BioMicroCenter:PacBio
| HOME -- | SEQUENCING -- | LIBRARY PREP -- | HIGH-THROUGHPUT -- | COMPUTING -- | DATA MANAGEMENT -- | OTHER TECHNOLOGY |
PACBIO SEQUEL
|
 |
The Sequel works by detecting DNA replication in real time. Fluors are attached to the gamma-phosphate of the nucleotides. A PacBio library with a single polymerase is loaded into each well. When a nucleotide enters the active site, it can be detected by the sequencer. Basepaired nucleotides have a longer dwell time allowing them to be detected.
The Sequel is able to monitor >500,000 incorporations occurring simultaneously on a SMRTcell. Each run creates a “movie” of the incorporation of bases, collecting data on the intensity of each fluorophore in each well over time. On board computation converts this movie into basecall files.
Using an analysis mode called Circular Consensus Sequencing, high fidelity (HiFi) subreads can be assembled when libraries are sequenced end to end in rolling fashion multiple times. In CCS, a 2 kbp amplicon can easily be sequenced over 10 times end to end over the course of a 20 hr movie. A mode called Continuous Long Read Sequencing simply sequences a subread sequenced over the length of the run, either 10 or 20 hours. In CLR mode, rare lengths of 175 kbp have been seen, but in the Center, it is more typical to see read lengths 50-70 kbp: the longest reads are typically outliers since the average subread length is typically <50 kbp.
EXPECTED RESULTS
TYPICAL RESULTS FROM SMRTCELLS:
- 20kb fragments – 50,000 active sites – 400Mbp sequence
- 2kb fragments – 80,000 active sites – 500Mbp sequence
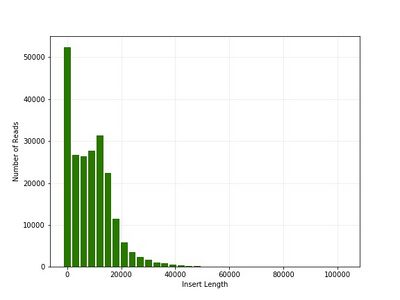
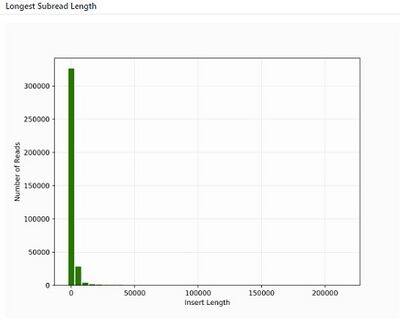
Typical Read Lengths
APPLICATIONS
There are several applications for which PacBio sequencing are well suited (and many it is not!). The major ones are below:
Assembly is the biggest strength of PacBio. While the individual base error rate is quite high (~14%), the errors are random and so resequencing rapidly lowers the error rate. Using only PacBio reads, 60x coverage can give very good assembly – typically completing genomes in 1 shot . A 4 Mbp genome can be done on a single SMRT cell.
A second strategy is to use PacBio reads to supplement Illumina reads to join scaffolds. Here, standard MiSeq reads make up the bulk of the assembly but the separate scaffolds are spanned by PacBio reads. 7x reads has been the convention for this type of sequencing. 40 Mbp of genomic DNA can be done on a single SMRT cell. Indexing cannot be done easily with PacBio samples - particularly long reads - and so the 'quanta' for SMRTcells should be considered as 1 genome per SMRTcell.
PacBio reads can also be used to looks for variants in specific regions. The long reads allow for better detection of large rearrangements and understanding repetitive regions of the genome.
The use of long reads in assembly can also establish phasing of mutations. Short Illumina reads will typically not be able to span multiple mutations on a single read. PacBio reads are long enough to enable detection of multiple mutations as coming from the same strand.
Indexing is possible with resequencing amplicons but indexes should be added during PCR and not during library preparation.
Unlike Illumina RNAseq, which requires fragmenting of the RNA, full length cDNAs can be sequenced on the PacBio Sequel. This allows direct detection of different splice isoforms. The low number of reads makes using this as a counting method challenging.
One aspect that makes PacBio sequencing unique is its ability to recognize chemical modifications of bases. Because the instrument works by measuring the time a nucleotide is in the active site, it has an axis of “time”. Basepairing modified bases has different kinetics which allows the sequencer to provide a likelihood score that any given base has been modified. Methylcytosine, Hydroxymethylcytosine, 6-methyl adenosine and many others have been detected with the RSII.