Yaniv Maddahi Journal Week 4
From OpenWetWare
Jump to navigationJump to search
Purpose
- The purpose of this weeks assignment is to obtain sequence data for the SARS-CoV2 spike protein, compare it using multiple sequence alignments, and analyze it for phylogenetic relationships using trees. Further, we are continuing the discussion from last time regarding scientific data and research. We will be creating our own phylogenetic trees as well as sequence a set of spike protein sequences that differ slightly from those of the article read beforehand. We will be analyzing our trees to relate the ways in which branching occurs to the amino acid sequence of specific proteins and further comparing and contrasting our own sequence and tree to the sequence and tree of the article.
Methods and Results
Part 1
- I chose one of the GenBank records DQ071615: Bat SARS coronavirus Rp3, complete genome.
- GenBank records contain information regarding the genomic sequence of a virus, its locus, publications, source, organism and classifications, references, information regarding publishing of the virus, codons within the virus, translation sqeuence, and more.
- I then downloaded the nucleotide sequence in FASTA format to my local hard drive.
- I clicked the "Send to link" in the upper right of the page, selected Complete Record, file as the Destination, and FASTA as the format, and created the file.
- I opened the file that I saved with a word processor to confirm that I had the sequence and that it is in the FASTA format. In the FASTA format each sequence is preceded by a label which begins with the greater than sign (>). I searched for the GenBank record associated with that sequence and located the spike protein accession number in the GenBank record. (Note that the spike protein is sometimes called the "S" protein.)
- I then downloaded my assigned protein sequence in FASTA format, just like I did for the whole genome sequence.
- I began every line with a space character as it will be interpreted as a fixed width font and the sequences will line up nicely on the page.
Part 2
- In order to analyze sequence data I used www.phylogeny.fr, a free, simple to use web service dedicated to reconstructing and analysing phylogenetic relationships between molecular sequences.
- In my browser, I went to the website www.phylogeny.fr. I scrolled down on the page to the section labeled ‘Phylogeny analysis’, and clicked on the text ‘One Click’.
- I then clicked in the large text field labeled ‘Upload your set of sequences in FASTA, EMBL, or NEXUS format' and copied the list of sequences from the talk page and used Ctrl-V (or command-V) to paste my sequences here. I then clicked the “Submit” button.
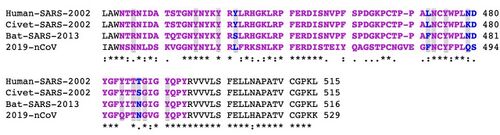
- After my alignment was complete,I found the numbered tabs located just beneath the text One Click Mode, and clicked on the tab labeled 3. Alignment. Within the alignment, individual positions are color-coded to indicate their conservation, or how similar the sequences are to each other at that position. Blue highlighting indicates high conservation (i.e., the sequences are identical or at least very similar), while gray highlighting indicates lower conservation and white highlighting indicates little if any conservation.
- Near the bottom of the page, under Outputs, I clicked on Alignment in Clustal format. This displayed the alignment in a text-only format in which each position's conservation is indicated by a symbol underneath the alignment block (“*” for invariant, “:” for highly conserved, “.” for weakly conserved, and a space for not conserved).
Spike Protein Sequence
QDF43825.1 ---------MKLLVLV-----FATLVSSYTIEKCTDFD------DRTPPSNTQFLSSHRG
AGZ48818.1 ---------MKLLVLV-----FATLVSSYTIEKCLDFD------DRTPPANTQFLSSHRG
ALK02457.1 ----------MFIFLF-----FLTLTSGSDLESCTTFD------DVQAPNYPQHSSSRRG
AAS10463.1 ----------MFIFLL-----FLTLTSGSDLDRCTTFD------DVQAPNYTQHTSSMRG
AAP13441.1 ----------MFIFLL-----FLTLTSGSDLDRCTTFD------DVQAPNYTQHTSSMRG
AAP13567.1 ----------MFIFLL-----FLTLTSGSDLDRCTTFD------DVQAPNYTQHTSSMRG
QHD43416.1 ----------MFVFLV-----LLPLVSSQ----CVNLT------TRTQLPPAYTNSFTRG
