JT Correy Journal Week 5
Week 5
Purpose
The purpose of this week’s assignment is to investigate the interaction between the ACE2 protein and the protein spike of the 2019-nCOV virus through different modeling software that emphasizes certain parts of the protein.
Tasks
- I went to UniProt Knowledgebase (UniProt KB) to utilize their two forms of DNA and protein sequencing. They have a Swis-Prot which is curated and reviews by humans, and they have TrEMBL which is analyzed and filtered by a computer program.
- After searching for “sars-cov-2” on UniProt I got a result of 1616 different results with 18 of them being reviewed by a curator. Not all of the proteins came from the virus, there were also results coming from humans and mice.
- After searching the accession number “P0DTC2” provides information such as the name and taxonomy, subcellular location, pathology and biotech, processing, and structure.
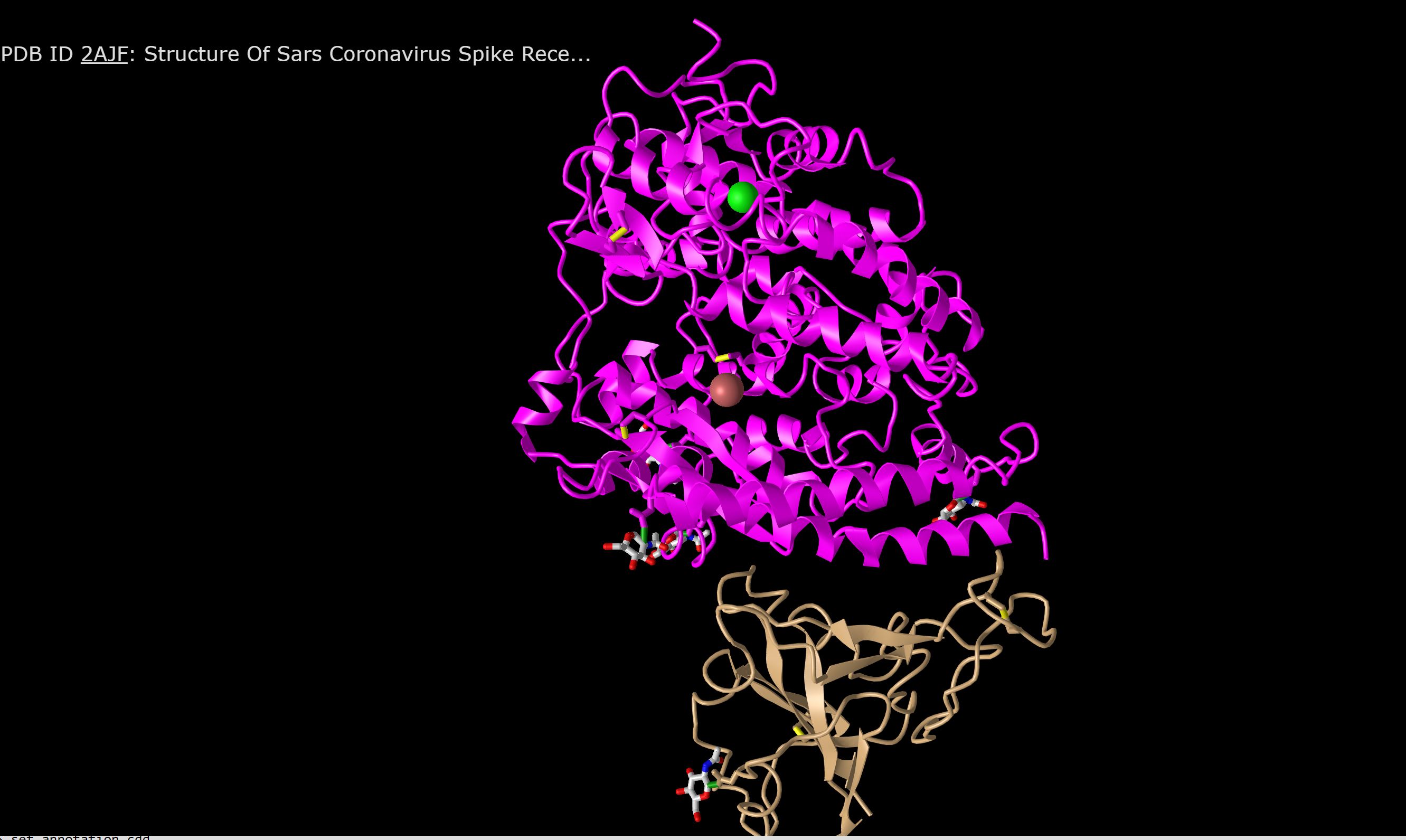
- Utilizing the modeling system of the SARS-CoV protein spike from NCBI Structure Database we analyzed Figure 1A SARS-CoV RBD (year 2002) complexed with human ACE2: 2AJF.
- We adjusted the model so the positioning aligned with that of Figure 2A from the Wan et al. (2020) study.
