Lkelly9 Week 10
From OpenWetWare
Jump to navigationJump to search
Navigation Links
- Lauren M. Kelly
- Assignment Page
- Individual Journal Entry
- Shared Journal Page
Purpose
- The purpose of this assignment was to read and outline the assigned paper and prepare a journal club presentation to demonstrate our understanding and mastery of the content.
Journal Club 2 Presentation
Journal Club 2 - Lauren M. Kelly and Cameron M. Rehmani Seraji
Definitions
- In vivo: (Of a biological process) made to occur within the living organism
- Diurnal: A biological rhythm that primarily express a periodicity during daylight hours
- Mannoproteins: yeast cell wall components that are proteins with large numbers of mannose groups attached; highly antigenic.
- Transcriptome: all the RNAs present in a cell type
- Motif: The smallest group of atoms in a polymer that, when under the influence of a rotation-translation operator, will assemble the rest of the atoms in the chain.
- Biogenesis:(1) The process in which life forms arise from similar life forms., (2) It asserts that living things can only be produced by another living thing, and not by a non-living thing.
- Desaturase: Any of several enzymes that put double bonds into the hydrocarbon areas of fatty acids.
- Prototrophic Strains: Strain's that have the same nutritional requirements as the wild-type strain.
- Orthologues: Any gene that can be found in two or more different species that can be traced back to the same common ancestor.
- Exogenous: Developed or originating outside the organism, as exogenous disease.
Outline
- What is the main result presented in this paper?
- DNA microarray analysis showed that 494 genes in the glucose-limited cultures and 806 genes in the nitrogen-limited cultures had significantly different transcript levels at the two temperatures.
- Other environmental parameters have a significant effect on the transcriptional response to a stimulus.
- Cannot change one parameter without any impact on others.
- The use of both glucose- and ammonium-limited cultures enabled the identification of core sets of genes that have a context-independent response to single environmental stimuli.
- Growing evidence that the specific growth rate affects genome-wide transcription
- Acclimatized growth at low temperatures does not involve a Msn2/Msn4-complex regulatory system. It is based on something else that has not been identified.
- Identified a set of 235 genes that showed a consistent transcriptional response to low temperature, regardless of the limiting nutrient.
- A group of genes involved in lipid metabolism was the only group that was clearly regulated in both low-temperature chemostats and batch cultures.
- HSP26 and HSP42 were down regulated at low temperature in the chemostat culture and up regulated at low temperature in the batch cultures.
- Study overall demonstrates that responses to low temperatures and low specific growth rate can be separated by using chemostat cultures.
- What is the importance or significance of this work?
- This work pointed out important differences between batch cultures and chemostat cultures, such as the importance of discriminating different phases in the adaptation to environmental change. The chemostat cultures are able to control more variables.
- It is also emphasized that low-temperature acclimation of yeast involves more than transcriptional reprogramming.
- Intracellular metabolite levels are also adjusted.
- Create a flow chart to describe their methods.
- 4 Different Growth Conditions
- 12°C, glucose-limited
- 12°C, ammonium-limited
- 30°C, glucose-limited
- 30°C, ammonium-limited
- Utilized various analytical methods and microarray analysis and compared their data to other S. cerevisiae low-temperature transcriptome datasets.
- 4 Different Growth Conditions
- How did they treat the cells (what experiment were they doing?)
- They aimed to see how yeast cells acclimate to low temperature in chemostat environments with different nutrient-limited conditions.
- What strain(s) of yeast did they use? Was the strain haploid or diploid?
- Strain: prototrophic, haploid reference S. cerevisiae strain CEN.PK113-7D (MATa)
- What media did they grow them in? Under what conditions and temperatures?
- 2.0 L chemostats, with a working volume of 1.0 L
- dilution rate (D) = 0.03 h-1
- pH kept at 5.0
- Stirrer speed = 600 rpm
- Grown at both 12°C and 30°C.
- Defined synthetic medium limited by carbon or by nitrogen. All other growth requirements were in excess.
- Anaerobic conditions
- Before sampling, biomass dry weight, metabolites, dissolved oxygen, and gas profiles were constant for at least 3 volume changes.
- 2.0 L chemostats, with a working volume of 1.0 L
- What controls did they use?
- The yeast grown at 30°C acted as a control to compare the yeast grown at 12°C to.
- A combination of both glucose- and ammonium-limited cultures reduced the impact of secondary effects.
- How many replicates did they perform per condition?
- In the microarray analysis, the results for each growth condition were derived from three independently cultured replicates.
- It is unclear how many chemostats with each set of conditions were used.
- It appears to be two glucose-limited (12°C and 30°C) and two ammonium-limited (12°C and 30°C).
- What mathematical/statistical method did they use to analyze the data?