AVP78031.1 -----------MLFFL-----FLQFALVN--SQCVNLT------GRTPLNPNYTNSSQRG
ABD75323.1 --------MKILIFAF-----LVTLVKAQ--EGCGVIN------LRTQPKLTQVSSSRRG
QDF43835.1 --------MKVLIVLL-----CLGLVTAQ--DGCGHIS------TKPQPLLDKFSSSRRG
ABD75332.1 --------MKVLIFAL-----LFSLAKAQ--EGCGIIS------RKPQPKMEKVSSSRRG
QDF43820.1 --------MKILIFAF-----LVTLVEAQ--EGCGIIS------RKPQPKMAQVSSSRRG
AAZ67052.1 --------MKILILAF-----LASLAKAQ--EGCGIIS------RKPQPKMAQVSSSRRG
AFS88936.1 ----MIHSVFLLMFLLTPTESYVDVGPDSVKSACIEVDIQQTFFDKTWPRPIDVSKA-DG
YP_0010399 MTLLMCLLMSLLIFVRGCDSQFVDMSPASNTSECLESQVDAAAFSKLMWPYPIDPSKVDG
::. . * . *
QDF43825.1 VYYPDDIFRSNVLHLVQDHFLPFDSNVTRFITFGLN-------------FDN---PIIPF
AGZ48818.1 VYYPDDIFRSNVLHLVQDHFLPFDSNVTRFITFGLN-------------FDN---PIIPF
ALK02457.1 VYYPDEIFRSDTLYLTQDLFLPFYSNVTGFHTINHR-------------FDN---PVIPF
AAS10463.1 VYYPDEIFRSDTLYLTQDLFLPFYSNVTGFHTINHT-------------FDD---PVIPF
AAP13441.1 VYYPDEIFRSDTLYLTQDLFLPFYSNVTGFHTINHT-------------FGN---PVIPF
AAP13567.1 VYYPDEIFRSDTLYLTQDLFLPFYSNVTGFHTINHT-------------FDN---PVIPF
QHD43416.1 VYYPDKVFRSSVLHSTQDLFLPFFSNVTWFHAIHVS------GTNGTKRFDN---PVLPF
AVP78031.1 VYYPDTIYRSDTLVLSQGYFLPFYSNVSWYYSLTTN-------NAATKRTDN---PILDF
ABD75323.1 VYYNDDIFRSDVLHLTQDYFLPFHSNLTQYFSLNIE-------SDKIVYFDN---PILKF
QDF43835.1 VYYNDDIFRSDVLHLTQDYFLPFDTNLTRYLSFNMD-------SATKVYFDN---PTLPF
ABD75332.1 VYYNDDIFRSDVLHLTQDYFLPFDSNLTQYFSLNID-------SNKYTYFDN---PILDF
QDF43820.1 VYYNDDIFRSDVLHLTQDYFLPFDSNLTQYFSLNVD-------SDRYTYFDN---PILDF
AAZ67052.1 VYYNDDIFRSNVLHLTQDYFLPFDSNLTQYFSLNVD-------SDRFTYFDN---PILDF
AFS88936.1 IIYPQGRTYSNITITYQGLF-PYQGDHGDMYVYSAG--HATGTTPQKLFVANYSQDVKQF
YP_0010399 IIYPLGRTYSNITLAYTGLF-PLQGDLGSQYLYSVSHAVGHDGDPTKAYISNYSLLVNDF
: * *. . * * : : *
QDF43825.1 RDGVYF----AATEKSNVIRG-------------WVFGSTMNNKSQ---------SVIIM
AGZ48818.1 KDGIYF----AATEKSNVIRG-------------WVFGSTMNNKSQ---------SVIIM
ALK02457.1 KDGVYF----AATEKSNVVRG-------------WVFGSTMNNKSQ---------SVIII
AAS10463.1 KDGIYF----AATEKSNVVRG-------------WVFGSTMNNKSQ---------SVIII
AAP13441.1 KDGIYF----AATEKSNVVRG-------------WVFGSTMNNKSQ---------SVIII
AAP13567.1 KDGIYF----AATEKSNVVRG-------------WVFGSTMNNKSQ---------SVIII
QHD43416.1 NDGVYF----ASTEKSNIIRG-------------WIFGTTLDSKTQ---------SLLIV
AVP78031.1 KDGIYF----AATEHSNIIRG-------------WIFGTTLDNTSQ---------SLLIV
ABD75323.1 GDGVYF----AATEKSNVIRG-------------WVFGSTFDNTTQ---------SAIIV
QDF43835.1 GDGIYF----AATEKSNVVRG-------------WIFGSTMDNTTQ---------SAIIV
ABD75332.1 GDGVYF----AATEKSNVIRG-------------WIFGSSFDNTTQ---------SAIIV
QDF43820.1 GDGVYF----AATEKSNVIRG-------------WIFGSTFDNTTQ---------SAVIV
AAZ67052.1 GDGVYF----AATEKSNVIRG-------------WIFGSTFDNTTQ---------SAVIV
AFS88936.1 ANGFVVRIGAAANSTGTVIISPSTSATIRKIYPAFMLGSSVGNFSDGKMGRFFNHTLVLL
YP_0010399 DNGFVVRIGAAANSTGTIVISPSVNTKIKKAYPAFILGSSLTNTSAGQ-PLYANYSLTII
:*. . *:.. ..:: . :::*::. . : : ::
QDF43825.1 NNSTNLVIRACNFELCDNPFFVVLRSNNTQIPSY------IFNNAFN-CTFEYVSKDFNL
AGZ48818.1 NNSTNLVIRACNFELCDNPFFVVLKSNNTQIPSY------IFNNAFN-CTFEYVSKDFNL
ALK02457.1 NNSTNVVIRACNFELCDNPFFAVSKPTGTQTHTM------IFDNAFN-CTFEYISDSFSL
AAS10463.1 NNSTNVVIRACNFELCDNPFFVVSKPMGTRTHTM------IFDNAFN-CTFEYISDAFSL
AAP13441.1 NNSTNVVIRACNFELCDNPFFAVSKPMGTQTHTM------IFDNAFN-CTFEYISDAFSL
AAP13567.1 NNSTNVVIRACNFELCDNPFFAVSKPMGTQTHTM------IFDNAFN-CTFEYISDAFSL
QHD43416.1 NNATNVVIKVCEFQFCNDPFLGVYY--HKNNKSWMESEFRVYSSANN-CTFEYVSQPFLM
AVP78031.1 NNATNVIIKVCNFDFCYDP-YLSGY--YHNNKTWSIREFAVYSSYAN-CTFEYVSKSFML
ABD75323.1 NNSTHIIIRVCYFNLCKDPMYTVSA--GTQKSSW------VYQSAFN-CTYDRVEKSFQL
QDF43835.1 NNSTHIIIRVCYFNLCKEPMYAISN--EQHYKSW------VYQNAYN-CTYDRVEQSFQL
ABD75332.1 NNSTHIIIRVCNFNLCKEPMYTVSK--GTQQSSW------VYQSAFN-CTYDRVEKSFQL
QDF43820.1 NNSTHIIIRVCNFNLCKEPMYTVSR--GTQQSSW------VYQSAFN-CTYDRVERSFQL
AAZ67052.1 NNSTHIIIRVCNFNLCKEPMYTVSR--GAQQSSW------VYQSAFN-CTYDRVEKSFQL
AFS88936.1 PDGCGTLLRAFYCIL--EPRSGNHCPAGNSYTSF-----ATYHTPATDCSDGNYNRNASL
YP_0010399 PDGCGTVLHAFYCIL--KPRTVNRCPSGTGYVSY-----FIYETVHNDCQ-STINRNASL
:. ::.. : .* : : . . * . :
QDF43825.1 DIGEKPGNFKDLREFVFRNKDG--------FLHVYSGYQPISAASGLPTGF--NALKPIF
AGZ48818.1 DLGEKPGNFKDLREFVFRNKDG--------FLHVYSGYQPISAASGLPTGF--NALKPIF
ALK02457.1 DVAEKSGNFKHLREFVFKNKDG--------FLYVYKGYQPIDVVRDLPSGF--NILKPIF
AAS10463.1 DVSEKSGNFKHLREFVFKNKDG--------FLYVYKGYQPIDVVRDLPSGF--NTLKPIF
