Samantha M. Hurndon Week 6
From OpenWetWare
Jump to navigationJump to search
DNA Glycosylase
Intro
- We will explore the relationship between a protein’s structure and its function in a human DNA glycosylase (hOGG1)
Methods
- I first got started by opening the program StarBiochem and opened the Sample “DNA glycosylase hOGG! w/ DNA – H. sapiens (1EBM)”
- The structure was in a ball-and-stick model, which Is the defult in biochem. This allows us to see how atoms of the structure are bonded together. Although that is a nice thing to see we want to see the space each atom fills.
- We did this by looking at the space-filled model, here size is represented of the physical space each atom occupies.
- A series of questions was completed below using the StartBiochem program (seen below)
- We then logged onto Cn3D answered a series of questions (below)
- Lastly, We were to compare the two program and decide which was better to work with.
Exercise: StarBiochem
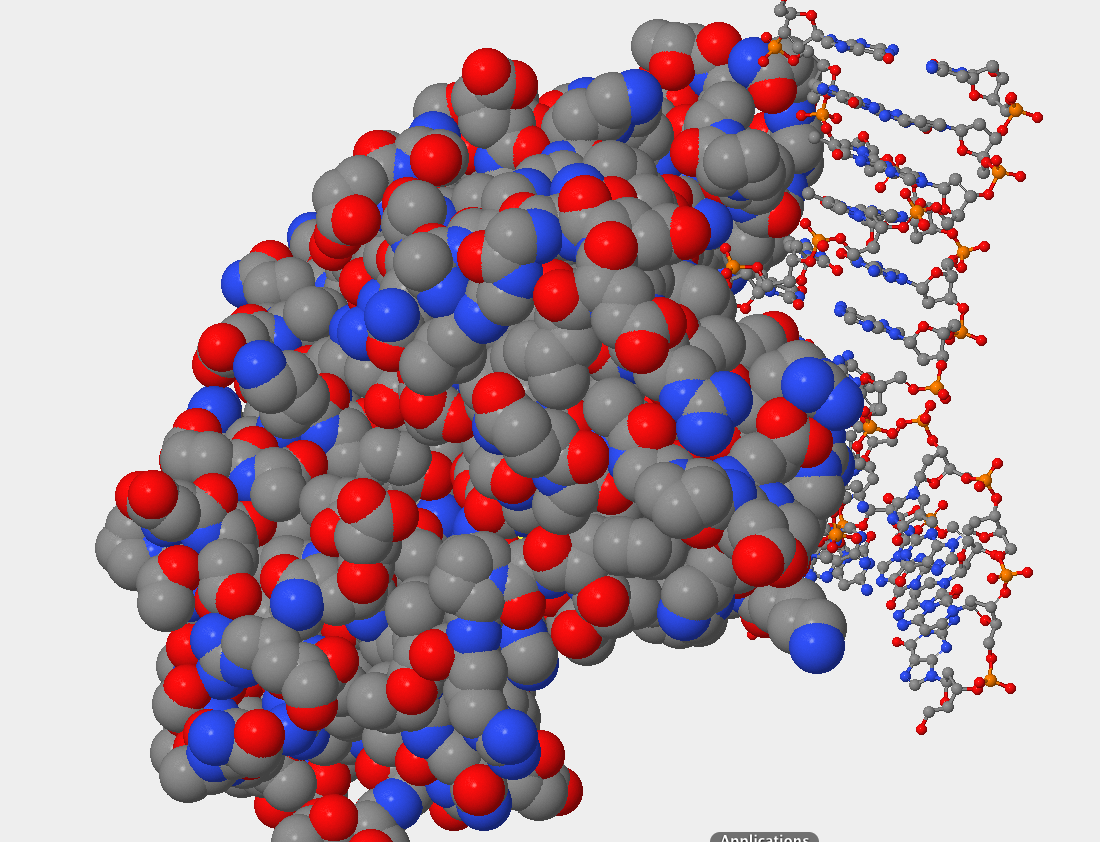
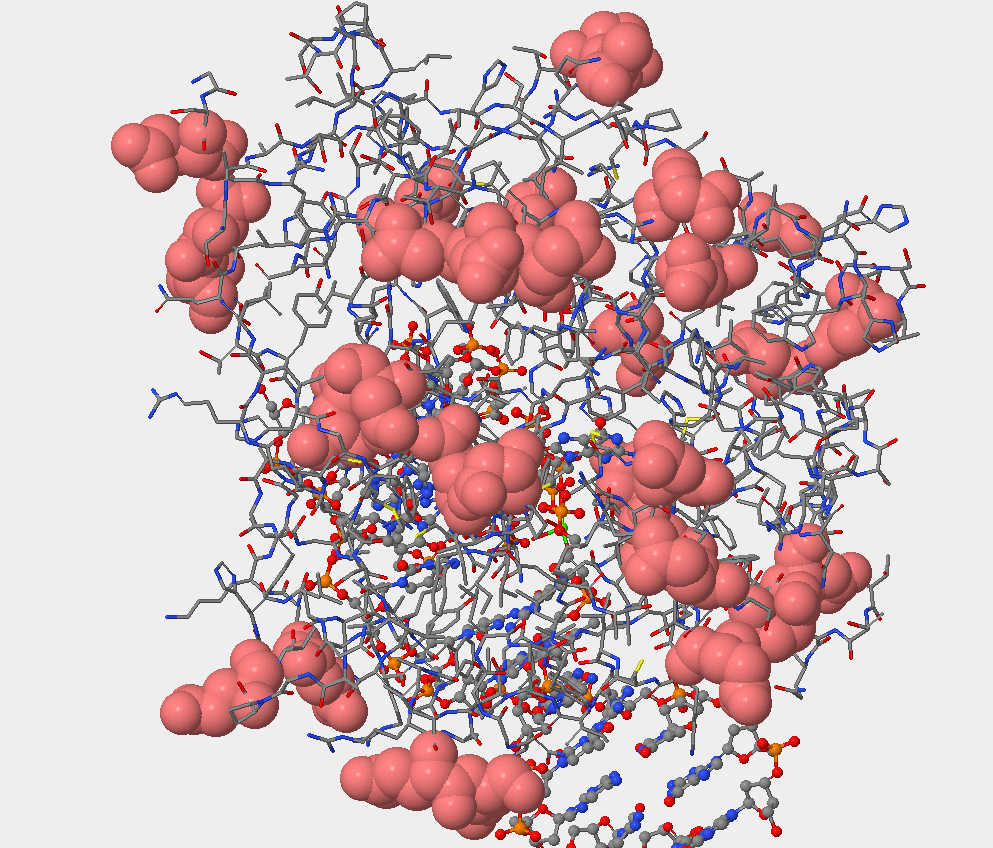
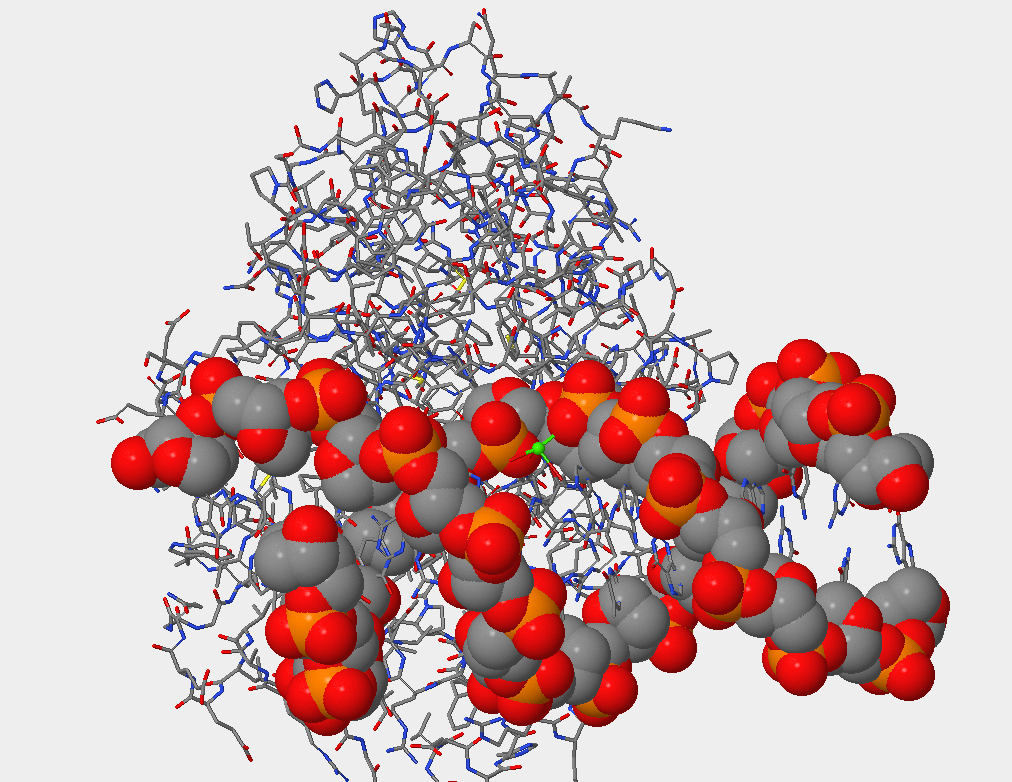
- Can you identify the DNA and the hOGG1 protein in the structure. Where is the DNA segment? In the picture below you can see that the DNA occupies less space than the protein atoms. Also, what is seen is the double helix that indicates DNA.
