CAMRI:HowToScan
CAMRI Policy on Unexpected EventsIf an unexpected, unusual or untoward event occurs during a scan, please send an e-mail with a brief description of the event to camri@bcm.edu as soon as possible. This will allow CAMRI to be responsive and proactive in addressing any issues. The e-mail does not need to be lengthy but is required for both scanner-related and subject-related events. Example e-mails:
In addition to specific problems that you encounter, we always welcome user feedback on CAMRI operations. Adverse Event Reporting to IRBThe CAMRI requirement to report unexpected events is distinct from the enduring requirement to notify the IRB of any adverse event occurring to human subjects enrolled in a protocol. An adverse event is defined as “any untoward medical occurrence in a participant, which does not necessarily have a causal relationship with the trial intervention. An adverse event (AE) can, therefore, be any unfavorable and unintended sign (including an abnormal laboratory finding), symptom, or disease.” For more information, please review the highlighted section (pages 150 - 151) in the BCM IRB Manual. Media:irb-manual.pdf Saving DataThere are three ways to save the data that you have collected at great expense. CAMRI does not back up data. Users are responsible for saving their own data! Method 1 (Easiest): Use the Flywheel ServerPlease click on the Flywheel tab above or go to https://openwetware.org/wiki/CAMRI:Flywheel for further instructions on backing up data using that resource. Method 2: Use BoxThis requires that your lab have a Box account; contact Dave Lee (dlee@bcm.edu) to obtain one.
Method 3: Burning a Backup Disk
Human Subjects ProtectionTo scan at CAMRI, approval from your home institution institutional review board (IRB) or committee for the protection of human subjects (CPHS) is required. Click here for an IRB proposal template. MRI Data AnalysisClick here for more information on MRI data analysis. Change LogNotes on Recent Changes to Our Equipment. Updated scanner 5 to new computer and imager. Still running at VB17 software. Scheduling and Visitor policyTo schedule or cancel time on the scanner, you must use the iLab web site https://bcm.corefacilities.org/service_center/show_external/3684 Click here for detailed instructions.
If bringing subjects in after hours please use the Cullen entrance to check yourself and your participants in. About CAMRI's Siemens ScannersThe BCM CAMRI has 2 whole-body 3T Siemens Scanners labeled 3T-4 and 3T-5. Scanner 3T-5 is a Magnetom Trio that has a 12 channel and 32 channel head coils, Spine coil and flex body coils. Scanner 3T-4 is a Magnetom PRISMA Fit with a Head/Neck 20 channel, Head/Neck 64 channel, Spine 32 channel, Flex 4 Large, Flex 4 Small, and body coil. More information about the difference between the coils can be found here: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3538896/ Both scanners have a bore inner tunnel diameter of 60 cm. TroubleshootingReport all issues to camri@bcm.edu http://openwetware.org/images/0/01/CAMRI_MRI_Safety_Manual.pdf There are a few helpful videos that you can watch on YOUTUBE that will help you become a little more familiarized with our systems here at CAMRI. Here are the links: https://www.youtube.com/watch?v=pi1OLUflhTU
Having problems with...
The sequence of operation is important so make sure it is done in this order. Response RecordingResponses are recorded with a fORP system. Link to Jung Hwan's user guide here: fORP Configuration
Bold Screen SetupFor Scanner 5 Bold Screen Setup: Media:Scanner5.docx For Scanner 3 Bold Screen Setup: Media:Scanner3.docx
Console CamThe Console Cam is provided for users to obtain scanner console support remotely from CAMRI staff without the staff member having to enter the room. It will broadcast an image of the console and allow voice communications via Zoom conferencing. Please use the iMac in the control room that is next to the Scanner Console to use the Console Cam. You can use this account to log into the iMac; account: camri
password: camri
After you are logged in start a new meeting in Zoom and copy the invite to the clipboard. You can then use your own web mail to paste the invite info and send the email to the person you are requesting support from. The arm of the Console Cam is flexible and you may have to adjust it so that the screen is centered so that the viewer can see the entire screen. Visual Stimulus Displays3T-3 and 3T-5 display visual stimuli on direct view LCD screens.
