Kvescio Week 10 Assignment
Weekly Assignments
- user:kvescio ~ Instructions: BIOL368/S20:Week 1
- kvescio Week 2 Assignment ~ Instructions: BIOL368/S20:Week 2
- kvescio Week 3 Assignment ~ Instructions: BIOL368/S20:Week 3
- kvescio Week 4 Assignment ~ Instructions: BIOL368/S20:Week 4
- kvescio Week 5 Assignment ~ Instructions: BIOL368/S20:Week 5
- kvescio Week 6 Assignment ~ Instructions: BIOL368/S20:Week 6
- kvescio Week 8 Assignment ~ Instructions: BIOL368/S20:Week 8
- kvescio Week 10 Assignment ~ Instructions: BIOL368/S20:Week 10
- kvescio Week 11 Assignment ~ Instructions: BIOL368/S20:Week 11
- kvescio Week 13 Assignment ~ Instructions: BIOL368/S20:Week 13
- kvescio Week 14 Assignment ~ Instructions: BIOL368/S20:Week 14
Class Journals
- BIOL368/S20:Class Journal Week 1
- BIOL368/S20:Class Journal Week 2
- BIOL368/S20:Class Journal Week 3
- BIOL368/S20:Class Journal Week 4
- BIOL368/S20:Class Journal Week 5
- BIOL368/S20:Class Journal Week 6
- BIOL368/S20:Class Journal Week 10
- BIOL368/S20:Class Journal Week 11
- BIOL368/S20:Class Journal Week 13
- BIOL368/S20:Class Journal Week 14
Purpose
The Purpose of this assignment is to investigate a scientific database, and analyze how they organize and demonstrate their data. This is useful in order to further be able to analyze scientific research that will be important for this class, and for future scientific endeavors.
Questions
General information about the database
- What is the name of the database? (link to the home page)
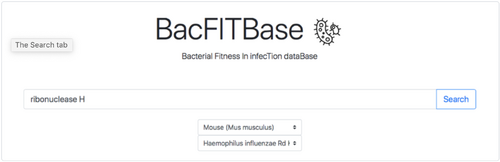
- BacFITBase: a database to assess the relevance of bacterial genes during host infection (http://www.tartaglialab.com/bacfitbase/)
- What type (or types) of database is it?
- Bacterial fitness in infection database. (http://www.tartaglialab.com/bacfitbase/)
- What biological information (type of data) does it contain? (sequence, structure, model organism, or specialty [what?])
- " It now covers the infection processes of 15 pathogenic bacteria in 5 model vertebrates across 10 different tissues" (http://www.tartaglialab.com/bacfitbase/about).
- What type of data source does it have?
- primary versus secondary ("meta")?
- Secondary. They list primary databases on their "about" page.
- curated versus non-curated?
- "BacFITBase is a manually curated database of bacterial genes that collates in vivo information on their relevance during host infection, as measured by transposon mutagenesis" (http://www.tartaglialab.com/bacfitbase/about)
- primary versus secondary ("meta")?
- if curated, is it electronic versus human curation?
- Human curation; team of scientists.
- if human curation, is it in-house staff versus community curation?
- In-house staff.
- What individual or organization maintains the database?
- Javier Macho, Benjamin Lang, Gian Gaetano Tartaglia, and Marc Torrent maintain the database. (http://www.tartaglialab.com/bacfitbase/about)
- public versus private
- Public. Freely available.
- large national or multinational entity or small lab group
- Small lab group.
- What is their funding source(s)?
- "This study has been funded by the Spanish Ministerio de Ciencia, Innovación y Universidades (SAF2015-72518-EXP, SAF2017-82158-R and RYC-2012-09999) and a Research Grant 2016 by the European Society of Clinical Microbiology and Infectious Diseases (ESCMID).
- European Union This project has also received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Sklodowska-Curie grant agreement No 793135." (http://www.tartaglialab.com/bacfitbase/about).
Scientific quality of the database
- Does the content appear to completely cover its content domain?
- How many records does the database contain?
- "Overall, BacFITBase includes information on 15 pathogenic bacteria and 5 host vertebrates across 10 different tissues"(http://www.tartaglialab.com/bacfitbase/about).
- How many records does the database contain?
- What claims do the database owners make about coverage in the corresponding paper?
- In the article, the authors state that it contains more than 90,000 entries with information on the contribution of individual genes to bacterial fitness under in vivo infection conditions. The data were collected from 15 different studies where transposon mutagenesis was performed.
- What species are covered in the database? (If it is a very long list, summarize.)
- Any host species can be searched, but also under the search bar one can specifically search under the hosts cow, chicken, mouse, rabbit and pig.
- Is the database content useful? I.e., what biological questions can it be used to answer?