AAP13441.1 DVSEKSGNFKHLREFVFKNKDG--------FLYVYKGYQPIDVVRDLPSGF--NTLKPIF
AAP13567.1 DVSEKSGNFKHLREFVFKNKDG--------FLYVYKGYQPIDVVRDLPSGF--NTLKPIF
QHD43416.1 DLEGKQGNFKNLREFVFKNIDG--------YFKIYSKHTPINLVRDLPQGF--SALEPLV
AVP78031.1 NISGNGGLFNTLREFVFRNVDG--------HFKIYSKFTPVNLNRGLPTGL--SVLQPLV
ABD75323.1 DTSPKTGNFTDLREFVFKNRDG--------FFTAYQTYTPVNLLRGLPSGL--SVLKPIL
QDF43835.1 DTAPQTGNFKDLREYVFKNKDG--------FLSVYNAYSPIDIPRGLPVGF--SVLKPIL
ABD75332.1 DTAPKTGNFKDLREYVFKNKGG--------FLRVYQTYTAVNLPRGFPAGF--SVLRPIL
QDF43820.1 DTAPKTGNFKDLREYVFKNRDG--------FLSVYQTYTAVNLPRGLPIGF--SVLRPIL
AAZ67052.1 DTAPKTGNFKDLREYVFKNRDG--------FLSVYQTYTAVNLPRGLPIGF--SVLRPIL
AFS88936.1 NSFKE---YFNLRNCTFMYTYNITEDEILEWFGITQTAQGVHLFSSRYVDLYGGNMFQFA
YP_0010399 NSFK---SFFDLVNCTFFNSWDITADETKEWFGITQDTQGVHLYSSRKGDLYGGNMFRFA
: : * : .* . : . : . .: . : :
QDF43825.1 KLPLGINITNFRTLLTAF------PPNPGYWGTSAAAYFVGYLKPTTFMLKYDENGTITD
AGZ48818.1 KLPLGINITNFRTLLTAF------PPRPDYWGTSAAAYFVGYLKPTTFMLKYDENGTITD
ALK02457.1 KLPLGINITNFRAILTAF------LPAQDTWGTSAAAYFVGYLKPATFMLKYDENGTITD
AAS10463.1 KLPLGINITNFRAILTAF------SPAQDTWGTSAAAYFVGYLKPTTFMLKYDENGTITD
AAP13441.1 KLPLGINITNFRAILTAF------SPAQDIWGTSAAAYFVGYLKPTTFMLKYDENGTITD
AAP13567.1 KLPLGINITNFRAILTAF------SPAQDTWGTSAAAYFVGYLKPTTFMLKYDENGTITD
QHD43416.1 DLPIGINITRFQTLLALHRSYLTPGDSSSGWTAGAAAYYVGYLQPRTFLLKYNENGTITD
AVP78031.1 ELPVSINITKFRTLLTIHRGD---PMPNNGWTAFSAAYFVGYLKPRTFMLKYNENGTITD
ABD75323.1 KLPFGINITSFRVVMAMF------SKTTSNYVPESAAYYVGNLKQSTFMLSFNQNGTIVD
QDF43835.1 KLPIGINITSFKVVMSMF------SRTTSNFLPEVAAYFVGNLKYSTFMLNFNENGTITD
ABD75332.1 KLPFGINITSYRVVMTMF------SQFNSNFLPESAAYYVGNLKYTTFMLSFNENGTITD
QDF43820.1 KLPFGINITSYRVVMAMF------SQTTSNFLPESAAYYVGNLKYTTFMLRFNENGTITD
AAZ67052.1 KLPFGINITSYRVVMAMF------SQTTSNFLPESAAYYVGNLKYTTFMLSFNENGTITN
AFS88936.1 TLPVYDTIKYYSIIPHSIRSI---QSDRKAW----AAFYVYKLQPLTFLLDFSVDGYIRR
YP_0010399 TLPVYEGIKYYTVIPRSFRSK---ANKREAW----AAFYVYKLHQLTYLLDFSVDGYIRR
**. *. : : : **::* *: *::* :. :* *
QDF43825.1 AVDCSQNPLAELKCSVKSFEIDKGIYQTSNFRVAPSKEVVRFPNITNLCPFGEVFNATTF
AGZ48818.1 AVDCSQNPLAELKCSVKSFEIDKGIYQTSNFRVAPSKEVVRFPNITNLCPFGEVFNATTF
ALK02457.1 AVDCSQNPLAELKCSVKSFEIDKGIYQTSNFRVAPSKEVVRFPNITNLCPFGEVFNATTF
AAS10463.1 AVDCSQNPLAELKCSVKSFEIDKGIYQTSNFRVVPSGDVVRFPNITNLCPFGEVFNATKF
AAP13441.1 AVDCSQNPLAELKCSVKSFEIDKGIYQTSNFRVVPSGDVVRFPNITNLCPFGEVFNATKF
AAP13567.1 AVDCSQNPLAELKCSVKSFEIDKGIYQTSNFRVVPSGDVVRFPNITNLCPFGEVFNATKF
QHD43416.1 AVDCALDPLSETKCTLKSFTVEKGIYQTSNFRVQPTESIVRFPNITNLCPFGEVFNATRF
AVP78031.1 AVDCALDPLSETKCTLKSLTVQKGIYQTSNFRVQPTQSVVRFPNITNVCPFHKVFNATRF
ABD75323.1 AVDCSQDPLAELKCTTKSFNVSKGIYQTSNFRVSPVTEVVRFPNITNLCPFDKVFNATRF
QDF43835.1 AIDCAQNPLSELKCTIKNFNVSKGIYQTSNFRVSPTHEVIRFPNITNRCPFDKVFNASRF
ABD75332.1 AVDCSQNPLAELKCTIKNFNVSKGIYQTSNFRVTPTQEVVRFPNITNRCPFDKVFNASRF
QDF43820.1 AIDCAQNPLAELKCTIKNFNVSKGIYQTSNFRVSPTQEVVRFPNITNRCPFDKVFNASRF
AAZ67052.1 AIDCAQNPLAELKCTIKNFNVSKGIYQTSNFRVSPTQEVIRFPNITNRCPFDKVFNATRF
AFS88936.1 AIDCGFNDLSQLHCSYESFDVESGVYSVSSFEAKPSGSVVEQAEGVE-CDFSPLLSGTP-
YP_0010399 AIDCGHDDLSQLHCSYTSFEVDTGVYSVSSYEASATGTFIEQPNATE-CDFSPMLTGVA-
*:**. : *:: :*: .: :..*:*..*.: . . .: .: .: * * ::..
QDF43825.1 PSVYAWERKRISNCVADYSVLYNSTSFSTFKCYGVSATKLNDLCFSNVYADSFVVKGDDV
AGZ48818.1 PSVYAWERKRISNCVADYSVLYNSTSFSTFKCYGVSATKLNDLCFSNVYADSFVVKGDDV
ALK02457.1 PSVYAWERKRISNCVADYSVLYNSTSFSTFKCYGVSATKLNDLCFSNVYADSFVVKGDDV
AAS10463.1 PSVYAWERKRISNCVADYSVLYNSTSFSTFKCYGVSATKLNDLCFSNVYADSFVVKGDDV
AAP13441.1 PSVYAWERKKISNCVADYSVLYNSTFFSTFKCYGVSATKLNDLCFSNVYADSFVVKGDDV
AAP13567.1 PSVYAWERKKISNCVADYSVLYNSTFFSTFKCYGVSATKLNDLCFSNVYADSFVVKGDDV
QHD43416.1 ASVYAWNRKRISNCVADYSVLYNSASFSTFKCYGVSPTKLNDLCFTNVYADSFVIRGDEV
AVP78031.1 PSVYAWERTKISDCIADYTVFYNSTSFSTFKCYGVSPSKLIDLCFTSVYADTFLIRFSEV
ABD75323.1 PSVYAWERTKISDCVADYTVFYNSTSFSTFNCYGVSPSKLIDLCFTSVYADTFLIRFSEV
QDF43835.1 PNVYAWERTKISDCVADYTVLYNSTSFSTFKCYGVSPSKLIDLCFTSVYADTFLIRSSEV
ABD75332.1 PNVYAWERTKISDCVADYTVLYNSTSFSTFKCYGVSPSKLIDLCFTSVYADTFLIRSSEV
QDF43820.1 PNVYAWERTKISDCVADYTVLYNSTSFSTFKCYGVSPSKLIDLCFTSVYADTFLIRSSEV
AAZ67052.1 PNVYAWERTKISDCVADYTVLYNSTSFSTFKCYGVSPSKLIDLCFTSVYADTFLIRSSEV
AFS88936.1 PQVYNFKRLVFTNCNYNLTKLLSLFSVNDFTCSQISPAAIASNCYSSLILDYFSYPLSMK
YP_0010399 PQVYNFKRLVFSNCNYNLTKLLSLFAVDEFSCNGISPDSIARGCYSTLTVDYFAYPLSMK
..** ::* :::* : : : . .. *.* :*. : *::.: * * .