3T-3 uses a [24" display with resolution of 1920x1200] Link to manual here: Cambridge Research Systems BOLD Screen User Guide The BOLDScreen has a built-in color profile that can make images appear too dark. Instead, use the default color profile as follows:
Ricky Savjani has redone the gamma calibration on the 3T-5 BOLDscreen, this can be accessed in PsychToolbox as follows. First, Click here to load the table % load in the gamma table
load CRS_32BOLDSCREEN/gammaTable_Spline_fit.mat
% after opening a window, call this to load the load the gamma table to to your Screen
Screen('LoadNormalizedGammaTable', win, gammaTable2*[1 1 1]);
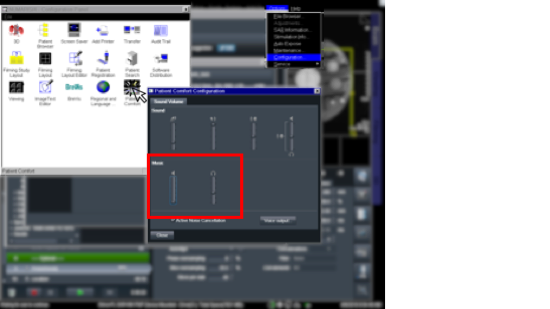
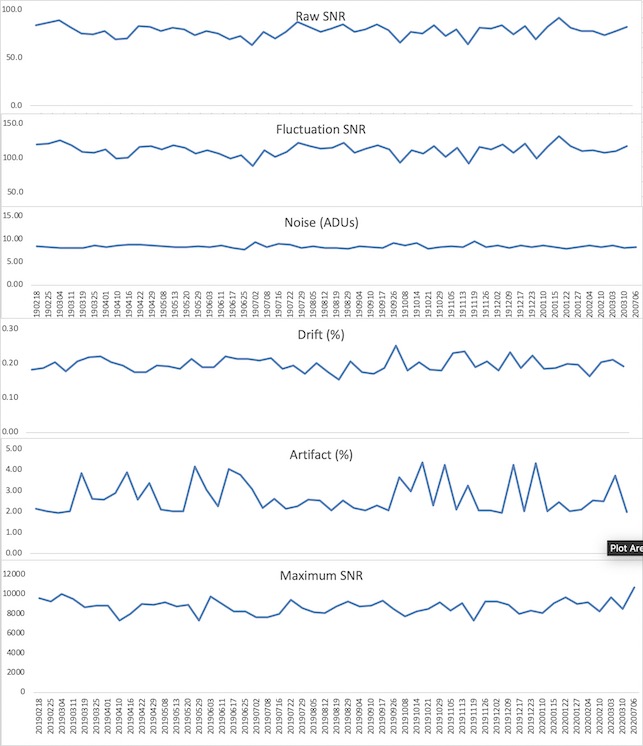
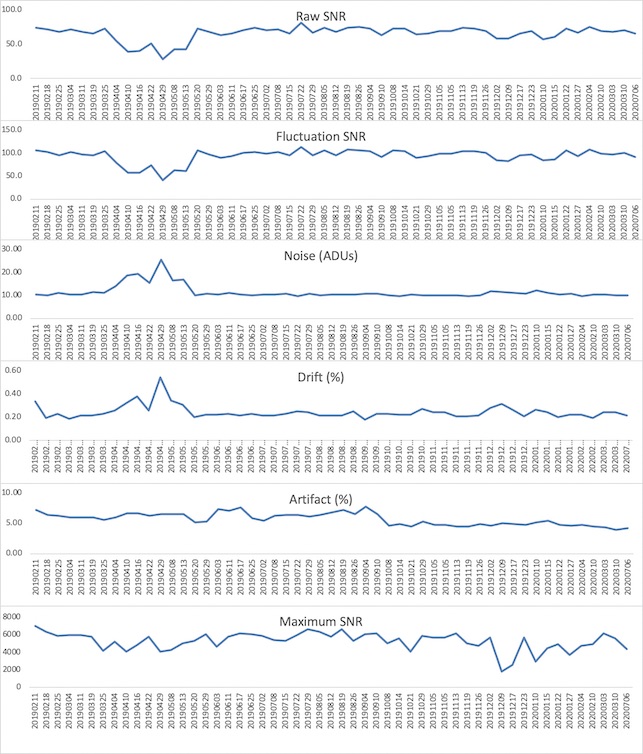
Now, desired intensity levels will be mapped appropriately to the screen. Rough calculation of image size in degrees of visual angle: roughly 76.5 cm from screen to eyeball height of screen is 13.7 in = 34. 8 cm width of screen is 34.8 * (1024/768) = 46.4 cm atan(23.2/76.5) = 0.29 rad = 16.6 degrees ~30 degrees full width Audio (headphones)Oneway