Schure et al. paper.
Workflow
- Based on Dr. Fitzpatrick's notes and looking at the equations again I decided to remove the glucose consumption variable from my system of equations as it is redundant given the last term in the dc2/dt equation.
- Based on the above, this renders the m variable as also not needing to be used
- Rest of equation workflow handwritten in order to develop equations. can be found here
- Was in the middle of running the model and then the power conveniently went out
- Re-wrote MATLAB code based on changes made in do over:
function dx = Week8Dynamics(t,x) global q u1 u2 V1 K1 K2 dx = zeros(size(x)); dx(1) = (q*u1) - (q*x(1)) - ((x(3)*V1)*(x(1)/(x(1)+K1))*(x(2)/(x(2)+K2))); dx(2) = (q*u2) - (q*x(1)) - ((x(3)*(((q*u2)-(q*x(2))+ (q*x(4)))*x(3)))*(x(1)/(x(1)+K1))*(x(2)/(x(2)+K2))); dx(3) = ((x(3)*(((-q*u1)-(q*u2)+(q*x(4)))*x(3)))*(x(1)/(x(1)+K1))*(x(2)/(x(2)+K2)))- (q*x(3)); dx(4) = ((x(3)*x(4)*(((q*u2)-(q*x(2))+ (q*x(4)))*x(3)))*(x(1)/(x(1)+K1))*(x(2)/(x(2)+K2)))- (q*x(4)); end
- And re-wrote script to be:
global q u1 u2 V1 K1 K2
q = 0.15;
u1 =70;
u2 = 100;
V1 = 110;
K1 = 10;
K2 = 50;
tt = 0:0.1:10;
x0= [1,1,1,1];
[t,x] = ode45('Week8Dynamics',tt,x0);
plot(t,x)
xlabel('time')
ylabel('abundance')
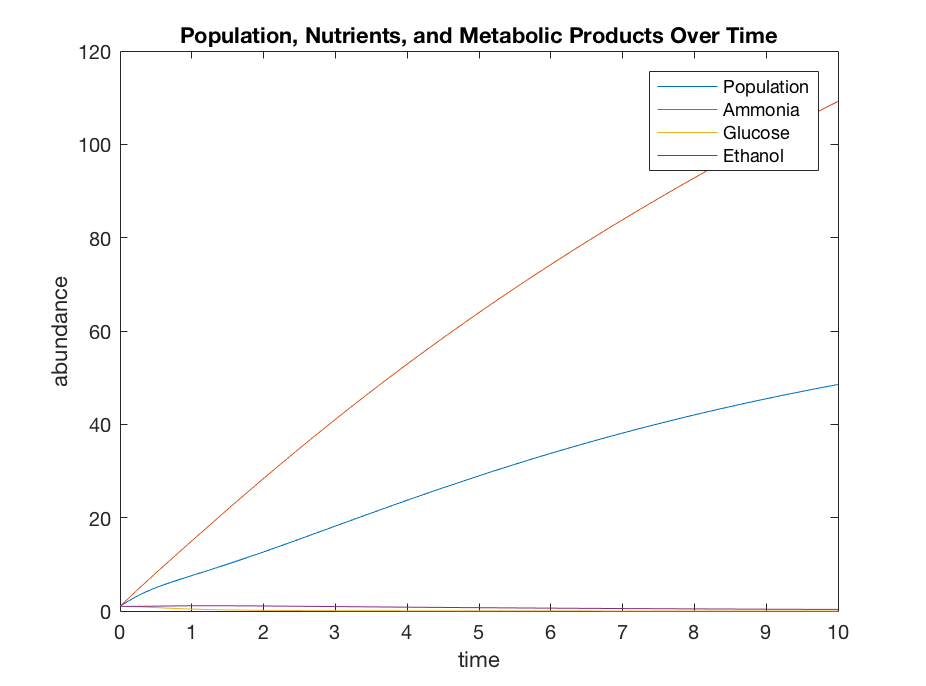
title('Population, Nutrients, and Metabolic Products Over Time')
legend('Population','Ammonia','Glucose','Ethanol')
Conclusions
The re-visited model doesn't seem to be producing a terribly different result from what my original model was producing, so if pursued further I would look more into the best way to model the R and V2 terms in terms of alcohol and yeast population. Overall I think I'm on a better track at least conceptually in terms of what the model was attempting to do in reducing the fermentation term to just being in terms of alcohol without the metabolic terms included.
References
Albertin, W., Marullo, P., Aigle, M., Dillmann, C., de Vienne, D., Bely, M., & Sicard, D. (2011). Population Size Drives Industrial Saccharomyces cerevisiae Alcoholic Fermentation and Is under Genetic Control .Applied and Environmental Microbiology, 77(8), 2772–2784. http://doi.org/10.1128/AEM.02547-10
Dahlquist, Kam D. (2017) BIOL398-05/S17:Week 5. Retrieved from http://www.openwetware.org/wiki/BIOL398-05/S17:Week_5 on 1 March 2017
Brauer, M. J., Saldanha, A. J., Dolinski, K., & Botstein, D. (2005). Homeostatic adjustment and metabolic remodeling in glucose-limited yeast cultures. Molecular biology of the cell, 16(5), 2503-2517. doi: 10.1091/mbc.E04-11-0968
ter Schure, E. G., Sillje, H. H., Verkleij, A. J., Boonstra, J., & Verrips, C. T. (1995). The concentration of ammonia regulates nitrogen metabolism in Saccharomyces cerevisiae. Journal of bacteriology, 177(22), 6672-6675.
Acknowledgments
- Dr. Fitzpatrick for the notes he gave at the end of class on March 16, 2017 helped in re-structuring my thinking on this project
- Worked with Lauren Kelly on our respective equations in Seaver 120 on March 22, 2017
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
Margaret J. Oneil 03:13, 23 March 2017 (EDT)