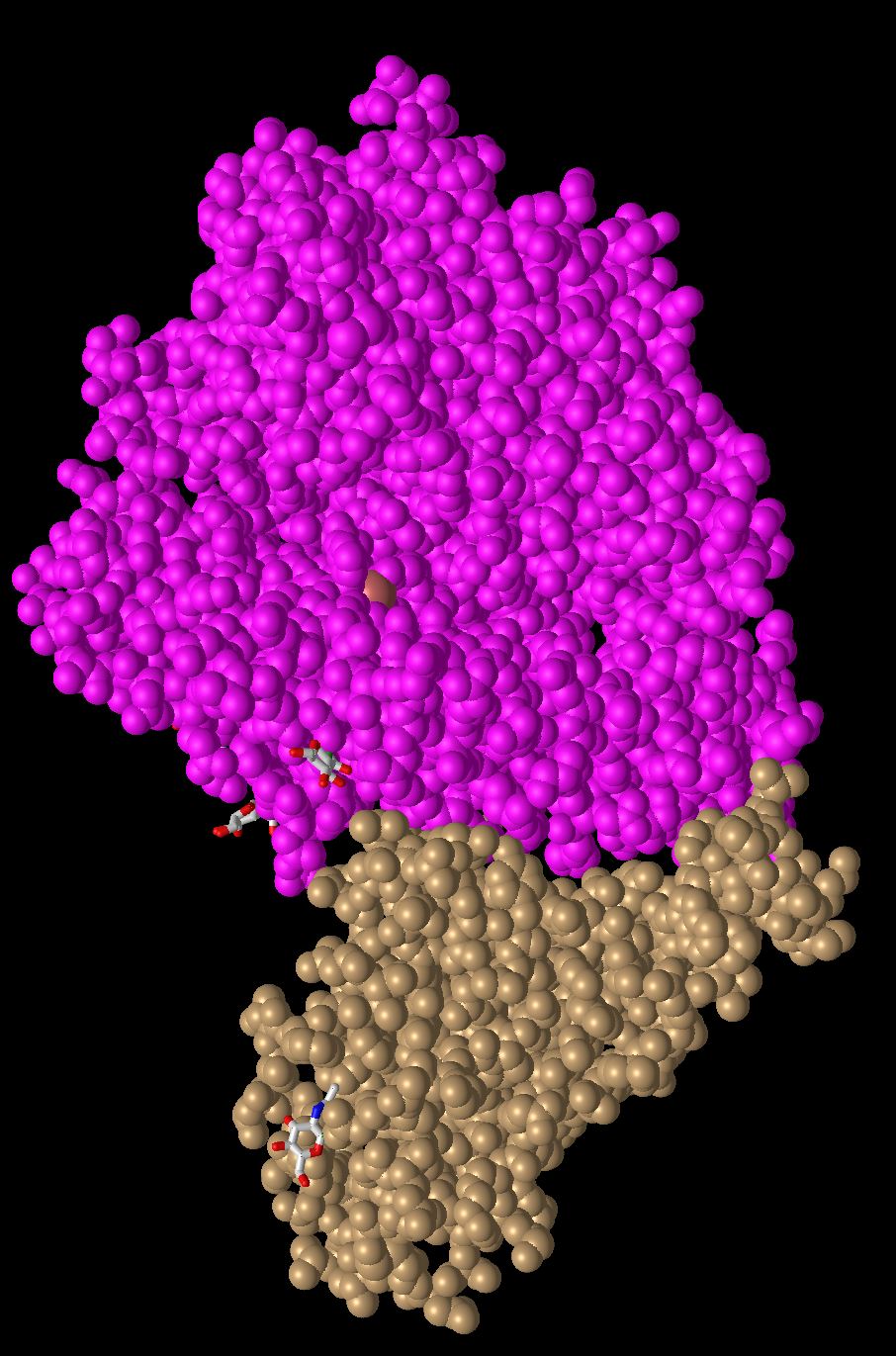
- This is quaternary structure because there is interaction between more than one protein. There is non-covalent bonds that are interacting between the two.
- There are 2 domains in this figure because there are 2 proteins each with their own function and structure. Each is identified by their own color, one being brown and one being pink. These structures can separated without causing either chain to break.
- The protein model was then modified by selecting different styles in the "Style" then "Proteins" menu.
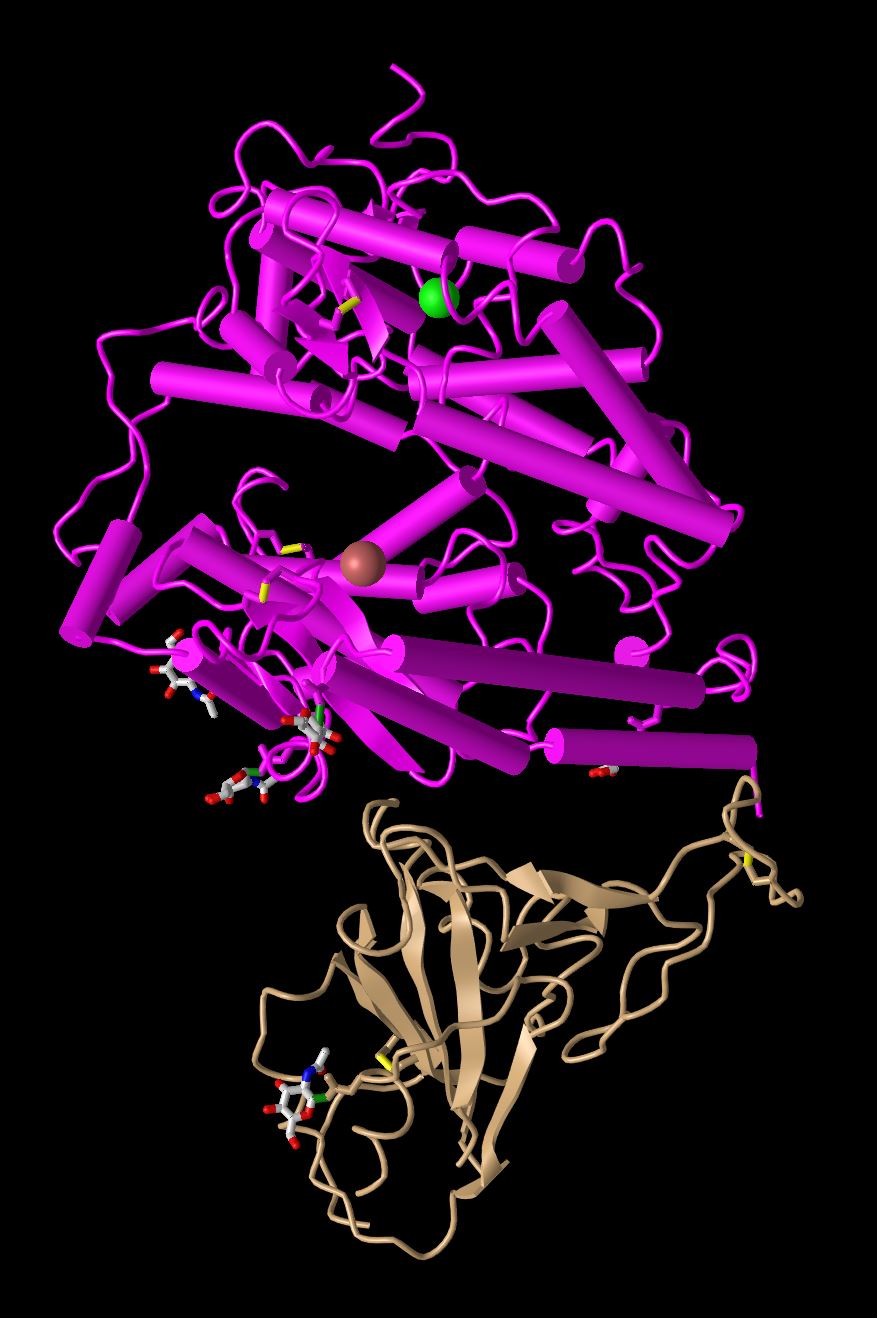
- Cylinder and plate
- In this diagram the alpha helices are condensed into solid cylinders and the rest of the protein is unchanged. This allows the overall structure to have the helices stand out more and we can easily identify them.