to deliver auditory stimuli to your participants is through the headphones. These are Siemens equipment and are controlled by the scanner. First, make sure the headphone cord is plugged into to your stimulus presentation computer (laptop, desktop, boombox, etc). Then, plug the other end into the back of the intercom box (the same one that's used to speak with the participant while they're in the scanner.) If you're not hearing anything, go to Options -> Configuration. A new window will pop up with icons. Choose "Patient Comfort." In the next window, make sure the volume bars under "Music" are not at the very bottom (which is the default.) . The speaker icon indicates sounds send to the room, and the headphone bar is specific to sound sent through the headphones. ASL HelpThis ppt describes the units when you draw an ROI on the relCBV map (BW). You cannot draw on the Perfusion Color map the units are not scaled. Media:ASL.ppt Quality AssuranceWeekly QA scans are performed on both the Prisma and Trio scanners. Below are the results of the previous 52 week EPI QA runs.
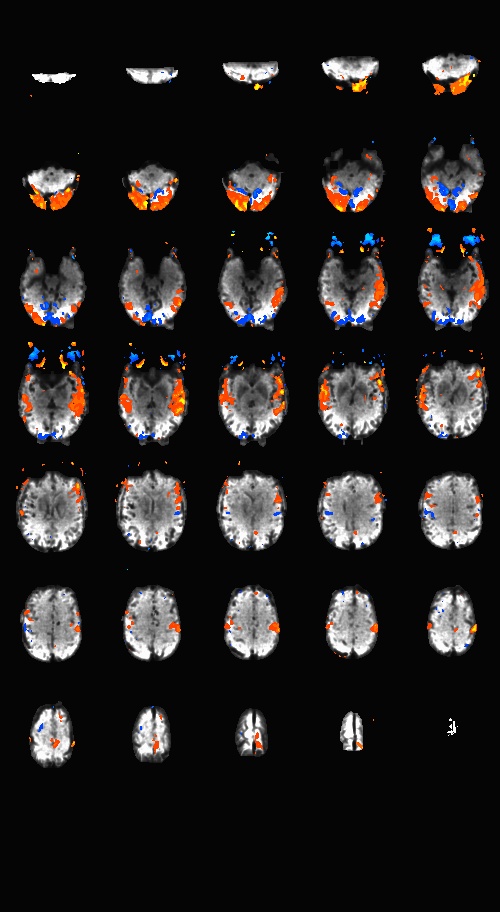