- Cells sampled from chemostats, preparation of probes, and hybridization to Affymetrix Genechip microarrays. (Precise methods described in Piper et al. (2002))
- Agilent 2100 Bioanalyzer: determined RNA quality
- Microsoft Excel running the significance analysis of microarrays add-in was used for pair-wise comparisons.
- Expressionist Analyst version 3.2: Generated venn diagrams and heat-map visualizations of transcript data
- Regulatory Sequence Analysis (RSA) Tools: performed promoter analysis
- Database for Annotation, Visualization and Integrated Discovery (DAVID) 2006: Statistical assessment of over-representation of GO biological processes categories among sets of significantly changed transcripts
- Fisher's test: overrepresentation of transcription-factor binding sites as defined by chromatin immunoprecipitation (ChIP)-on-chip analysis
- Probability was calculated using the equation on the right.
- What transcription factors did they talk about?
- Msn2/Msn4
- Known to regulate key genes in carbohydrate synthesis via STREs
- Their results indicated that low-temperature acclimatized growth does not involve a Msn2/Msn4-complex regulatory role but is based on a different mode of regulation.
- Skf1p and Stt4p
- involved in a pathway required to generate phosphatidylinositol, as well as in other cellular processes.
- Ino2/Ino4
- control YEL073C; regulate phospholipid biosynthesis genes
- Mbp1p
- Hap2-Hap1
- Hap3-Hap1
- Fhl1p
- Sfp1p
- Gln3p
- Gln3-Dal82
- Hap2-Dal82
- Aft2p
- Hsf1p
- Nrg1p
- Phd1p
- Rcs1p
- Rox1p
- Sok2p
- Nrg1-Aft2
- Phd1-Nrg1
- Rox1-Phd1
- Sok2-Nrg1
- Msn2/Msn4
- Briefly state the result shown in each of the figures and tables.
- Table 1. Physiological Characteristics of S. Cerevisiae grown in ammonium- and glucose-limited anaerobic chemostat cultures

Table. 1 - The values in this table are the mean +/- SD of data from three chemostat cultivations
- Biomass yields and fermentation rates were similar at 12 and 30°C in both glucose- and ammonium-limited chemostat cultures
- growth efficiency was not severely affected by growth temperature
- In cultures grown at 12°C, the residual concentrations of glucose and ammonium were 7.5-10-fold higher than in the cultures frown at 30°C. This indicates temperature dependence of the residual nutrient concentration.
- Figure 1. Global transcriptome responses to anaerobic growth at 12 and 30°C in anaerobic glucose- and ammonium-limited chemostat cultures (D = 0.03 h-1)
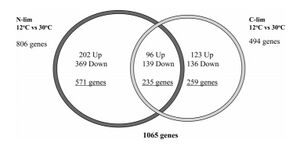
Figure 1. - Displays the significant differently expressed genes between 12 and 30°C in both C and N limitations
- The left side shows the number of significant differently expressed genes between the N-limited 12 and 30°C chemostat cultures, and the right side shows the number of significant differently expressed genes between the C-limited 12 and 30°C chemostat cultures. The center area represents the number of genes that were differently expressed in both N-limited and C-limited conditions.
- Figure 2. Heat map representing the transcript level ratio of 1065 differently expressed genes in anaerobic glucose- and ammonium-limited chemostat cultures grown at 12 and 30°C.

Figure 2. - Used to identify regulatory networks involved in the acclimation of the yeast to low temperature
- Temperature-responsive genes were screened for enrichment of specific funtional categories.
- Categories include lipid metabolism, carbohydrate transport, rRNA processing, electron transport, amino acid transport, hexose metabolism, protein synthesis/protein complex assembly, ribosome biogenesis and assembly/RNA processing/rRNA processing, nitrogen compound metabolism and catabolism/amine transport/allantoin metabolism, polysaccharide and trehalose metabolism, M phase of mitotic cell cycle and chromosome segregation, cellular morphogenesis, response to stimulus, nuclear export, carbohydrate metabolism, and transport.
- The genes belong to the enriched GO categories
- Table 2. Protein and storage carbohydrates contents of S. cerevisiae biomass grown in ammonium- and glucose-limited anaerobic chemostat cultures.

Table 2. - Displays measurements of biomass dry weight, whole cell protein, biomass nitrogen content, trehalose, and glycogen.
- In ammonium-limited cultures, trehalose and glycogen contents were significantly lower at 12 than at 30°C.
- In glucose-limited cultures, trehalose contents were lower at 12°C, but the glycogen content was higher than it was at 30°C.
- Yeast acclimation to low temperature does not require accumulation of glycogen and trehalose and the transcriptional induction of the genes involved in the synthesis of these compounds.