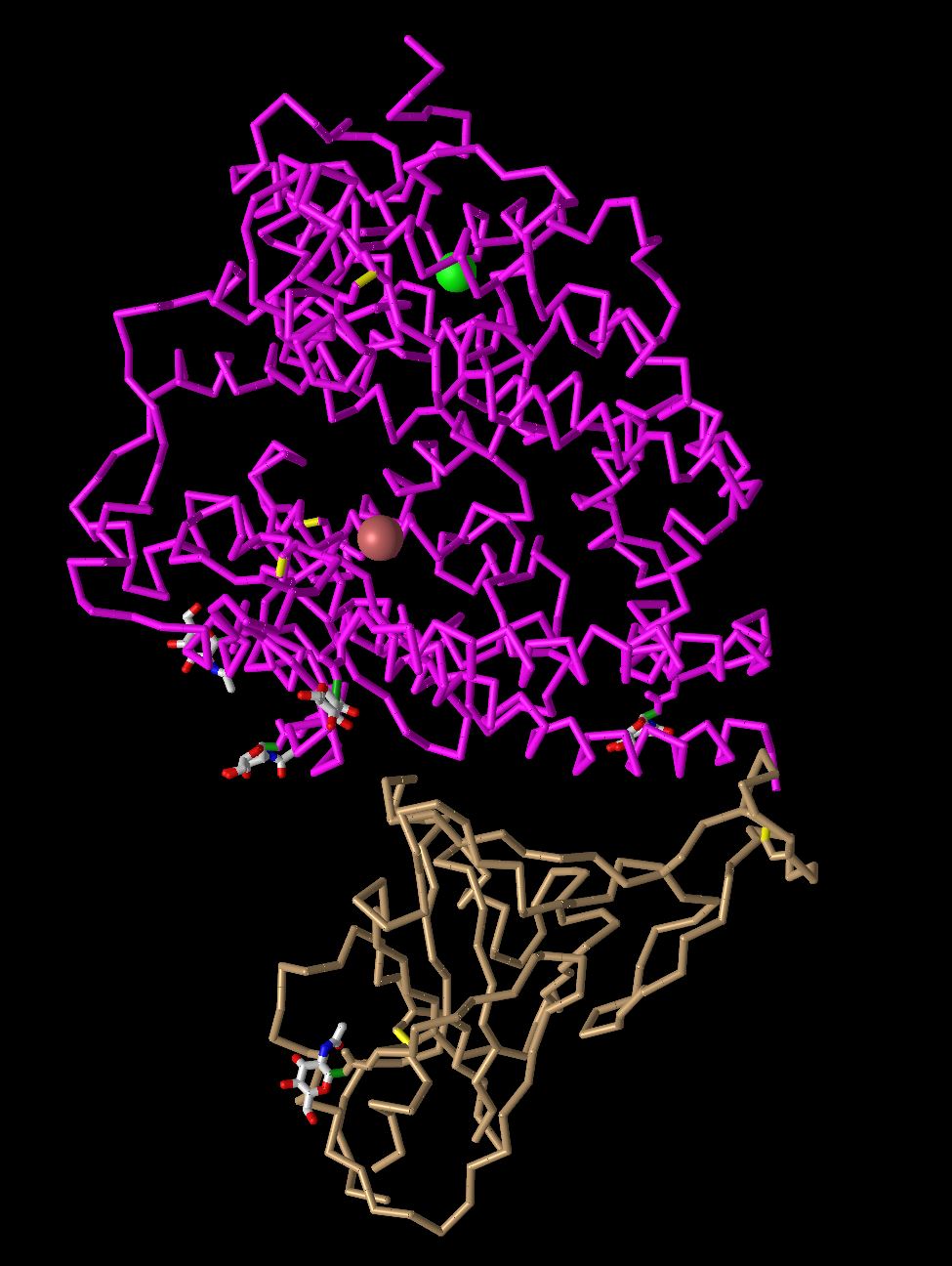
- C alpha trace
- In this model the program displays what seems to be the carbon backbone of the protein. It omits most of the secondary structures and just simply follows the backbone. It also tells what amino acid is present at any given position.
