Lurbinah Week 6
Assignments
- BIOL368/S20:Week 1
- BIOL368/S20:Week 2
- BIOL368/S20:Week 3
- BIOL368/S20:Week 4
- BIOL368/S20:Week 5
- BIOL368/S20:Week 6
- BIOL368/S20:Week 8
- BIOL368/S20:Week 9
- BIOL368/S20:Week 10
- BIOL368/S20:Week 13
- BIOL368/S20:Week 14
Weekly Assigments
- lurbinah Week 2
- lurbinah Week 3
- lurbinah Week 4
- lurbinah Week 5
- lurbinah Week 6
- lurbinah Week 7
- lurbinah Week 8
- lurbinah Week 9
- lurbinah Week 10
- lurbinah Week 11
- lurbinah Week 12
- lurbinah Week 13
- lurbinah Week 14
Class Journal Assigments
- BIOL368/S20:Class Journal Week 1
- BIOL368/S20:Class Journal Week 2
- BIOL368/S20:Class Journal Week 3
- BIOL368/S20:Class Journal Week 4
- BIOL368/S20:Class Journal Week 5
- BIOL368/S20:Class Journal Week 6
- BIOL368/S20:Class Journal Week 7
- BIOL368/S20:Class Journal Week 8
- BIOL368/S20:Class Journal Week 9
- BIOL368/S20:Class Journal Week 10
- BIOL368/S20:Class Journal Week 11
- BIOL368/S20:Class Journal Week 12
- BIOL368/S20:Class Journal Week 13
- BIOL368/S20:Class Journal Week 14
Purpose
The purpose of this assigment is to answer and analyze the question proposed on preovoius week using data provided by Markham etl.
Question
Are the interim visits important to record and analyze the decline of CD4 T cells in order to accurately map the progression of HIV evolution?
Methods/Results
Statistical Analysis of CD4 T-cells count and sequenced data.
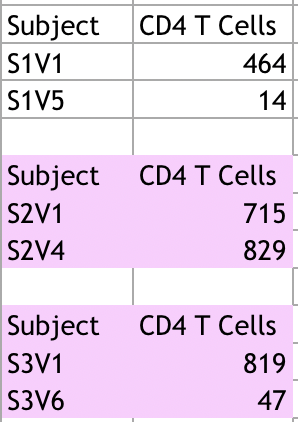
- We analyzed the data collected from Markham et al. 1998 data table, and gather the information needed to make our own table and graph
- The data used for this porject was the first and last CD4 T cell count with its corresponding sequence
- Some subjects had more visits but not record of DNA being sequenced, these values of CD4 T cells counts were not used in the analysis.
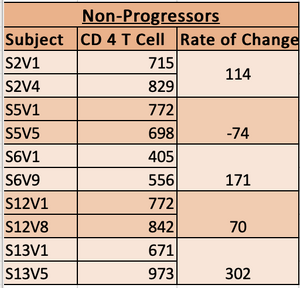
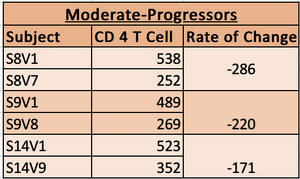
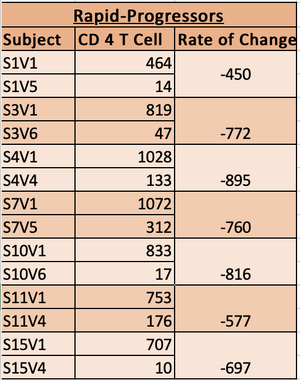
- Table used to graph and record data. The highligted subjects had more CD4 T cells counts record that were not used, due to the lack of DNA sequence.
Graph and calculate the slope in order to classify the subjects.
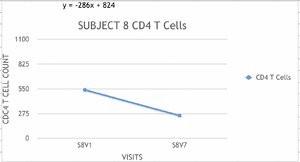
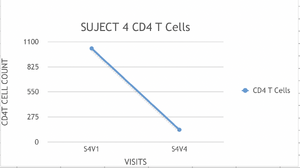
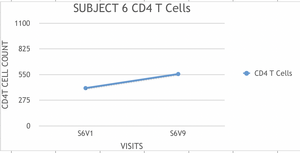
- Based on each of the data collected for each subject, a graph was elaborated to calculate the rate (slope) of change of the CD4 T cell counts.
- The scale used was the same in all the graphs.
- Example of Graphs
Classidication of Subjects
- Based on the rate of change the subjects were classified into Non-Progressor, Moderate Progressors and Rapid Progressors.
- Rules for Classification
- Subjects with slope greater than -100 were classified as non-progressors.
- Subjects with slope from -100 to -300 were classified as moderate progressors.
- Subjects with slopes with a minimun of -350 were classified as rapid Progressors.
- Tables with the Classification
Compare and contrast our classification and Markham’s.
- The only subjects with a discrepancy from Marks classification are 5, 6, 7
- Subject 5,6,7 were classified as moderate progressor and in our analysis 5 and 6 fall under non Progressor and and 7 as rapid progressor.
- This could be due to the CD4 T cell count values that were not taken into consideration.
- Subject 5,6,7 were classified as moderate progressor and in our analysis 5 and 6 fall under non Progressor and and 7 as rapid progressor.
