Week 11 Individual Journal William P Fuchs
From OpenWetWare
Jump to navigationJump to search
Question
- What are the visible protein sequence changes that correlate with a decline in CD4-T cell count and how are those changes altering the characteristic of the amino acid sequences?
Hypothesis
- Given that the amino acids of the clones of subject 10 are changing simultaneously within the V3 loop and gp120 contact surfaces which are significant in immune system interaction, there will be a relationship of those regions with the HIV population's immunogenicity.
Purpose
- Our purpose was to continue using the programs from BIOL368/F16:Week 9 to better investigate our question and hypothesis and to continue the HIV amino acid project initiated in Week 10.
Methods
- Access Biology Workbench and and upload Subject 10's clone sequences for amino acids and nucleotides and across all visits.
- Run a CLUSTALW multiple sequence alignment and generate a rooted tree.
- Calculate S and Theta for amino acid and nucleic acid
- Collect Min and Max values amino acid and nucleic acid
- Select the Multiple Sequence alignment tool and run a CLUSTALDIST test on the subject for amino acids.
- Find the 3 most divergent visits and clone sequences by resourcing the CLUSTALDIST matrix.
- Quantify the "most divergent visits" by measuring number of non-consensus amino acid changes.
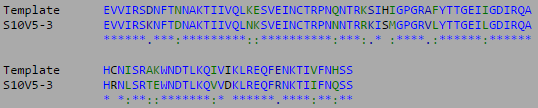
- Isolate the compared amino acid sequences to the V3 target sequence provided by the Huang et al. (2005) data set.
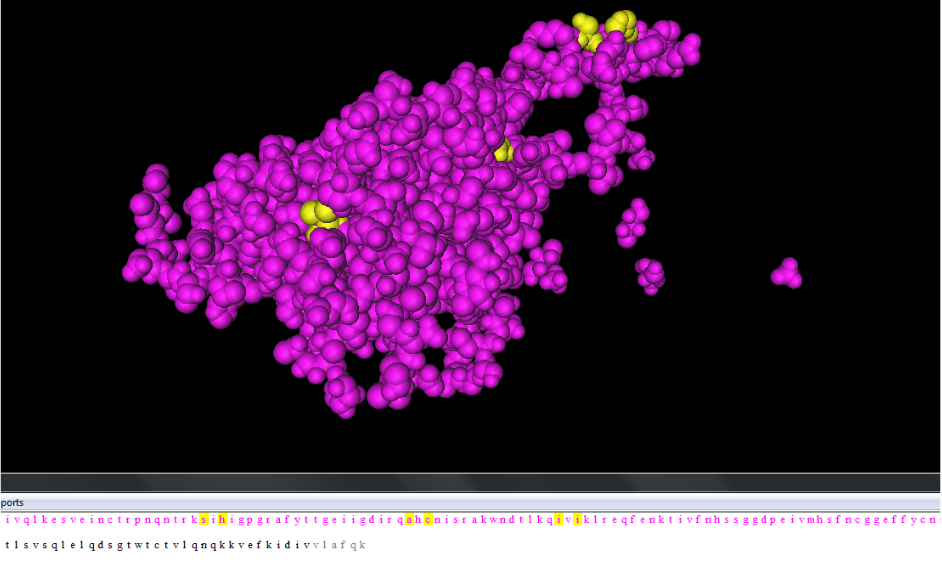
- Produce a 3D rendering of that protein with the amino acid changes of the clones which is highlighted on the CnD3 program.
- Identify where the non-consensus amino acids occur and analyze those locations and their influence on immunogenicity.