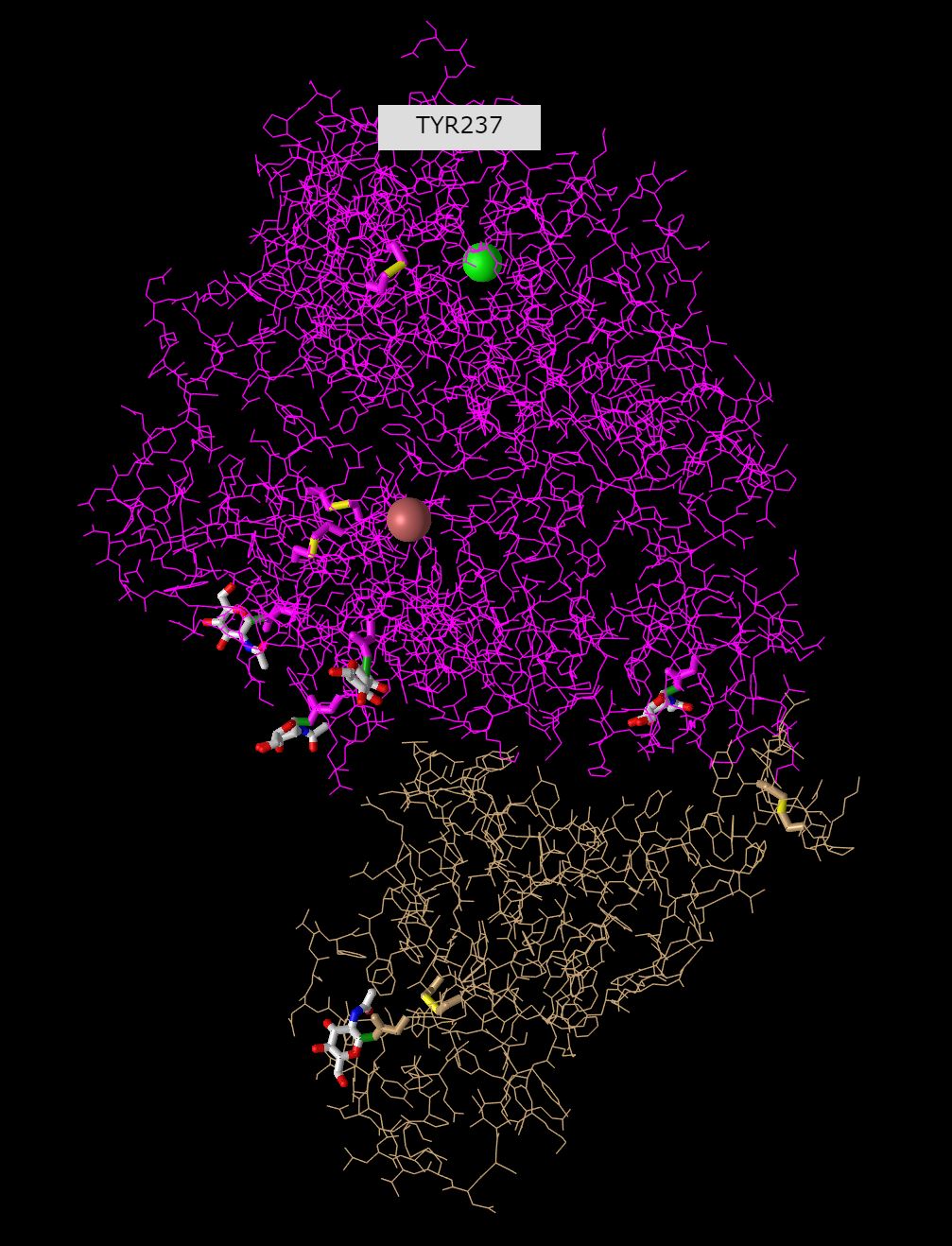
- Lines
- In the model all the carbons are displayed, including the r-groups of each amino acid. It is very clustered and dense so it is hard to analyze the overall structure, and almost impossible to identify and secondary structures. It is useful to analyze one small set of amino acids, but looking at the overall protein in this format is not very clear.