- Table 3. Idenfification of (A) significantly overrepresented cis-regulatory binding motifs in 5' upstream regions and (B) significantly overrepresented promoter elements that bind known transcription factors (TF) or TF pairs according to ChiP-on-chip analysis in low temperature up- and down-regulated gene clusters derived from C-Lim and N-Lim chemostat experiments.
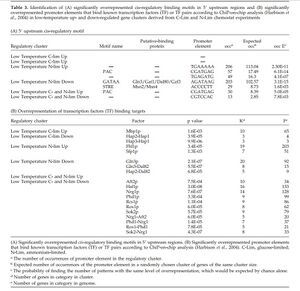
Table 3. - Overrepresentation of STRE elements in the upstream regions of genes that showed a reduced transcript level at 12°C in the nitrogen-limited cultures.
- Figure 3. Genes differentially expressed in batch cultures during adaptation to low temperature. (A) Venn diagram showing the number of genes that are common to three batch-cultire studies on low-temperature transcriptional adaptation. (B) Heat map representing the transcript ratio of 259 genes found in common in the three batch-culture low-temperature transcriptome datasets.
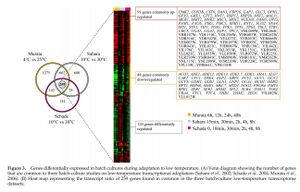

Figure 3. - Response of genes was not always consistent.
- Figure 4. Comparison of the transcript ratio of 259 genes common to three batch-culture low-temperature transcriptome datasets with the 12°C/30°C ratio of the genes specifically up- and down-regulated in anaerobic glucose- and ammonium-limited chemostat cultures. Venn diagrams show the number of low-temperature-responsive genes common to the batch-culture and chemostat-based datasets. The heat map shows the expression rations of the genes common to batch and chemostat datasets. The genes indicated between brackets show a consistent transcriptional regulation at low temperature in all datasets.
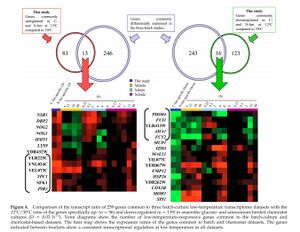
Figure 4. - 29 genes were transcriptionally regulated during both adaptation and acclimation to low temperature; only 11 showed a consistent pattern of regulation in all four situations.
- Three genes encoding transporters were found among the five consistently down-regulated genes.
- Figure 5. Comparison of the genes specifically up- or down-regulated during acclimation (this study) or adaptation (overlap of other datasets) to low temperature with the growth rate-dependent genes identified by Castrillo et al (2007) snf Regenberg et al (2006).
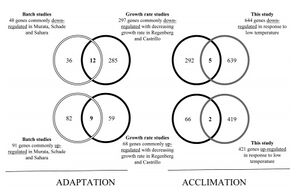
Figure 5. - The altered transcript levels of 25% of the low temperature down-regulated genes and 10% of the low temperature up-regulated genes are likely to have been primarily related to specific growth rate, rather than to temperature.
- Figure 6. Comparison of the genes specifically up- or down- regulated during acclimation (this study) or adaptation (overlap datasets from other studies) to low temperature with the ESR genes identified by Gasch et al. (2000).
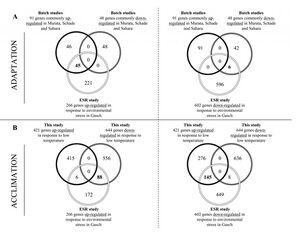
Figure 6. - An extensive overlap was revealed. 50% of the consistently low temperature up-regulated genes and 13% of the low temperature down-regulated genes found in the batch culture studies were also found in the ESR genes identified by Gasch et al. (2000).
- 1/3 of the low temperature-responsive genes found in the three batch culture studies could be linked to ESR.
- Several genes up- or down-regulated in the Gasch et al. (2000) study showed an opposite transcriptional response in the low-temperature chemostat cultures
- ESR is not an obligatory response to growth at low temperature, but occurs during adaptation upon a sudden exposure to sub-optimal temperatures.
- Table 1. Physiological Characteristics of S. Cerevisiae grown in ammonium- and glucose-limited anaerobic chemostat cultures
Acknowledgments
- Worked with Cameron M. Rehmani Seraji in Seaver 120. We discussed the journal club power point presentation.
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
Lauren M. Kelly 19:21, 29 March 2017 (EDT)
References
- Dahlquist, Kam D. (2017) BIOL398-05/S17:Week 10. Retrieved from http://www.openwetware.org/wiki/BIOL398-05/S17:Week_10 on 28 March 2017.
- Tai, S. L., Daran-Lapujade, P., Walsh, M. C., Pronk, J. T., & Daran, J. M. (2007). Acclimation of Saccharomyces cerevisiae to low temperature: a chemostat-based transcriptome analysis. Molecular biology of the cell, 18(12), 5100-5112. doi: 10.1091/mbc.E07-02-0131