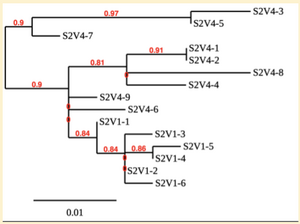
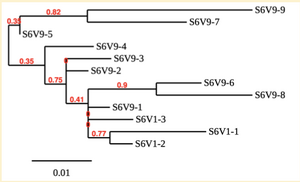
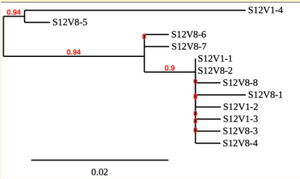
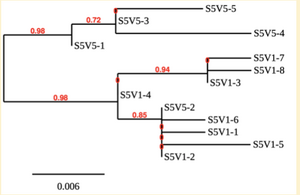
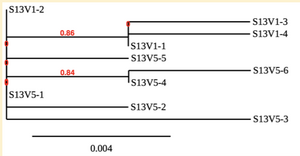
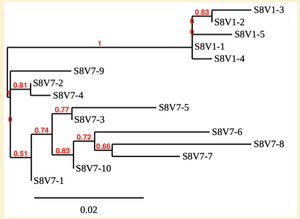
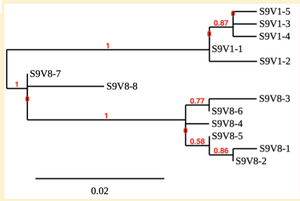
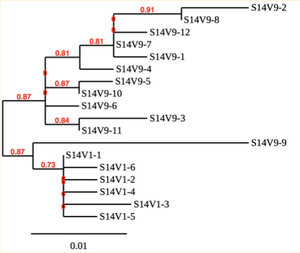
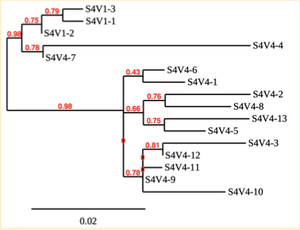
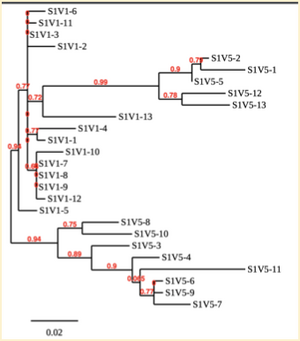
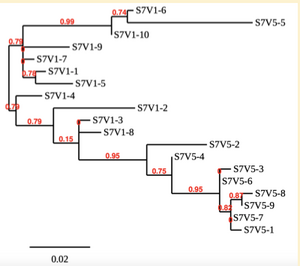
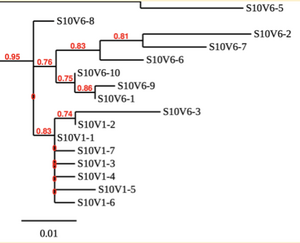
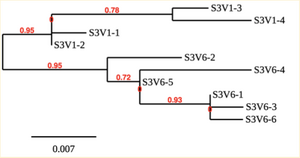
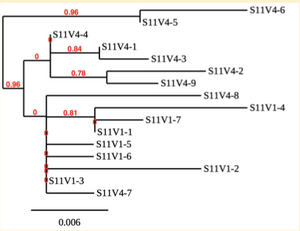
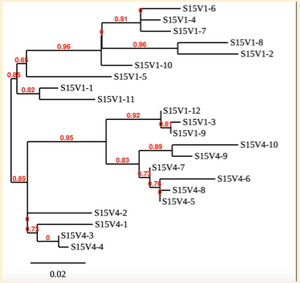
Creation of the phylogenetic trees of each subject.
- Phylogenetic tree where elobarated for each subject using their first and last visit sequences to look for similarities and differences among and within each group.
Data and Files
- subject 1 tree
- subject 2 tree
- subject 3 tree
- subect 4 tree
- subject 5 tree
- subject 6 tree
- subject 7 tree
- subject 8 tree
- subject 9 tree
- subject 10 tree
- subject 11 tree
- subject 12 tree
- subject 13 tree
- subject 14 tree
- subject 15 tree
- non progressor chart
- moderate progressor chart
- rapid progressor chart
Conclusions
CD4 T-cell decline is directly proportional to the evolution of HIV. Additonally each group responded differently to the ability or selection to change. Non-Progressors favored absence of change, maintaining stable CD4 T-cell counts.This was concluded from the trend showed in the phylogetic trees; Non-Progressors showed less divergence and diversity. However, based on the increase in diversity and divergence in the phylogenect trees moderate and Rapid Progressors showed; progressors favored against absence of change, revealing a relationship between mutation and decline in CD4 T-cell counts. Lastly, the exclusion of analysis of interim visits did not affect the results of overall CD4 T-cell decline in relatioship to the mapping of the evolution of HIV
References
- Markham, R.B., Wang, W.C., Weisstein, A.E., Wang, Z., Munoz, A., Templeton, A., Margolick, J., Vlahov, D., Quinn, T., Farzadegan, H., & Yu, X.F. (1998). Patterns of HIV-1 evolution in individuals with differing rates of CD4 T cell decline. Proc Natl Acad Sci U S A. 95, 12568-12573. doi: 10.1073/pnas.95.21.12568 (PubMed ID: 9770526)
- OpenWetWare. (2020). BIOL368/S20:Week 6. Retrieved February 20, 2020, from https://openwetware.org/wiki/BIOL368/S20:Week_6
- Phylogeny.fr. (2020) Phylogeny.fr:Home. Retrieved February 20, 2020, from http://www.phylogeny.fr/.
Aknowledgement
- I copied and modified directions and protocol from Week 6.
- I worked with Karina Vescio and Sahil in class and outside of class to analyze data and create a presentation.
- Copied and pasted nucleotide sequences to create phylogenetic trees from PubMed.
- Made phylogenetic trees and sequence Clustal forms via Phylogeny.fr
- Used data provieded in Nucleotide Sequence Data
- Communicated for guidance in class and outside class Kam Dahlquist in how to proceed with our project.
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source. Lurbinah (talk) 23:39, 26 February 2020 (PST)