QDF43825.1 RQIAPGQTGVIADYNYKLPDDFMGC-VLAWNTRNIDATSTGNYNYKYRSLRHGKLRPFER
AGZ48818.1 RQIAPGQTGVIADYNYKLPDDFTGC-VLAWNTRNIDATQTGNYNYKYRSLRHGKLRPFER
ALK02457.1 RQIAPGQTGVIADYNYKLPDDFTGC-VLAWNTRNIDATQTGNYNYKYRSLRHGKLRPFER
AAS10463.1 RQIAPGQTGVIADYNYKLPDDFMGC-VLAWNTRNIDATSTGNYNYKYRYLRHGKLRPFER
AAP13441.1 RQIAPGQTGVIADYNYKLPDDFMGC-VLAWNTRNIDATSTGNYNYKYRYLRHGKLRPFER
AAP13567.1 RQIAPGQTGVIADYNYKLPDDFMGC-VLAWNTRNIDATSTGNYNYKYRYLRHGKLRPFER
QHD43416.1 RQIAPGQTGKIADYNYKLPDDFTGC-VIAWNSNNLDSKVGGNYNYLYRLFRKSNLKPFER
AVP78031.1 RQVAPGQTGVIADYNYKLPDDFTGC-VIAWNTAKQD---VGNYF--YRSHRSTKLKPFER
ABD75323.1 RQVAPGQTGVIADYNYKLPDDFTGC-VIAWNTAKQD---VGSYF--YRSHRSSKLKPFER
QDF43835.1 RQVAPGETGVIADYNYKLPDDFTGC-VIAWNTAKQD---QGQYY--YRSSRKTKLKPFER
ABD75332.1 RQVAPGETGVIADYNYKLPDDFTGC-VIAWNTAQQD---QGQYY--YRSYRKEKLKPFER
QDF43820.1 RQVAPGETGVIADYNYKLPDDFTGC-VIAWNTAKQD---TGHYY--YRSHRKTKLKPFER
AAZ67052.1 RQVAPGETGVIADYNYKLPDDFTGC-VIAWNTAKQD---QGQYY--YRSHRKTKLKPFER
AFS88936.1 SDLSVSSAGPISQFNYKQSFSNPTC-LILATVPHNLTTITKPLKYSYINKCSRLLSDDRT
YP_0010399 SYIRPGSAGNIPLYNYKQSFANPTCRVMASVLANVTITKPHAYG--YIS-KCSRLTGANQ
: ..:* *. :*** . * :: : * *
QDF43825.1 DISNVPFSPDGKPCTPP-AF-NCYW-----------PLNDYGFFTTNGIGYQPYRVVVLS
AGZ48818.1 DISNVPFSPDGKPCTPP-AF-NCYW-----------PLNDYGFYITNGIGYQPYRVVVLS
ALK02457.1 DISNVPFSPDGKPCTPP-AF-NCYW-----------PLNDYGFYITNGIGYQPYRVVVLS
AAS10463.1 DISNVPFSPDGKPCTPP-AP-NCYW-----------PLNGYGFYTTSGIGYQPYRVVVLS
AAP13441.1 DISNVPFSPDGKPCTPP-AL-NCYW-----------PLNDYGFYTTTGIGYQPYRVVVLS
AAP13567.1 DISNVPFSPDGKPCTPP-AL-NCYW-----------PLNDYGFYTTTGIGYQPYRVVVLS
QHD43416.1 DISTEIYQAGSTPCNGVEGF-NCYF-----------PLQSYGFQPTNGVGYQPYRVVVLS
AVP78031.1 DLSSDE---------------NGVR-----------TLSTYDFNPNVPLEYQATRVVVLS
ABD75323.1 DLSSEE---------------NGVR-----------TLSTYDFNQNVPLEYQATRVVVLS
QDF43835.1 DLTSDE---------------NGVR-----------TLSTYDFYPNVPIEYQATRVVVLS
ABD75332.1 DLSSDE---------------NGVY-----------TLSTYDFYPSIPVEYQATRVVVLS
QDF43820.1 DLSSDDG--------------NGVY-----------TLSTYDFNPNVPVAYQATRVVVLS
AAZ67052.1 DLSSDE---------------NGVR-----------TLSTYDFYPSVPVAYQATRVVVLS
AFS88936.1 EVPQLVNANQYSPCVSI-VP-STVWEDGDYYRKQLSPLEGGGWLVASGSTVAMTEQLQMG
YP_0010399 DVETPLYINPGEYSICRDFSPGGFSEDGQVFKRTLTQFEGGGLLIGVGTRVPMTDNLQMS
:: . :. . : :.
QDF43825.1 FELL----NAPATVC-----GPKLSTDLIKNQCVNFNFNGLTGTGVLTPSSKRFQPFQQF
AGZ48818.1 FELL----NAPATVC-----GPKLSTDLIKNQCVNFNFNGLTGTGVLTPSSKRFQPFQQF
ALK02457.1 FELL----NAPATVC-----GPKLSTDLIKNQCVNFNFNGLTGTGVLTPSSKRFQPFQQF
AAS10463.1 FELL----NAPATVC-----GPKLSTDLIKNQCVNFNFNGLTGTGVLTPSSKRFQPFQQF
AAP13441.1 FELL----NAPATVC-----GPKLSTDLIKNQCVNFNFNGLTGTGVLTPSSKRFQPFQQF
AAP13567.1 FELL----NAPATVC-----GPKLSTDLIKNQCVNFNFNGLTGTGVLTPSSKRFQPFQQF
QHD43416.1 FELL----HAPATVC-----GPKKSTNLVKNKCVNFNFNGLTGTGVLTESNKKFLPFQQF
AVP78031.1 FELL----NAPATVC-----GPKLSTQLVKNQCVNFNFNGLKGTGVLTDSSKRFQSFQQF
ABD75323.1 FELL----NAPATVC-----GPKLSTSLVKNQCVNFNFNGFKGTGVLTDSSKTFQSFQQF
QDF43835.1 FELL----NAPATVC-----GPKLSTGLVKNQCVNFNFNGLRGTGVLTDSSKRFQSFQQF
ABD75332.1 FELL----NAPATVC-----GPKLSTQLVKNQCVNFNFNGLRGTGVLTTSSKRFQSFQQF
QDF43820.1 FELL----NAPATVC-----GPKLSTQLVKNQCVNFNFNGLKGTGVLTDSSKRFQSFQQF
AAZ67052.1 FELL----NAPATVC-----GPKLSTQLVKNQCVNFNFNGLKGTGVLTESSKRFQSFQQF
AFS88936.1 FGITVQYGTDTNSVCPKLEFANDTKIASQLGNCVEYSLYGVSGRGVFQNCTAVGVRQQRF
YP_0010399 FIISVQYGTGTDSVCPMLDLGDSLTITNRLGKCVDYSLYGVTGRGVFQNCTAVGVKQQRF
* : . :** . . . .:**::.: *. * **: .. *.*
QDF43825.1 GRDVSD-FTDSVRDPKTSEILDISPCSFGGVSVITPGTNTSSEVAVLYQDVNCTDVPVAI
AGZ48818.1 GRDVSD-FTDSVRDPKTSEILDISPCSFGGVSVITPGTNTSSEVAVLYQDVNCTDVPVAI
ALK02457.1 GRDVLD-FTDSVRDPKTSEILDISPCSFGGVSVITPGTNTSSEVAVLYQDVNCTDVPVAI
AAS10463.1 GRDVSD-FTDSVRDPKTSEILDISPCSFGGVSVITPGTNASSEVAVLYQDVNCTDVSTLI
AAP13441.1 GRDVSD-FTDSVRDPKTSEILDISPCSFGGVSVITPGTNASSEVAVLYQDVNCTDVSTAI
AAP13567.1 GRDVSD-FTDSVRDPKTSEILDISPCSFGGVSVITPGTNASSEVAVLYQDVNCTDVSTAI