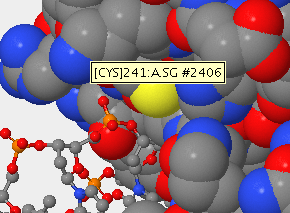
- Identify the name and sequence number of one of the amino acids in the structure that contains a sulfur atom.
- The sulfur atom is located in the side chain region of the amino acid, as seen below.
- Primary structure of the hOGG1 protein. Amino acids 105-117 complete amino acid name
- 105: Threonine
- 106: Leucine
- 107: Alanine
- 108:Glutamine
- 109: Leucine
- 110: Tyrosine
- 111: Histidine
- 112: Histidine
- 113: Tryptophan
- 114: Glycine
- 115: Serine
- 116: Valine
- 117: Aspartic acid
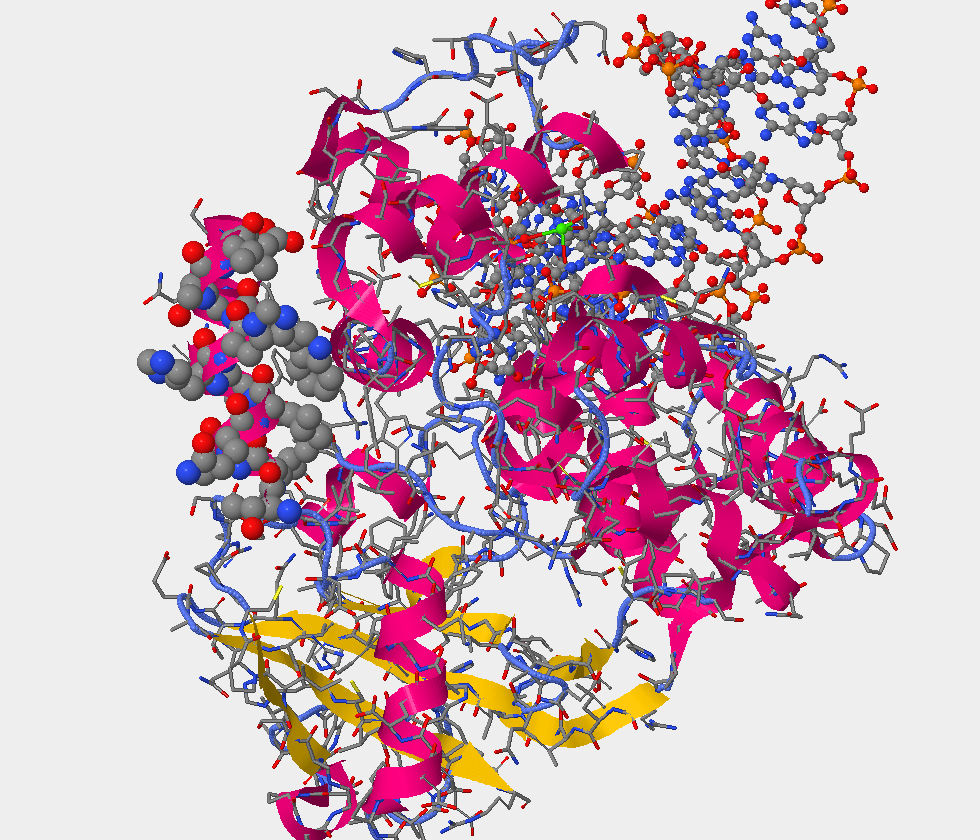
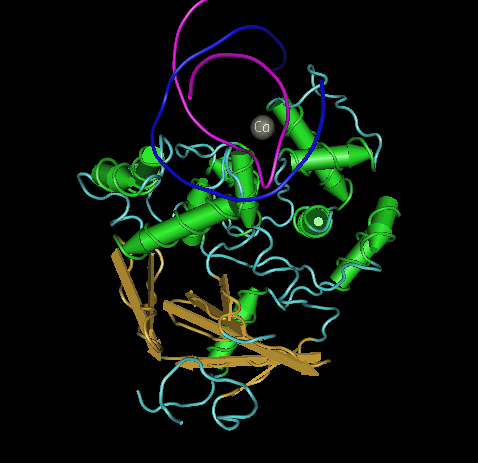
- Are helices, sheets or coils present in hOGG1? Describe the color that represents each secondary structure you observe. I observed helices seen with a pinkish color, sheets seen with a yellow color and coils were all present seen by a blue color(photo shown below)
- Which secondary structure does amino acids 105-117 fold into? They fold into a helice structure (shown below)
- Negatively charged amino acids are located on the outside (exposed) of this protein. Because it is known that negatively charged amino acids are hydrophilic, and it is observed that the negatively charged amino acids (pink color) are exposed to the outside environment, this indicates that these negatively charged amino acids wants to interact with the environment. (look at photo below)
- hOGG1’s interaction with DNA
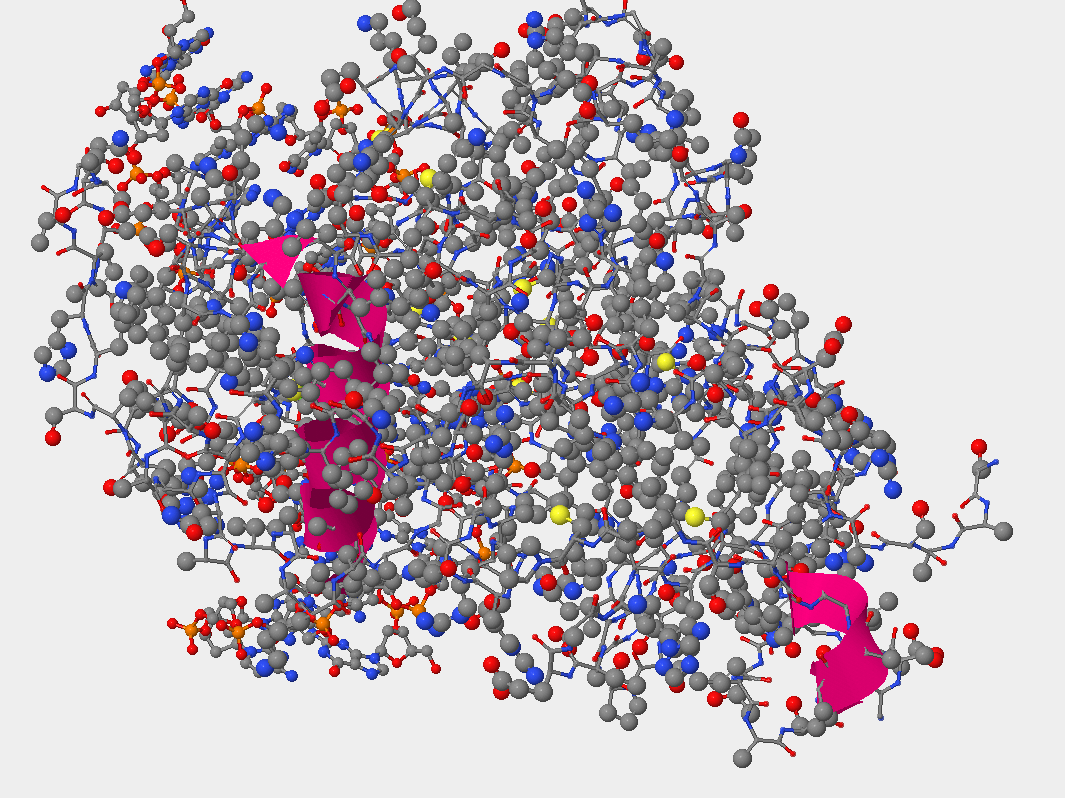