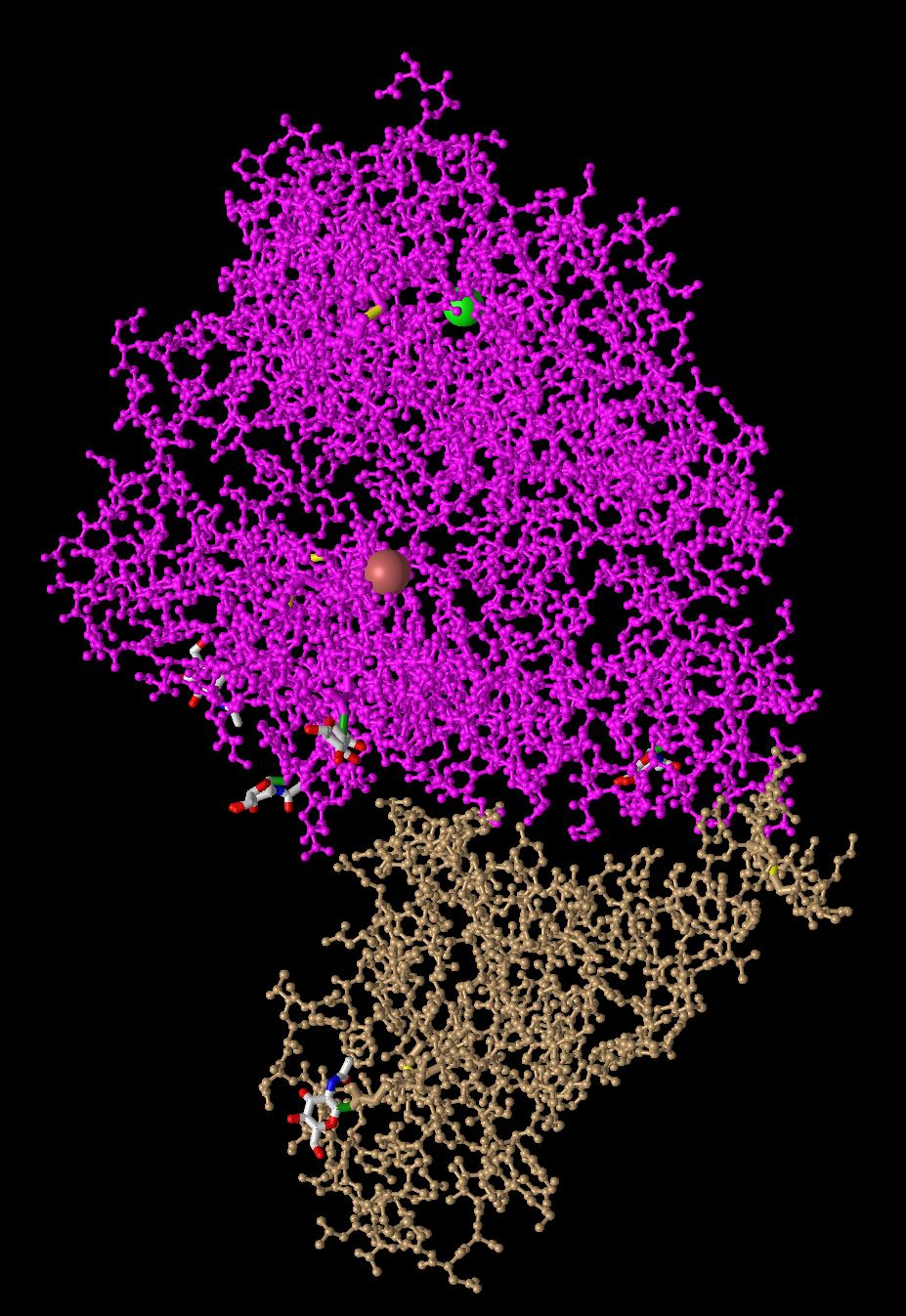
- Ball and Stick
- In this model all the atoms are displayed as balls and the bonds are displayed at sticks. It is useful to analyze small sections just like the previous model. But to clustered to analyze the protein overall. This is a good representation because we can analyze the structure of how the atoms align rather just the general shape.
- Spheres
- In this model each atom is its own sphere, it is very similar to the classic space-filling model. This is very useful to get any idea of the overall shape of the protein, but less useful for identifying smaller section of the protein. It also does a good job of displaying where the two domains come into contact with each other.
- Cylinder and plate
- The protein was then left in the "Spheres" style and the color scheme was altered by going to the color menu and selecting the following schemes.
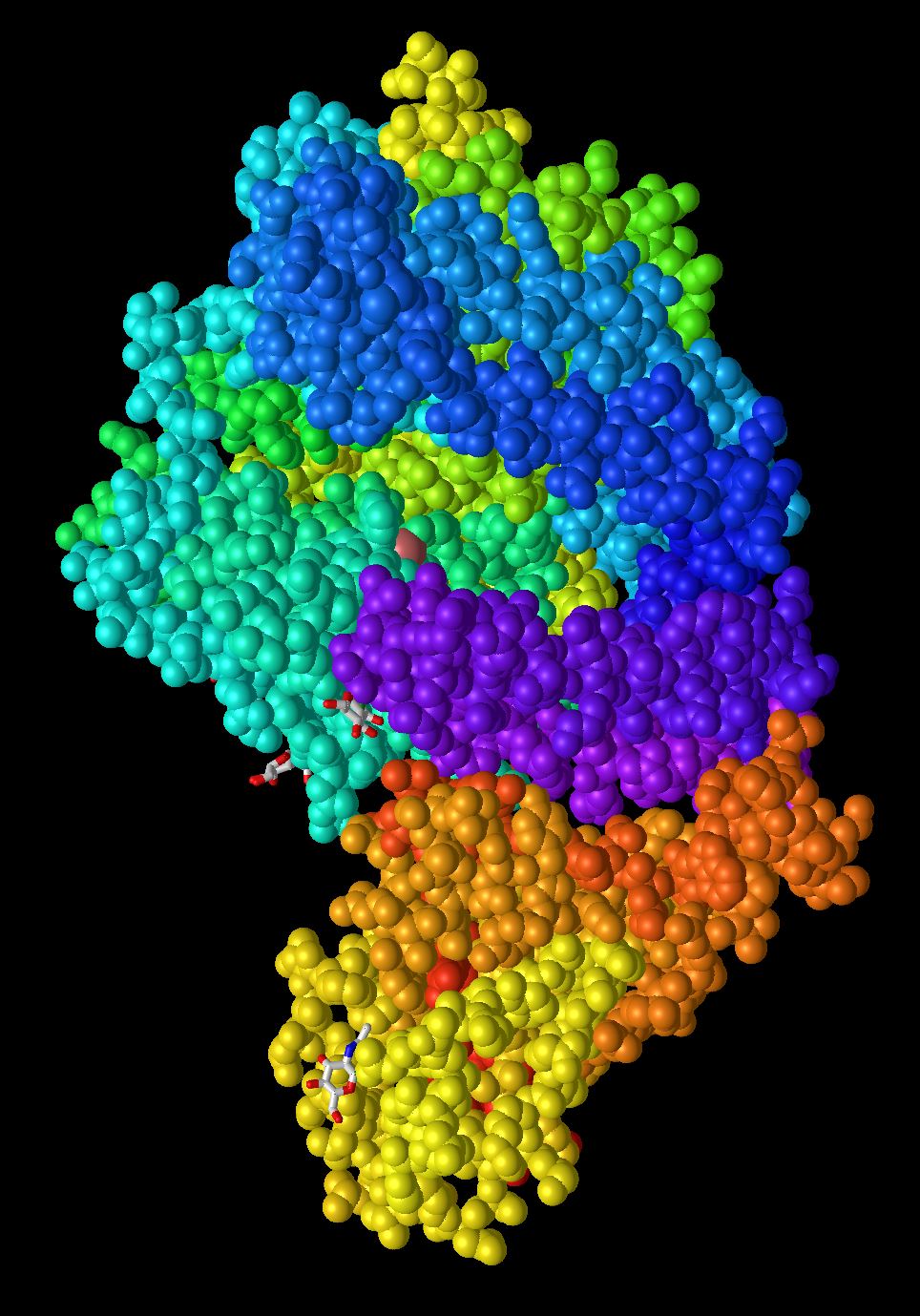
- Spectrum
- In this model the protein is organized in sections by adding different colors to areas on the protein. This model is helpful for breaking the protein into smaller subunits but challenging to identify what is occurring inside those groups. The method used to organize the protein into different colors is unknown.