QHD43416.1 GRDIAD-TTDAVRDPQTLEILDITPCSFGGVSVITPGTNTSNQVAVLYQDVNCTEVPVAI
AVP78031.1 GKDASD-FIDSVRDPQTLEILDITPCSFGGVSVITPGTNTSLEVAVLYQDVNCTDVPTTI
ABD75323.1 GRDASD-FTDSVRDPQTLRILDISPCSFGGVSVITPGTNTSSAVAVLYQDVNCTDVPRTI
QDF43835.1 GRDTSD-FTDSVRDPQTLEILDITPCSFGGVSVITPGTNASSEVAVLYQDVNCTDVPTAI
ABD75332.1 GRDTSD-FTDSVRDPQTLEILDISPCSFGGVSVITPGTNASSEVAVLYQDVNCTDVPTSI
QDF43820.1 GRDTSD-FTDSVRDPQTLEILDITPCSFGGVSVITPGTNASSEVAVLYQDVNCTDVPTAI
AAZ67052.1 GRDTSD-FTDSVRDPQTLEILDISPCSFGGVSVITPGTNASSEVAVLYQDVNCTDVPAAI
AFS88936.1 VYDAYQNLVGYYSDDGNYYCLR--ACVSVPVSVIY--DKETKTHATLFGSVACEHISSTM
YP_0010399 VYDSFDNLVGYYSDDGNYYCVR--PCVSVPVSVIY--DKSTNLHATLFGSVACEHVTTMM
* : . * . : .* **** : : *.*: .* * :. :
QDF43825.1 --HADQLTPAWRIYSTGNNVFQTQAGCLIGAEHVDTSY---ECDIPIGAGICASYHTVSS
AGZ48818.1 --HADQLTPSWRVYSTGNNVFQTQAGCLIGAEHVDTSY---ECDIPIGAGICASYHTVSS
ALK02457.1 --HADQLTPSWRVYSTGNNVFQTQAGCLIGAEHVDTSY---ECDIPIGAGICASYHTVSS
AAS10463.1 --HAEQLTPAWRIYSTGNNVFQTQAGCLIGAEHVDTSY---ECDIPIGAGICASYHTVSS
AAP13441.1 --HADQLTPAWRIYSTGNNVFQTQAGCLIGAEHVDTSY---ECDIPIGAGICASYHTVSL
AAP13567.1 --HADQLTPAWRIYSTGNNVFQTQAGCLIGAEHVDTSY---ECDIPIGAGICASYHTVSL
QHD43416.1 --HADQLTPTWRVYSTGSNVFQTRAGCLIGAEHVNNSY---ECDIPIGAGICASYQTQTN
AVP78031.1 --HADQLTPAWRIYATGTNVFQTQAGCLIGAEHVNASY---ECDIPIGAGICASYHTASI
ABD75323.1 --QADQLAPSWRVYTTGPYVFQTQAGCLIGAEHVNASY---QCDIPIGAGICASYHTASH
QDF43835.1 --RADQLTPAWRVYSTGINVFQTQAGCLIGAEHVNASY---ECDIPIGAGICASYHTAST
ABD75332.1 --HADQLTPAWRVYSTGVNVFQTQAGCLIGAEHVNASY---ECDIPIGAGICASYHTASV
QDF43820.1 --RADQLTPAWRVYSTGVNVFQTQAGCLIGAEHVNASY---ECDIPIGAGICASYHTAST
AAZ67052.1 --HADQLTPAWRVYSTGTNVFQTQAGCLIGAEHVNASY---ECDIPIGAGICASYHTAST
AFS88936.1 SQYSRSTRSMLKRRDSTYGPLQTPVGCVLGL--VNSSLFVEDCKLPLGQSLCALPDTPST
YP_0010399 S-QFSRLTQSNLRRRDSNIPLQTAVGCVIGLS--NNSLVVSDCKLPLGQSLCAV-PPVST
:** .**::* : * :*.:*:* .:** . :
QDF43825.1 ----LRSTS----QKSI--------VAYTMSLGADSSIAYSNNTIAIPTNFSISITTEVM
AGZ48818.1 ----LRSTS----QKSI--------VAYTMSLGADSSIAYSNNTIAIPTNFSISITTEVM
ALK02457.1 ----LRSTS----QKSI--------VAYTMSLGADSSIAYSNNTIAIPTNFSISITTEVM
AAS10463.1 ----LRSTS----QKSI--------VAYTMSLGADSSIAYSNNTIAIPTNFSISITTEVM
AAP13441.1 ----LRSTS----QKSI--------VAYTMSLGADSSIAYSNNTIAIPTNFSISITTEVM
AAP13567.1 ----LRSTS----QKSI--------VAYTMSLGADSSIAYSNNTIAIPTNFSISITTEVM
QHD43416.1 SPRRARSVA----SQSI--------IAYTMSLGAENSVAYSNNSIAIPTNFTISVTTEIL
AVP78031.1 ----LRSTS----QKAI--------VAYTMSLGAENSIAYANNSIAIPTNFSISVTTEVM
ABD75323.1 ----LRSTG----QKSI--------VAYTMSLGAENSVAYANNSIAIPTNFSISVTTEVM
QDF43835.1 ----LRSVG----QKSI--------VAYTMSLGAENSIAYANNSIAIPTNFSISVTTEVM
ABD75332.1 ----LRSTG----QKSI--------VAYTMSLGAENSIAYANNSIAIPTNFSISVTTEVM
QDF43820.1 ----LRSVG----QKSI--------VAYTMSLGAENSIAYANNSIAIPTNFSISVTTEVM
AAZ67052.1 ----LRSVG----QKSI--------VAYTMSLGAENSIAYANNSIAIPTNFSISVTTEVM
AFS88936.1 ----LTPRS----VRSVPGEMRLASIAFNHPIQVDQ-LNSSYFKLSIPTNFSFGVTQEYI
YP_0010399 ----FRSYSASQFQLAV--------LNYTSPIVV-TPINSSGFTAAIPTNFSFSVTQEYI
. . :: : :. .: . : : . :*****::.:* * :
QDF43825.1 PVSMAKTSVDCNMYICGDSTECANLLLQYGSFCTQLNRALSGIAVEQDRNTREVFAQVKQ
AGZ48818.1 PVSMAKTSVDCNMYICGDSTECANLLLQYGSFCTQLNRALSGIAVEQDRNTREVFAQVKQ
ALK02457.1 PVSMAKTSVDCNMYICGDSTECANLLLQYGSFCTQLNRALSGIAVEQDRNTREVFAQVKQ
AAS10463.1 PVSMAKTSVDCNMYICGDSTECANLLLQYGSFCRQLNRALSGIAAEQDRNTREVFVQVKQ
AAP13441.1 PVSMAKTSVDCNMYICGDSTECANLLLQYGSFCTQLNRALSGIAAEQDRNTREVFAQVKQ
AAP13567.1 PVSMAKTSVDCNMYICGDSTECANLLLQYGSFCTQLNRALSGIAAEQDRNTREVFAQVKQ
QHD43416.1 PVSMTKTSVDCTMYICGDSTECSNLLLQYGSFCTQLNRALTGIAVEQDKNTQEVFAQVKQ
AVP78031.1 PVSMAKTSVDCTMYICGDSIECSNLLLQYGSFCTQLNRALSGIAIEQDKNTQEVFAQVKQ
ABD75323.1 PVSMAKTSVDCTMYICGDSLECSNLLLQYGSFCTQLNRALSGIAVEQDKNTQEVFAQVKQ
QDF43835.1 PVSMSKTSVDCTMYICGDSQECSNLLLQYGSFCTQLNRALTGIAIEQDKNTQEVFAQVKQ
ABD75332.1 PVSIAKTSVDCTMYICGDSLECSNLLLQYGSFCTQLNRALTGIAIEQDKNTQEVFAQVKQ
QDF43820.1 PVSMAKTSVDCTMYICGDSQECSNLLLQYGSFCTQLNRALTGVALEQDKNTQEVFAQVKQ