- Yes, it is useful. It is used for the "systematic characterization of bacterial proteins relevant for host infection aimed to enable the identification of new antibiotic targets" (https://academic.oup.com/nar/article/48/D1/D511/5608989).
- Is the database content timely?
- Yes. See below for answers.
- Is there a need in the scientific community for such a database at this time?
- Yes, very much so. The authors state, "Bacterial infections have been on the rise world-wide in recent years and have a considerable impact on human well-being in terms of attributable deaths and disability-adjusted life years."
- Is the content covered by other databases already?
- Yes, some other databases have the content, but not as particular to the way BacFITBase is.
- How current is the database?
- Database is fairly new. From 2019.
- When did the database first go online?
- Database first went online in March of 2019
- How often is the database updated?
- Does not say how often it is updated, since it is fairly new. However, they do say that they will continue to update.
- When was the last update?
- Last update was in January of 2020
General utility of the database to the scientific community
- Are there links to other databases? Which ones?
- Yes, there are links to PubMed and ProViz. Also, http://pfam.xfam.org, https://blast.ncbi.nlm.nih.gov/Blast.cgi
- Is it convenient to browse the data?
- Yes, the Tool Bar and Search Engine are convenient and easy to understand. Also, the Tutorial tab is a useful way to learn how to use the database.
- Is it convenient to download the data?
- Yes, there is a download tab that navigates the user to the data bacfitbase_v1 that can be downloaded. Is currently being updated as well. (http://www.tartaglialab.com/bacfitbase/download).
- In what file formats are the data provided?
- FASTA
- What type of files, indicated by the file extension (e.g., .txt, .xml., etc.)?
- .tsv, .xlsx, .txt, .zip
- Are they standard or non-standard formats? (i.e., are they following an approved standard for that type of data)?
- Yes, .tsv and .xlsx are standard formats. .txt and .zip are non-standard for this type of data
Evaluate the “user-friendliness” of the database: can a naive user quickly navigate the website and gather useful information?
- Is the website well-organized?
- Yes, the website is very clean and well organized. There is not much clutter on any page, and it is easy to locate the tabs. The Tabs are Search, BLAST, Browse, Download, Tutorial and About.
- Does it have a help section or tutorial?
- Yes, there is a tutorial section: (http://www.tartaglialab.com/bacfitbase/tutorial).
- Are the search options sensible?
- Yes, there are different host and pathogen options. I do think they should add "human" under the host search tab, and they should continue to add more pathogens as more research is being done.
- Run a sample query. Do the results make sense?
- Access: Is there a license agreement or any restrictions on access to the database?
- No, but here is link to license agreement. (https://www.crg.eu/en/content/legal-notice-privacy-policy)
Summary judgment
- Would you direct a colleague unfamiliar with the field to use it?
- Yes, I would direct a colleague if they needed to search for this particular topic, but I would highly suggest they check out the Tutorial page first.
- Is this a professional or "hobby" database? The "hobby" analogy means that it was that person's hobby to make the database. It could mean that it is limited in scope, done by one or a few persons, and seems amateur.
- I think that BacFIT is more of a "hobby" database, because it was made by a few people, and it still has a lot of room for improvement. It is also a newer database, and they are still working to correct it and update the system. It has the ability to become more professional within the near future.
Scientific Conclusion
BacFITBase is a nucleic acid research database that searches the bacterial fitness in infections in the host. The fitness is measured by transposon mutagenesis. BacFITBase has many different entries with information about specific hosts and pathogens. The database is useful and can be understood, especially, under the Tutorial tab. It is privately made by a team of scientists, and is considered more of a "hobby" database, but as more research is done and data is added, the team says they will continue to update the database. In conclusion, BacFITBase is a good database, and the main goal is very important for the future of science and understanding bacterial infections. However, there is much room for improvement, and the creators should be constantly updating and working towards their ultimate goal.
Acknowledgements
- I copied and pasted the questions from Week 10 for this assignment.
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
(Kvescio (talk) 21:05, 1 April 2020 (PDT))
References
- OpenWetWare. (2020). BIOL368/S20:Week 10. Retrieved March 27, 2020, from https://openwetware.org/wiki/BIOL368/S20:Week_10
- Centre for Genomic Regulation: LEGAL NOTICE AND PRIVACY POLICY. Retrieved March 31, 2020, from https://www.crg.eu/en/content/legal-notice-privacy-policy
- Javier Macho Rendón, Benjamin Lang, Gian Gaetano Tartaglia, Marc Torrent Burgas, BacFITBase: a database to assess the relevance of bacterial genes during host infection, Nucleic Acids Research, Volume 48, Issue D1, 08 January 2020, Pages D511–D516, https://doi.org/10.1093/nar/gkz931
- Oxford Academic. Nucleic Acids Research (2019). Retrieved March 27,2020 from https://academic.oup.com/nar/issue/48/D1