- Secondary
- In this model the structure highlights the secondary structures of the protein. Alpha helices and beta sheets are each illuminated in a unique color. The unstructured chains between the structures are just shown in gray. This is very helpful in analyzing the overall structure of the protein as well as identifying where the secondary structures are.
- Charge [[week5picture9.JPG]
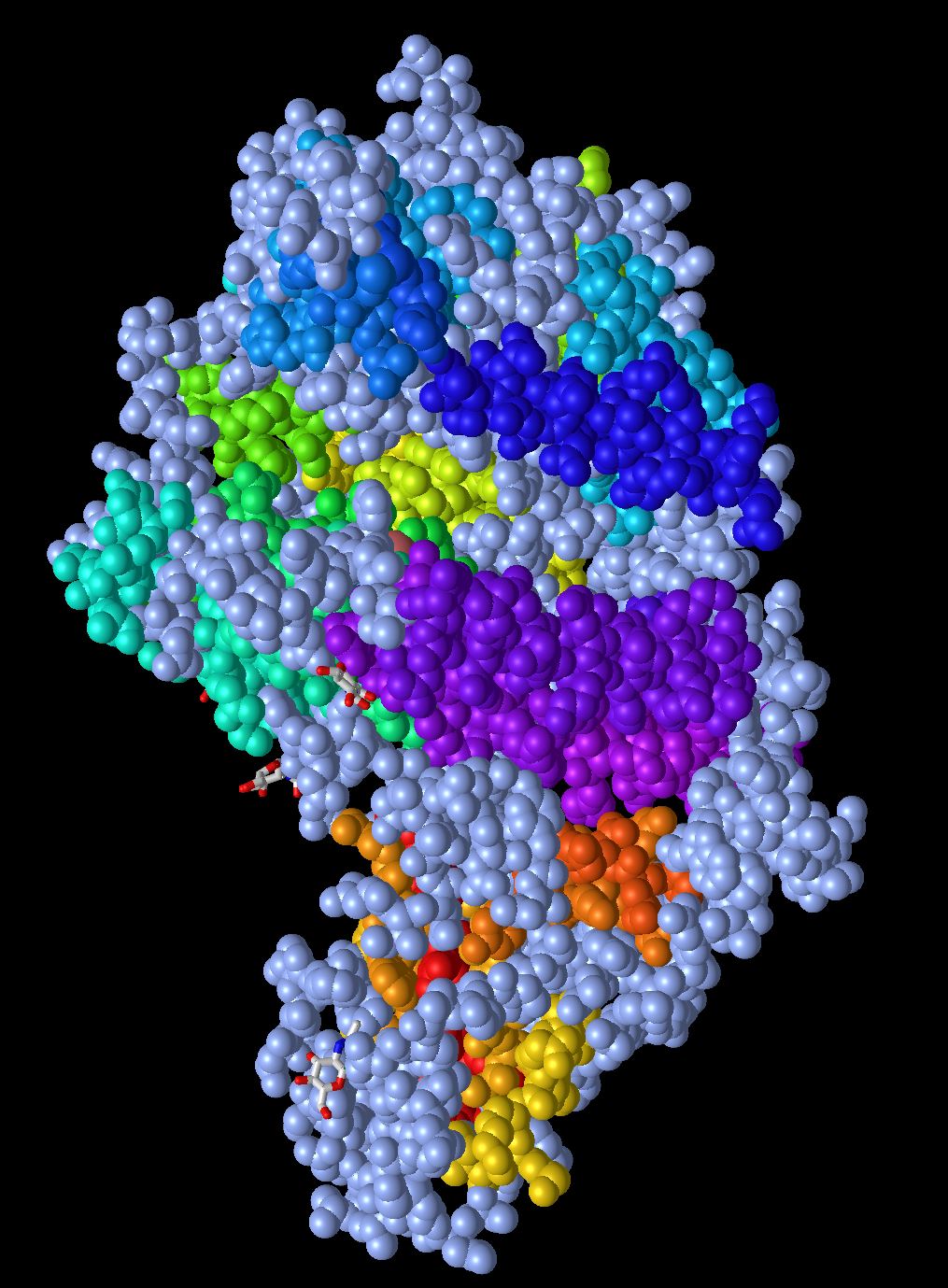
- In this model the different charges of the amino acids are displayed in colors. The positively charged amino acids are shown in blue and the negatively charged amino acids are shown in red. Histidine is a special case because its Pka is very close to biological pH so it is shown in a light blue color. All the uncharged and nonpolar amino acids are displayed in gray. This is helpful because it allows us to see how the protein may interact with itself and its environment.