AAZ67052.1 PVSMAKTSVDCTMYICGDSLECSNLLLQYGSFCTQLNRALSGIAIEQDKNTQEVFAQVKQ
AFS88936.1 QTTIQKVTVDCKQYVCNGFQKCEQLLREYGQFCSKINQALHGANLRQDDSVRNLFASVKS
YP_0010399 ETSIQKVTVDCKQYVCNGFTRCEKLLVEYGQFCSKINQALHGANLRQDESVYSLYSNIKT
.:: *.:***. *:*.. * :** :**.** ::*.** * ** .. .:: .:*
QDF43825.1 MYKTPTLKD-FGG-FNFSQILPDPLKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
AGZ48818.1 MYKTPTLKD-FGG-FNFSQILPDPLKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
ALK02457.1 MYKTPTLKD-FGG-FNFSQILPDPLKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
AAS10463.1 MYKTPTLKD-FGG-FNFSQILPDPLKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
AAP13441.1 MYKTPTLKY-FGG-FNFSQILPDPLKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
AAP13567.1 MYKTPTLKY-FGG-FNFSQILPDPLKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
QHD43416.1 IYKTPPIKD-FGG-FNFSQILPDPSKPSKRSF---IEDLLFNKVTLADAGFIKQYGDCL-
AVP78031.1 IYKTPPIKD-FGG-FNFSQILPDPSKPSKRSF---IEDLLFNKVTLADAGFIKQYGDCL-
ABD75323.1 MYKTPTIRD-FGG-FNFSQILPDPLKPTKRSF---IEDLLYNKVTLADAGFMKQYADCL-
QDF43835.1 MYKTPAIKD-FGG-FNFSQILPDPSKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
ABD75332.1 MYKTPAIKD-FGG-FNFSQILPDPSKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
QDF43820.1 MYKTPAIKD-FGG-FNFSQILPDPSKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
AAZ67052.1 MYKTPAIKD-FGG-FNFSQILPDPSKPTKRSF---IEDLLFNKVTLADAGFMKQYGECL-
AFS88936.1 SQSSPIIPG-FGGDFNLTLLEPVSISTGSRSARSAIEDLLFDKVTIADPGYMQGYDDCMQ
YP_0010399 T-STQTLEYGLNGDFNLTLLQVPQIGGSSSSYRSAIEDLLFDKVTIADPGYMQGYDDCMK
.: : :.* **:: : . * *****::***:**.*::: * :*:
QDF43825.1 -GDINARDLICAQKFNGLTVLPPLLTDDMIAAYTAALVSGTATAGWTFGAGAALQIPFAM
AGZ48818.1 -GDINARDLICAQKFNGLTVLPPLLTDDMIAAYTAALVSGTATAGWTFGAGAALQIPFAM
ALK02457.1 -GDINARDLICAQKFNGLTVLPPLLTDDMIAAYTAALVSGTATAGWTFGAGAALQIPFAM
AAS10463.1 -GDINARDLICAQKFNGLTVLPPLLTDDMIAAYTAALVSGTATAGWTFGAGAALQIPFAM
AAP13441.1 -GDINARDLICAQKFNGLTVLPPLLTDDMIAAYTAALVSGTATAGWTFGAGAALQIPFAM
AAP13567.1 -GDINARDLICAQKFNGLTVLPPLLTDDMIAAYTAALVSGTATAGWTFGAGAALQIPFAM
QHD43416.1 -GDIAARDLICAQKFNGLTVLPPLLTDEMIAQYTSALLAGTITSGWTFGAGAALQIPFAM
AVP78031.1 -GGISARDLICAQKFNGLTVLPPLLTDEMIAAYTAALISGTATAGWTFGAGAALQIPFAM
ABD75323.1 -GGINARDLICAQKFNGLTVLPPLLTDDMIAAYTAALISGTATAGWTFGAGAALQIPFAM
QDF43835.1 -GDINARDLICAQKFNGLTVLPPLLTDDMIAAYTAALVSGTATAGWTFGAGAALQIPFAM
ABD75332.1 -GDISARDLICAQKFNGLTVLPPLLTDEMIAAYTAALVSGTATAGWTFGAGSALQIPFAM
QDF43820.1 -GDINARDLICAQKFNGLTVLPPLLTDDMIAAYTAALVSGTATAGWTFGAGAALQIPFAM
AAZ67052.1 -GDISARDLICAQKFNGLTVLPPLLTDEMIAAYTAALVSGTATAGWTFGAGSALQIPFAM
AFS88936.1 QGPASARDLICAQYVAGYKVLPPLMDVNMEAAYTSSLLGSIAGVGWTAGLSSFAAIPFAQ
YP_0010399 QGPQSARDLICAQYVSGYKVLPPLYDPNMEAAYTSSLLGSIAGAGWTAGLSSFAAIPFAQ
* ******** . * .***** :* * **::*:.. *** * .: ****
QDF43825.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
AGZ48818.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
ALK02457.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
AAS10463.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
AAP13441.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
AAP13567.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
QHD43416.1 QMAYRFNGIGVTQNVLYENQKLIANQFNSAIGKIQDSLSSTASALGKLQDVVNQNAQALN
AVP78031.1 QMAYRFNGIGVTQNVLYENQKLIANQFNSAIGKIQESLTSTASALGKLQDVVNQNAQALN
ABD75323.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAITQIQESLTTTSTALGKLQDVVNQNAQALN
QDF43835.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
ABD75332.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
QDF43820.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
AAZ67052.1 QMAYRFNGIGVTQNVLYENQKQIANQFNKAISQIQESLTTTSTALGKLQDVVNQNAQALN
AFS88936.1 SIFYRLNGVGITQQVLSENQKLIANKFNQALGAMQTGFTTTNEAFQKVQDAVNNNAQALS
YP_0010399 SMFYRLNGVGITQQVLSENQKLIANKFNQALGAMQTGFTTSNLAFSKVQDAVNANAQALS
.: **:**:*:**:** **** ***:**.*: :* .:::: *: *:**.** *****.