Data and Files
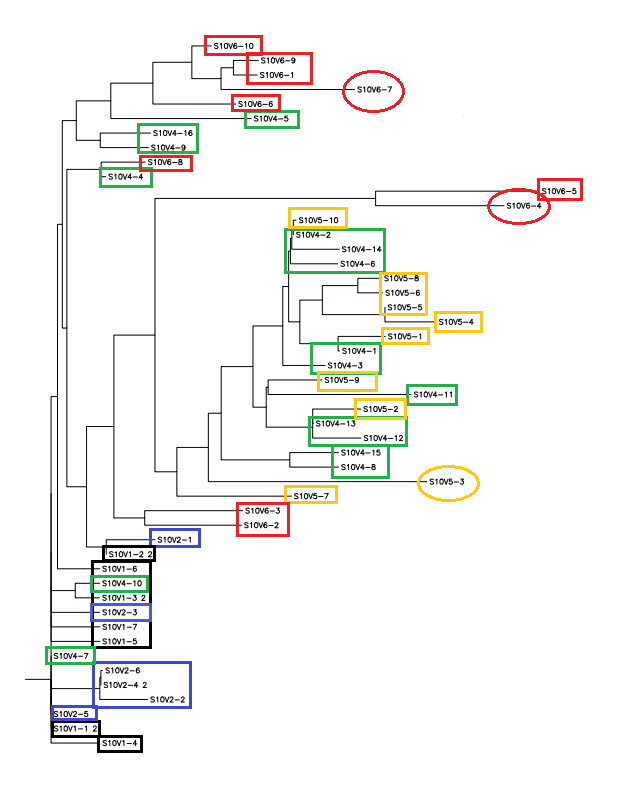
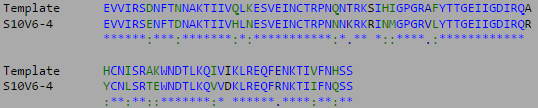
- Black: Visit 1, Indigo: Visit 2, Green: Visit 4, Orange: Visit 5, Red: Visit 6
- Yellow regions indicate non-consensus changes in amino acid.
- Powerpoint link: PDF
Results/Scientific Conclusion
- Week 10's work toward answering our question yielded interesting results. After preliminary testing of the cultivated clones for their amino acid sequences the clones that showed the highest divergence were data points S10V6-4, S10V5-3, and S10V6-7. Upon analyzing the phylogeny of these constituents since seroconversion they showed the highest divergence in terms of differences in amino acids since initial testings of the subject. Furthermore, there is an observed increase in divergence, since the first visit. Each sets of visits their clones associate in a sequential increase in divergence as further testings were conducted. High amino acid change in samples like S10V6-4, S10V6-5, and S10V6-7 may contribute to the worsening condition associated in the "rapid progressor" category of the Markham et al. (1998) study. There were several amino acids that could have a large impact on the immunogenicity of HIV given that these changes occured on the immunogenic tip and the surface of gp120. These changes can influence the interaction the virus has with the immune system and would therefore have a negative effect on CD4-T cell count.
Acknowledgments
I worked with Matt Oki in discussing the further directions of the project and the partial presentation of our current work and the results we've obtained thus far. We discussed these matters over the phone and in class While I worked with the people noted above, this individual journal entry was completed by me and not copied from another source. William P Fuchs 21:42, 14 November 2016 (EST)
References
- Week 11 Assignment Page
- Provided guidelines and instructions for this week's individual journal.
- ExPASY Translate tool
- UniProt Knowledgebase (UniProt KB)
- PredictProtein server
- Cn3D software site
- Huang, C. C., Tang, M., Zhang, M. Y., Majeed, S., Montabana, E., Stanfield, R. L., ... & Wyatt, R. (2005). Structure of a V3-containing HIV-1 gp120 core. Science, 310(5750), 1025-1028. doi: 10.1126/science.1118398.
- Markham, R.B., Wang, W.C., Weisstein, A.E., Wang, Z., Munoz, A., Templeton, A., Margolick, J., Vlahov, D., Quinn, T., Farzadegan, H., & Yu, X.F. (1998). Patterns of HIV-1 evolution in individuals with differing rates of CD4 T cell decline. Proc Natl Acad Sci U S A. 95, 12568-12573.doi: 10.1073/pnas.95.21.12568.
- Imported backbone for this journal entry from Week 10 Journal
Externals
Individual Journals
- WPF Week 2
- WPF Week 3
- WPF Week 4
- WPF Week 5
- WPF Week 6
- WPF Week 7
- WPF Week 8
- WPF Week 9
- WPF Week 10
- WPF Week 11
- WPF Week 12
- WPF Week 14
- WPF Week 15
Class Journals
- Week 1
- Week 2
- Week 3
- Week 4
- Week 5
- Week 6
- Week 7
- Week 8
- Week 9
- Week 10
- Week 11
- Week 12
- Week 13
- Week 14
- Week 15
Assignments
Useful Links
- Bio Class Page
- BIOL368/F16:People
- Will Fuchs
- Link to LMU: http://www.lmu.edu/