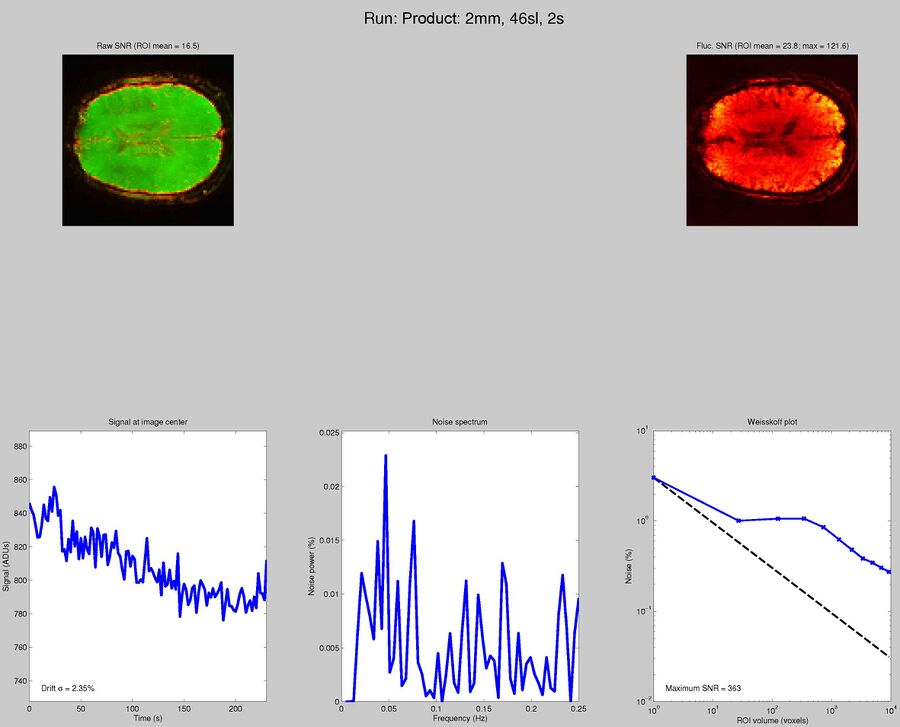
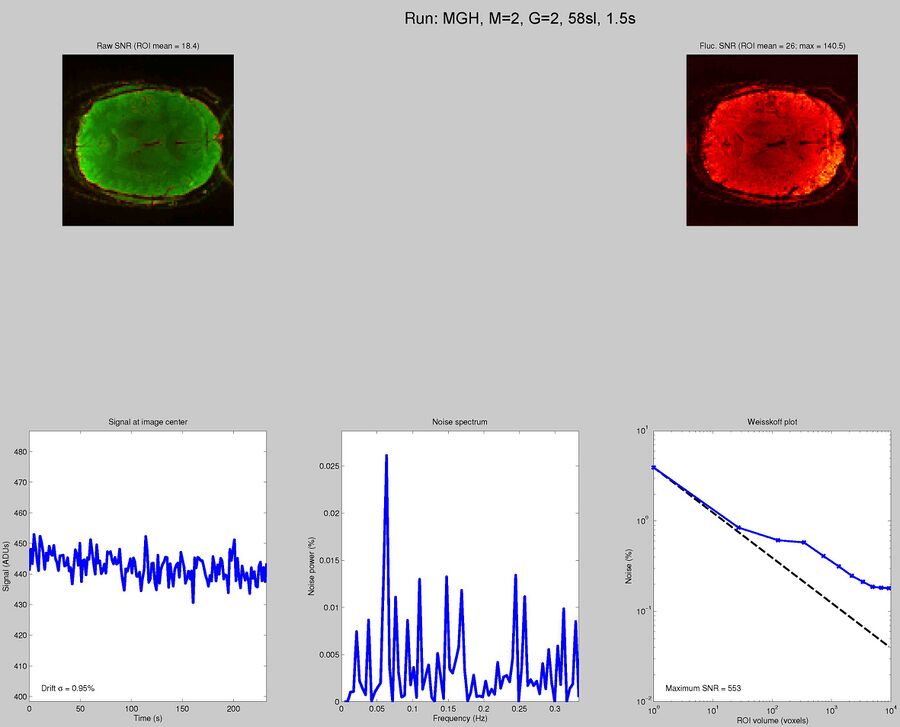
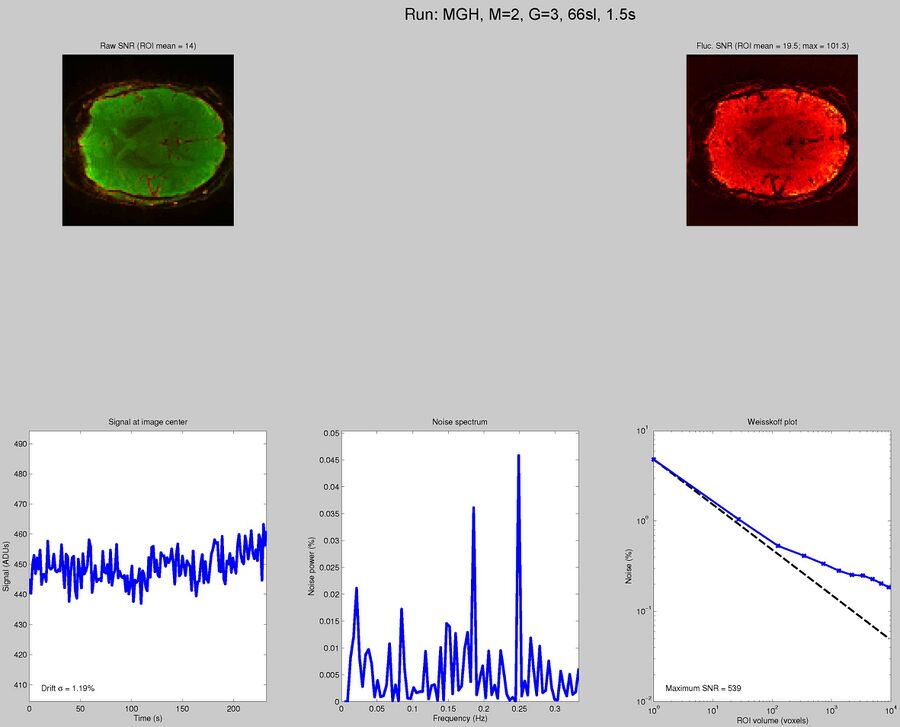
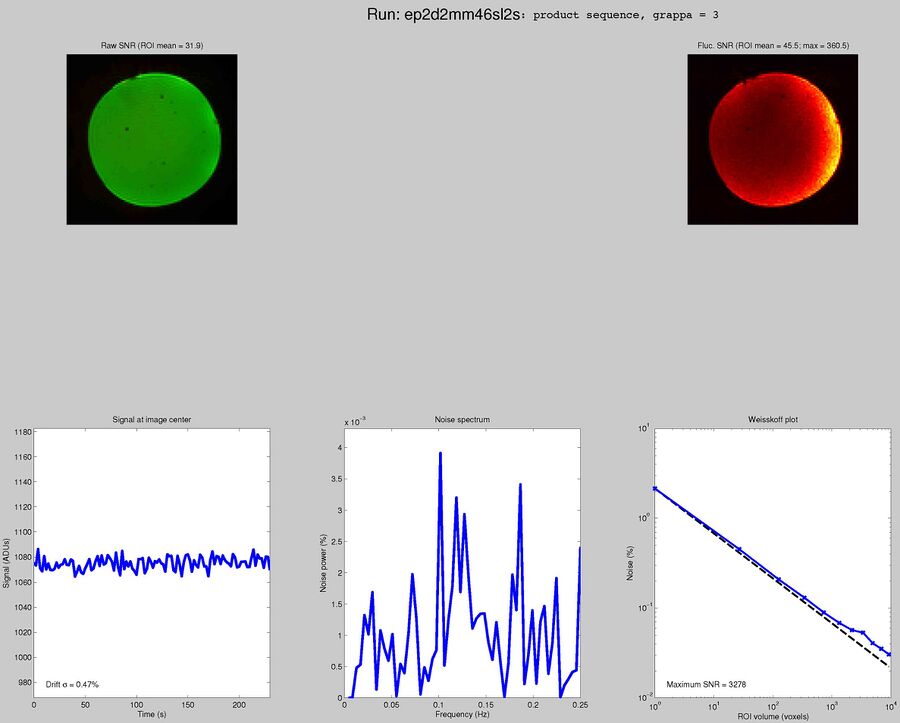
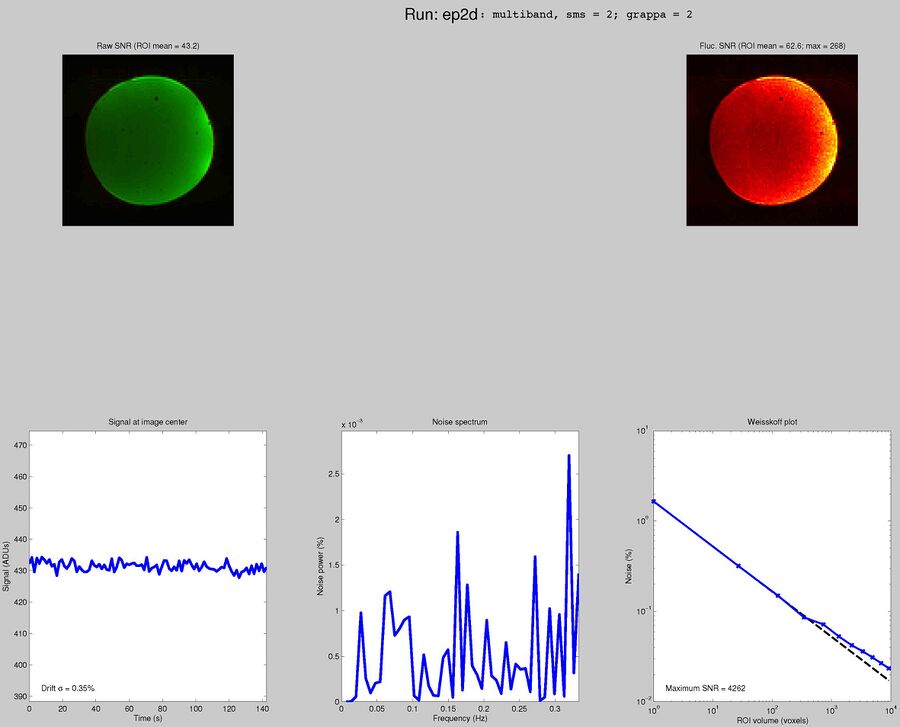
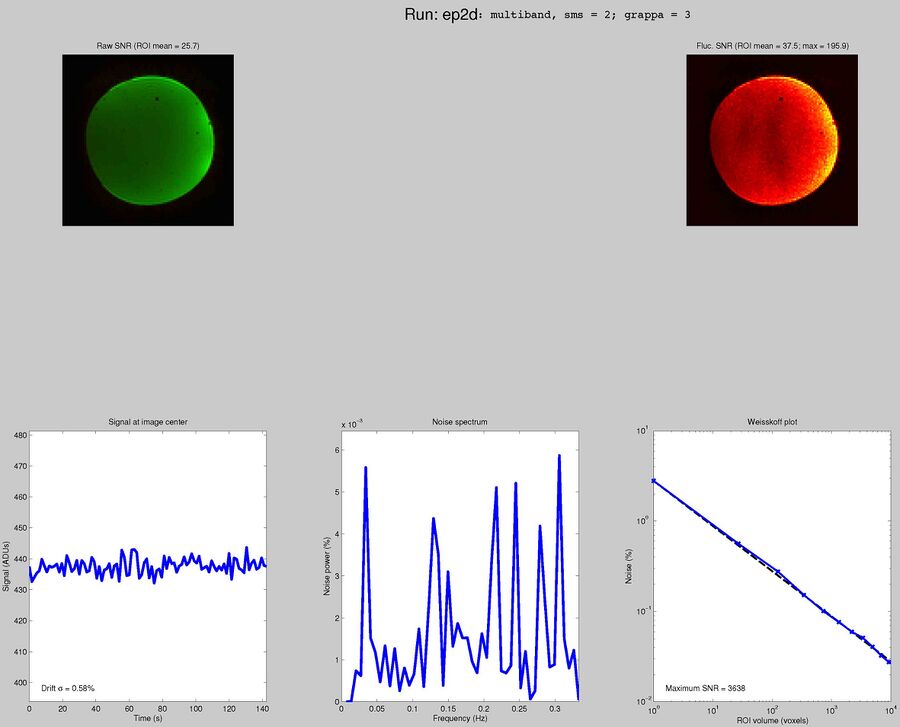
Sequences to UseWhat pulse sequence to use for fMRIThere are a variety of different pulse sequences that can be used for fMRI, including the standard Siemens EPI product sequences; multi-band EPI sequences from MGH and Univ. of Minnesota; and spiral sequences implemented by David Ress.