QDF43825.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
AGZ48818.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
ALK02457.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
AAS10463.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
AAP13441.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
AAP13567.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
QHD43416.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
AVP78031.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
ABD75323.1 TLVKQLSSNFGAISSALNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
QDF43835.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
ABD75332.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
QDF43820.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
AAZ67052.1 TLVKQLSSNFGAISSVLNDILSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRA
AFS88936.1 KLASELSNTFGAISASIGDIIQRLDVLEQDAQIDRLINGRLTTLNAFVAQQLVRSESAAL
YP_0010399 KLASELSNTFGAISSSISDILARLDTVEQDAQIDRLINGRLISLNAFVSQQLVRSETAAR
.*..:**..*****: :.**: *** :* :.******.*** :*:::*:***:*:
QDF43825.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQAAPHGVVFLHVTYVPSQERNFTTAPA
AGZ48818.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQAAPHGVVFLHVTYVPSQERNFTTAPA
ALK02457.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQAAPHGVVFLHVTYVPSQERNFTTAPA
AAS10463.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQAAPHGVVFLHVTYVPSQERNFTTAPA
AAP13441.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQAAPHGVVFLHVTYVPSQERNFTTAPA
AAP13567.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQAAPHGVVFLHVTYVPSQERNFTTAPA
QHD43416.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQSAPHGVVFLHVTYVPAQEKNFTTAPA
AVP78031.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQSAPHGVVFLHVTYIPSQEKNFTTAPA
ABD75323.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQSAPHGVVFLHVTYVPSQEKNFTTAPA
QDF43835.1 SANLAATKMSECVLGQSKRVDFCGRGYHLMSFPQAAPHGVVFLHVTYVPSQEKNFTTAPA
ABD75332.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQAAPHGVVFLHVTYVPSQERNFTTAPA
QDF43820.1 SANLAATKMSECVLGQSKRVDFCGRGYHLMSFPQAAPHGVVFLHVTYVPSQEKNFTTAPA
AAZ67052.1 SANLAATKMSECVLGQSKRVDFCGKGYHLMSFPQAAPHGVVFLHVTYVPSQERNFTTAPA
AFS88936.1 SAQLAKDKVNECVKAQSKRSGFCGQGTHIVSFVVNAPNGLYFMHVGYYPSNHIEVVSAYG
YP_0010399 SAQLASDKVNECVKSQSKRNGFCGSGTHIVSFVVNAPNGFYFFHVGYVPTNYTNVTAAYG
**:** *:.*** .**** .*** * *::** **:*. *:** * *:: :..:* .
QDF43825.1 ICHEGK---AYFPREGVFVFNGTS-------WFITQRNFFSPQIITTDNT-FVSGSCDVV
AGZ48818.1 ICHEGK---AYFPREGVFVFNGTS-------WFITQRNFFSPQIITTDNT-FVSGSCDVV
ALK02457.1 ICHEGK---AYFPREGVFVFNGTS-------WFITQRNFFSPQIITTDNT-FVSGSCDVV
AAS10463.1 ICHEGK---AYFPREGVFVFNGTS-------WFITQRNFFSPQIITTDNT-FVSGNCDVV
AAP13441.1 ICHEGK---AYFPREGVFVFNGTS-------WFITQRNFFSPQIITTDNT-FVSGNCDVV
AAP13567.1 ICHEGK---AYFPREGVFVFNGTS-------WFITQRNFFSPQIITTDNT-FVSGNCDVV
QHD43416.1 ICHDGK---AHFPREGVFVSNGTH-------WFVTQRNFYEPQIITTDNT-FVSGNCDVV
AVP78031.1 ICHEGK---AHFPREGVFVSNGTH-------WFVTQRNFYEPKIITTDNT-FVSGNCDVV
ABD75323.1 ICHEGK---AYFPREGVFVSNGSS-------WFITQRNFYSPQIITTDNT-FVAGSCDVV
QDF43835.1 ICHEGK---AYFPREGVFVSNGTS-------WFITQRNFYSPQIITTDNT-FVAGSCDVV
ABD75332.1 ICHEGK---AYFPREGVFVSNGTS-------WFITQRNFYSPQIITTDNT-FVAGNCDVV
QDF43820.1 ICHEGK---AYFPREGVFVSNGTF-------WFITQRNFYSPQIITTDNT-FVAGNCDVV
AAZ67052.1 ICHEGK---AYFPREGVFVSNGTS-------WFITQRNFYSPQIITTDNT-FVAGSCDVV
AFS88936.1 LCDAANPTNCIAPVNGYFIKTNNT--RIVDEWSYTGSSFYAPEPITSLNTKYVA--PQVT
YP_0010399 LCNNNNPPLCIAPIDGYFITNQTTTYSVDTEWYYTGSSFYKPEPITQANSRYVS--SDVK
:* : . * :* *: . . * * .*: *: ** *: :*: :*
QDF43825.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRL
AGZ48818.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEINRL
ALK02457.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRL
AAS10463.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQEEIDRL
AAP13441.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRL
AAP13567.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRL
QHD43416.1 IGIVNNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRL
AVP78031.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDIDLGDISGINASVVNIQKEIDRL
ABD75323.1 IGIINNTVYDPL---QPELDSFKQELDKYFKNHTSPDVDLGDISGINASVVDIQKEIDRL
QDF43835.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRL
ABD75332.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRL
QDF43820.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRL
AAZ67052.1 IGIINNTVYDPL---QPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRL
AFS88936.1 YQNISTNLPPPLLGNSTGID-FQDELDEFFKNVSTSIPNFGSLTQINTTLLDLTYEMLSL
YP_0010399 FDKLENNLPPPLLENSTDVD-FKDELEEFFKNVTSHGPNFAEISKINTTLLDLSDEMAML
:...: ** .. :* *::**:::*** :: ::..:: **:::::: *: *
QDF43825.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMVTILLCCMTSCCSCLKG
AGZ48818.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMVTILLCCMTSCCSCLKG
ALK02457.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMVTILLCCMTSCCSCLKG
AAS10463.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMVTILLCCMTSCCSCLKG
AAP13441.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMVTILLCCMTSCCSCLKG
AAP13567.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMVTILLCCMTSCCSCLKG
QHD43416.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYIWLGFIAGLIAIVMVTIMLCCMTSCCSCLKG
AVP78031.1 NEVARNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMVTILLCCMTSCCSCLKG
ABD75323.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLVGLFMAIILLCYFTSCCSCCKG
QDF43835.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMATILLCCMTSCCSCLKG
ABD75332.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMVTILLCCMTSCCSCLKG
QDF43820.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMATILLCCMTSCCSCLKG
AAZ67052.1 NEVAKNLNESLIDLQELGKYEQYIKWPWYVWLGFIAGLIAIVMVTILLCCMTSCCSCLKG
AFS88936.1 QQVVKALNESYIDLKELGNYTYYNKWPWYIWLGFIAGLVALALCVFFILCCTGCGTNCMG
YP_0010399 QEVVKQLNDSYIDLKELGNYTYYNKWPWYVWLGFIAGLVALLLCVFFLLCCTGCGTSCLG
::*.. **:* ***:***:* * *****:********:.: : ::: *.* : *
QDF43825.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
AGZ48818.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
ALK02457.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
AAS10463.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
AAP13441.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
AAP13567.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
QHD43416.1 CCSCGSCC-KFDEDDSEPVLKGVKLHYT
AVP78031.1 CCSCGSCC-KFDEDDSEPVLKGVKLHYT
ABD75323.1 MCSCGSCC-RFDEDDSEPVLKGVKLHYT
QDF43835.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
ABD75332.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
QDF43820.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
AAZ67052.1 ACSCGSCC-KFDEDDSEPVLKGVKLHYT
AFS88936.1 KLKCNRCCDRYEEYDLEP----HKVHVH
YP_0010399 KMKCKNCCDSYEEYDVE------KIHVH
.* ** ::* * * *:*
Phylogenetic Tree
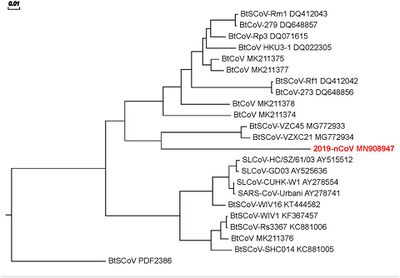
Figure 1: Phylogenetic Tree of Spike Protein Sequences. The length of each branch represents the percentage change in amino acid sequence occurring along that branch, relative to the scale bar shown at the bottom of the tree.