- Is helix 1 or 16 more likely to recognize damaged guanine bases? Helix 16 would be more likely to recognize damaged guanine bases for two reasons. (1) Helix 16 is much loser to the DNA helix than Helix 1. and (2)Helix 16 is near the glutamine (position 315), this position is very close to the gaunine that is damaged.
Exercise: CN3D Program
- This structure is a quaternary structure because more than one polypeptide is observed.
- DNA glycosylase has three different domains. A domain can function and exists independently from the rest of the protein chain and can often be independently stable and folded.
- I think that on starbiochem you can see much more, and you have much more options. Starbiochem shows you the list of all the amino acids. You can see the atoms in different views (ie: Ball-and-stick or space-filled view). I think starbiochem shows more and has a lot more to offer. If I am wrong and Cn3D has the same to offer, it is much more difficult to work with and I did not see the same options.
- I would choose starbiochem with no hesitation! I think that their tabs and they over set up of starbiochem is MUCH easier to follow. Also I think the program shows a better representation of the protein we were looking at, its options were much more diverse and just all around better! I really enjoyed starbiochem, I can’t say the same for Cn3D
Interesting Links
Bioinformatics Individual Journal Entries
Week 6: DNA Glycosylase Exercise
Week9: Amino Acid Sequences/Presentation
Week 10: DNA Microarray Introduction
Samantha M. Hurndon Week 11: Week 11: DNA Microarray Journal Club
Samantha M. Hurndon Week 12: DNA Microarray project
Samanhta M. Hurndon Week 14: Final Project
Class Assignments