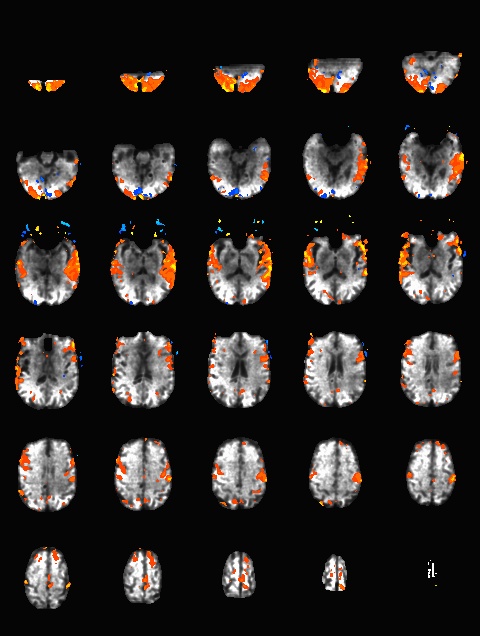
Here is another analysis of the same data by David Ress. The results are similar, with the G2, MB2 version winning by around 15% in fSNR over the product sequence. But, the shorter TR will also net you an improvement of ~5% in CNR in a similar length run. It is also interesting, however, that the G3 run had less aliasing artifact, suggesting that shorter acquisition times reconstruct more cleanly. Similar results were obtained with a phantom: What T1-weighted Anatomicals to UseMost studies collect at least one scan with structural contrast in order to align their other runs (functional, DTI, etc) to other participants and to published atlases. In some cases, measurements such as volume or cortical thickness are also derived from these scans. We have tested a few of these on the Prisma (Bay 4) and will run similar analyses on the Trio (Bay 5) soon. CAMRI (specifically, Meghan) recommends the following scans for new studies, and can easily set these up in your protocol: (1) 1mm isotropic scan. Acquisition time: 4:01. Contrast to noise ratio (CNR): ~2.0. This is a good choice if speed is of the essence, and your main goals are alignment of lower-resolution scans. (2) 1mm isotropic scan. Acquisition time 5:12. CNR: ~2.6. High quality T1 scan that will suit most studies. Very good CNR. (3) 800um isotropic scan. Acquisition time: 6:38. CNR: ~2.2. A happy medium between speed and resolution. (4) 700um isotropic scan. Acquisition time: 8:18. CNR ~ 2.2. Based on the HCP sequences. This is recommended if your study would benefit from higher resolution. For instance, if you plan to do cortical thickness studies or are interested in segmentation of fine structures, this could be a good choice. A note on CNR: This was calculated from only a handful of people (sometimes, just from one participant). CNR was calculated using FreeSurfer segmentations of the cortical ribbon and the white matter mask. CNR=(Mean(white) - mean(grey)) / mean(std(grey), std(white)). Signal was taken from the raw image, with prescan normalize turned ON. Misc NotesSlice OrderingDavid Ress' former student Andrew Florens examined the acquisition time stamps and found that axial slices are collected from Superior to Inferior (head to foot). If an odd number of slices is acquired, the slice order is 1, 3, 5, 7, etc. 2, 4, 6, 8 etc. If an even number of slices is acquired, the slice order is reversed: 2, 4, 6, 8, etc. 1, 3, 5, 7, etc. Editing Dicom HeadersInformation regarding how to edit and anonymize DICOM images can be found: Media:How_to_anonymize_and_edit_Dicom_headers_utilizing_Sante_Dicom_Editor.pdf |