My Tree and Sequences
- Looking at the tree created in our class alongside the sequences, it is evident that there are some aspects of the sequence that we see in the tree and vice versa. For example, AFS88936.1 and YP_0010399 are two sequences that seem to have sequences similar to each other yet very distinct from the other ones and it is also seen in the tree, both appearing as outliers. Furthermore, QDF43820.1 and QDF43835.1 have similar sequences to each other and, as expected, also appear close to each other on the tree; they both stem from a similar node and do not have very long branches. However, QHD43416.1 and AVP78031.1 have somewhat similar sequences a few differences and, as expected as well, no not appear close to each other in the tree. While they do both branch from the same node, the length of branches for the two are different, with QHD43416.1 appearing to be on a longer branch. Lastly, AAP13441.1 and AAP13567.1 appear to have similar sequences and further appear to be located very close to each other on the tree, both stemming from the same node and not branching very far.
My sequence and Article Sequence
- Looking at my sequence alignment in comparison to the article's, a few differences are seen. Aside from the fact that there are differences in the actual sequences used and presented, we do see a greater prevalence in the amount of colons in my sequences, indicating positions that have strongly conserved residues. This may be simply due to the region analyzed but it is noteworthy. Many of the regions in the article sequence appear to have asterisks, indicating positions that have a single, fully conserved residues. The article sequences seem to be starting with either Leucine or Tyrosine while my sequences vary greatly. Furthermore, a lot of my sequences appear to have gaps in them, while the article sequences do not. A similarity between my sequence and the article sequence is that there are in fact areas that were fully conserved and some that maintained the same amino acid throughout; this is seen in my sequence with E, representing Glutamic Acid, and Y, in the article sequence, representing Tyrosine, although mine also does have a prevalence of Tyrosine as well. Further, there is a region in which the sequence has several instances of Valine and mine shows that as well, although more so with Threonine.
My Tree and Article Tree
- Initially it is clear that my tree includes two out-groups, AFS88936.1 and YP_0010399, while the Article tree only contains one, BtSCoV PDF2368. Although that was expected as those were some of the previously-described differences among the sequences. Further, my tree includes spike protein sequences while the article tree includes genomic sequences. Within my tree, AAP13441.1 and AAP13567.1 appear to be located very close to each other on the tree, both stemming from the same node and not branching very far. This is seen in the article's tree, with AY278741 and AY278554; the sequences are different because the article cites genomic sequences while my tree contains spike protein sequences.
Reproducibility
- I do not believe enough information provided by Wan et al (2020) in their paper for anyone really to reproduce their analysis; their materials and methods section was incredibly short, providing little information to anyone who may want to attempt to reproduce the results and see it for themselves. As seen in our lab, a methods section should be reasonably thorough, enough to explain to someone with experience in the field how to conduct the lab and how results were obtained. While their methods did provide information regarding softwares used, without information on how to obtained the data to input into the software and without specific information on software configuration, simply providing the name of the software does not prove to be especially useful.
Conclusion
- I believe the purpose of this exercise was met as I was able to not only analyze and discuss the article but further conduct our own analysis and create our own sequences and phylogenetic trees to analyze ourselves. We created sequence data using spike protein analysis of several samples and further went on to create a phylogenetic tree. In class we discussed our opinions of the article and even spent a significant amount of time individually presenting the figures and tables. Furthermore, we were able to strengthen our skills with regards to scientific literary analyzes through both the discussion of the article and the activity. Not only, but we were able to strengthen our skills with regard to the use of bioinformatics databases.
Acknowledgements
- I acknowledge my professor, Kam D. Dahlquist, Ph.D., with whom I met several times via zoom to discuss questions regarding the purpose and format.
- I acknowledge my TA, Annika Dinulos, whom I contacted regarding a question for one of my assignments.
- I acknowledge my lab partner, JT Correy, whom I worked with and consulted while creating my sequence alignments and tree
- I copied and modified the protocol shown on the Week 1 page.
- I copied and modified the protocol shown on the Week 4 page.
- I copied and modified sequence data and a phylogenetic tree from Phylogeny.fr
- I copied and referred to data in the article by Wan et al (2020)
"Except for what is noted above, this individual journal entry was completed by me and not copied from another source." Yaniv Maddahi (talk) 16:44, 30 September 2020 (PDT)
References
- Yushun Wan, Jian Shang, Rachel Graham, Ralph S. Baric, Fang Li Journal of Virology Mar 2020, 94 (7) e00127-20; DOI: 10.1128/JVI.00127-20
- OpenWetWare. (2020). BIOL368/F20:Week 1. Retrieved September 30, 2020, from https://openwetware.org/wiki/BIOL368/F20:Week_1
- OpenWetWare. (2020). BIOL368/F20:Week 4. Retrieved September 30, 2020, from https://openwetware.org/wiki/BIOL368/F20:Week_4
- Phylogeny.fr: Home. (2020). Retrieved September 30, 2020, from https://www.phylogeny.fr/
- NCBI GenBank. (2020). Bat SARS coronavirus Rp3, complete genome - Nucleotide. Retrieved 1 October 2020, from https://www.ncbi.nlm.nih.gov/nuccore/DQ071615
- NCBI GenBank. (2020). spike protein [Bat SARS CoV Rp3/2004] - Protein. Retrieved 1 October 2020, https://www.ncbi.nlm.nih.gov/protein/72256271
Template
Assignment Week
- Week 1 Assignment
- Week 2 Assignment
- Week 3 Assignment
- Week 4 Assignment
- Week 5 Assignment
- Week 6 Assignment
- Week 7 Assignment
- Week 8 Assignment
- Week 9 Assignment
- Week 10 Assignment
- Week 11 Assignment
- Week 12 Assignment
- Week 14 Assignment
Individual Journal Pages
- Yaniv Maddahi Journal Week 1
- Yaniv Maddahi Journal Week 2
- Yaniv Maddahi Journal Week 3
- Yaniv Maddahi Journal Week 4
- Yaniv Maddahi Journal Week 5
- Yaniv Maddahi Journal Week 6
- Yaniv Maddahi Journal Week 7
- Allosteric Database Review
- Yaniv Maddahi Journal Week 9
- Yaniv Maddahi Journal Week 10
- Yaniv Maddahi Journal Week 11
- The Mutants Research Project Week 12
- The Mutants Research Project Week 14