- Atom week5picture10.JPG
- In this model the protein is organized by color coding each atom. Carbon is gray, oxygen is red, nitrogen is blue, and sulfur is gold. This display is helpful because it allows us to see any place that a sulfide bond is potentially available as well as areas why hydrogen bonding is likely to occur.
- Spectrum
- We adjusted the model so the positioning aligned with that of Figure 2A from the Wan et al. (2020) study.
- The secondary structures were then analyzed through the following procedure and the following results were found
Click on the link to interact with the structure in iCn3D
- Click on the Windows menu to “View Sequences & Annotations”
- In the new window that appears to the right, click on the “Details” tab to show the actual amino acid sequences
- There are 2 sets of ACE2-spike proteins because of the way the proteins crystallized.
- Focus on the pink and tan chains and orient them like is shown in Figure 4B
- We are going to make the amino acid side chains shown in the figure visible.
- In the sequence window go to sequence “Protein 3SCK_A” (in pink) and select the following amino acids
- T31
- E35
- E38
- T82
- K353
- The part of the ribbon that represents these amino acids should be highlighted in yellow in the structure
- Go to the Styles menu and select Proteins > Ball and Stick
- Go to the Color menu and select Atom
- You should see the side chains shown in the figure.
- Repeat this process for the tan spike protein sequence 3SCK_E for the following amino acids:
- T487
- R479
- G480
- Y442
- P472
- In the ACE2 protein
- There are beta sheets and alpha helices found in the ACE2Protein. By looking at the cylinder and plate model we can clearly see several alpha helices in the ACE2 protein. There are also arrows in the cylinder and plate model that represent the beta sheets. These structures can be easily located with the secondary color scheme.
- In the spike protein
- There are beta sheets in the spike protein that are represented by arrows in the model. There are no alpha helices in the spike protein and this can be seen by using the cylinder and plate diagram where there are clearly no cylinders in the protein.
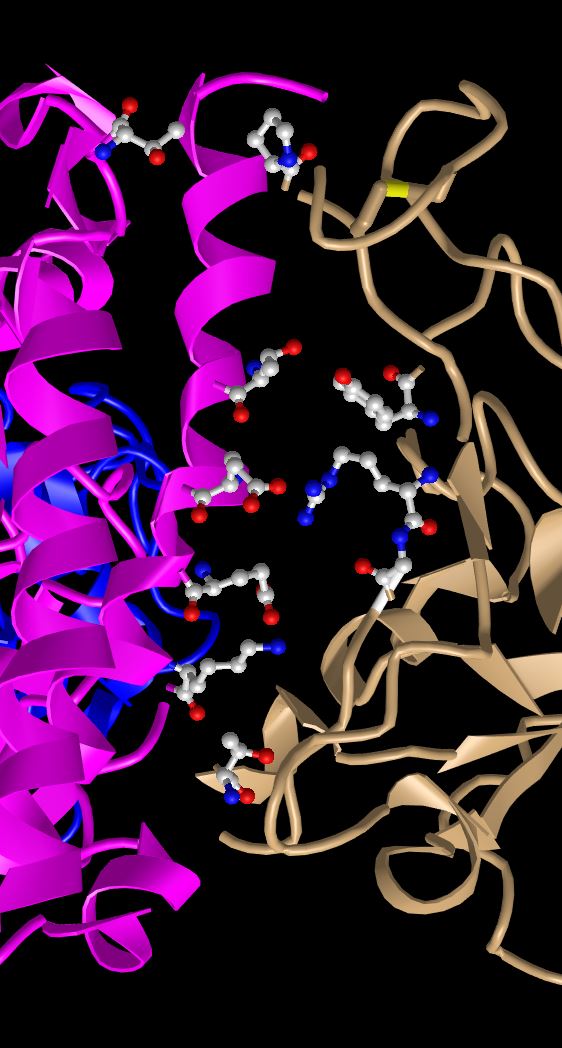
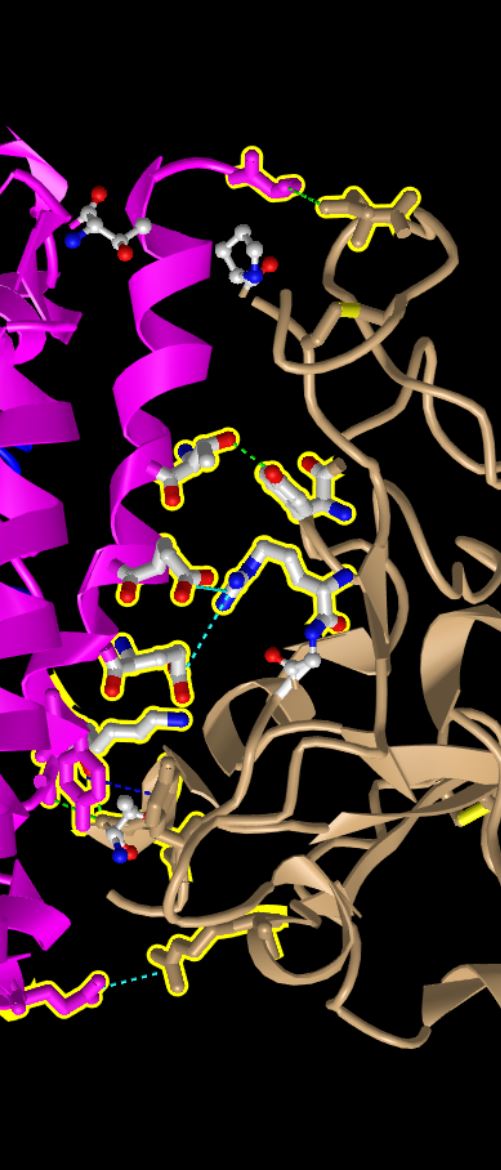
- The following were more observations about the characteristics of the side chains that interact between the two proteins.
- R479 is the positively charged sidechain and E35 is the negatively charged sidechain. We know this because nitrogens are displayed in blue when using the ball and stick model and oxygens are displayed in red when using the ball and stick model. Nitrogen commonly has an extra hydrogen bonded to it that it becomes basic and can donate a hydrogen. Oxygen commonly has a negative charge because it has one less hydrogen then optimal.
- Both chemical groups are uncharged polar chemical groups because thy both have hydroxyl (OH) groups that are uncharged. T31 corresponds to a threonine and Y442 corresponds to a tyrosine. We can tell by the red oxygen atoms on the model.
- Finally, the interaction between these side chains was visualized with the following process
- You can make the dashed lines for these bonds in iCn3D as follows:
- Go to the View menu and select H bonds & Interactions
- In part 1 of the window that appears, uncheck “Contacts/Interactions” leaving Hydrogen Bonds and Ionic Interaction checked
- In part 2 of the window, select the first set “3SCK_A” (pink)
- In part 3 of the window, select the second set “3SCK_E” (tan)
- In part 4 of the window, click the button “3D Display”
- You can make the dashed lines for these bonds in iCn3D as follows:
Beginning the research project
- How do differences in ACE2 receptors for host organisms affect binding strength with SARS-CoV?
- We will use the UniProt database and possibly GenBank for obtaining sequences. We will remain within the mammals category and possibly look at the sequences for ACE2 of Humans, Bats, Civet Cats, Pangolins, Mice, Squirrels, Rats, Birds, Dogs, Cats, Lions, Tigers, Ferrets, and Minks.
Conclusion
The interaction between the 2019-nCOV protein spike and the ACE2 protein in humans was displayed by a 3D interactive model. By using a 3D model to represent the interaction was easy to analyze through making slight alterations to the model. Through different color schemes and structure styles we were able to illuminate different section of the protein (like using the secondary color scheme to display the secondary structure or lines structure style to see the r-groups of the amino acids in the protein).
References
- OpenWetWare. (2020). BIOL368/F20:Week 5. Retrieved 7 October 2020, from https://openwetware.org/wiki/BIOL368/F20:Week_5
- OpenWetWare. (2020). BIOL368/F20:Week 1. Retrieved 7 October 2020, from https://openwetware.org/wiki/BIOL368/F20:Week_1
- OpenWetWare. (2020). Yaniv Maddahi Week 5. Retrieved u October 2020, https://openwetware.org/wiki/Yaniv_Maddahi_Journal_Week_5
- Foldit - Blogs. (2020). Retrieved 6 October 2020, from https://fold.it/portal/blog
- Foldit - Solve Puzzles for Science. (2020). Retrieved 7 October 2020, from https://fold.it/portal/
- iCn3D: Web-based 3D Structure Viewer 2AJF. (2020). Retrieved 7 October 2020, from https://www.ncbi.nlm.nih.gov/Structure/icn3d/full.html?&mmdbid=35213&bu=1&showanno=1
- iCn3D: Web-based 3D Structure Viewer 3SCK. (2020). Retrieved 7 October 2020, from https://www.ncbi.nlm.nih.gov/Structure/icn3d/full.html?pdbid=%203SCK
- Uniprot. (2020). S - Spike glycoprotein precursor - Severe acute respiratory syndrome coronavirus 2 (2019-nCoV) - S gene & protein. Retrieved 1 October 2020, from https://www.uniprot.org/uniprot/P0DTC2
- Wan, Y., Shang, J., Graham, R., Baric, R., & Li, F. (2020). Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. Journal Of Virology, 94(7). doi: 10.1128/jvi.00127-20
- Andersen, K.G., Rambaut, A., Lipkin, W.I. et al. The proximal origin of SARS-CoV-2. Nat Med 26, 450–452 (2020). https://doi.org/10.1038/s41591-020-0820-9
Acknowledgements
- Yaniv Maddahi
- Yaniv and I worked as homework partners for this week. We communicated and worked together both at the end of the week 5 lab and throughout the week to create our research question and assignment pages.
- Dr. Dahlquist
- Dr. Dahlquist served as a coach for how to begin our pages. She also instructed the class and provided us with the guiding homework document.
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
Jcorrey (talk) 21:13, 7 October 2020 (PDT)
JT Correy Template
Weekly Assignments
- Week 1 Assignment
- Week 2 Assignment
- Week 3 Assignment
- Week 4 Assignment
- Week 5 Assignment
- Week 6 Assignment
- Week 7 Assignment
- Week 8 Assignment
- Week 9 Assignment
- Week 10 Assignment
- Week 11 Assignment
- Week 12 Assignment
- Week 14 Assignment
Individual Journal Pages
- JT Correy
- JT Correy Journal Week 2
- JT Correy Journal Week 3
- JT Correy Journal Week 4
- JT Correy Journal Week 5
- JT Correy Journal Week 6
- JT Correy Journal Week 7
- CancerTracer Review
- JT Correy Journal Week 9
- JT Correy Journal Week 10
- JT Correy Journal Week 11
- JT Correy Journal Week 12
- JT Correy Journal Week 14
- The Mutants Week 14
Class Journal Pages
- BIOL368/F20:Class Journal Week 1
- BIOL368/F20:Class Journal Week 2
- BIOL368/F20:Class Journal Week 3
- BIOL368/F20:Class Journal Week 4
- BIOL368/F20:Class Journal Week 5
- BIOL368/F20:Class Journal Week 6
- BIOL368/F20:Class Journal Week 7
- BIOL368/F20:Class Journal Week 8
- BIOL368/F20:Class Journal Week 9
- BIOL368/F20:Class Journal Week 10
- BIOL368/F20:Class Journal Week 11
- BIOL368/F20:Class Journal Week 12
- BIOL368/F20:Class Journal Week 14