Samantha M. Hurndon Week 10
From OpenWetWare
Jump to navigationJump to search
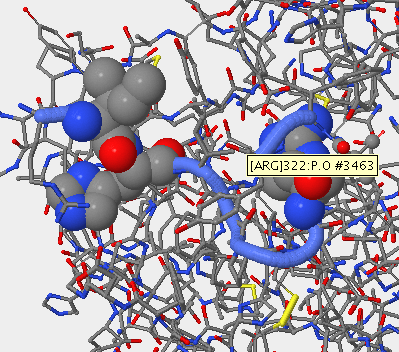
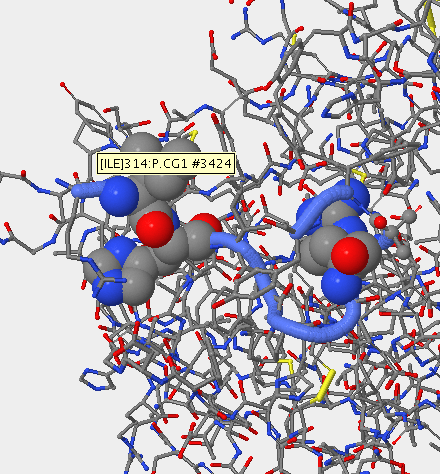
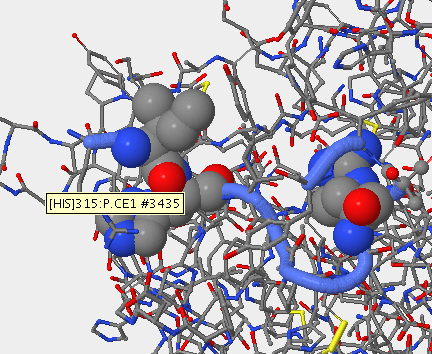
Mutation in V3 Region
- Subject 8 was sequenced and comparied to the V3 region on stanfeild's paper. The locations of the mutations were then mapped out using starbiochem.
Introduction to DNA Microarrays
- Number 5 from p. 110: Media:Number 5 p110.xlsx
- Number 6.b from p 110:
- Hour 1: X: Black Y: Black Z: Black
- Hour 3: X: Dark red Y: Red (still dark but not as dark as X) Z: Blackish-red
- Hour 5: X: Black Y: Black Z: Dark Red
- Hour 9: X: Blackish Green Y: Green Z: Dark Red
- Number 7 from pg 110: In 4.7b it looks as though up to hr 3 the genes of X, Y and Z were transcribed similarly, all having the ratios of 1.0. Beyond hour three a similarity is not seen between the three genes.
- Number 9 pg 118: Yellow indicates no change in the expression of the gene. Initially, the control and experiment have a surplus of glucose, therefore regulation can stay constant. However, when the culture grows, there is less glucose causing a need for regulation.
- Number 10 p. 118: In the experiment of TEF4, we can see that in the beginning of the experiment the TEF4 expression stayed constant. However,as time went we can see that TEF4 expression is being repressed. The cell took percausanary measures to prevent from starvation.TEF4's regulation was part of the cell's response to a reduction in available glucose because the cell probably predicted that it might starve, thus depleting the energy that would be available for translation.
- Number 11 p. 120: The TCA cycle produces energy (ATP), therefore, if is glucose is running low the cell could produce energy via TCA cycle(the cell will want to make as many ATP molecules as it can before the glucose runs out). This would cause expression of TCA genes.
- Number 12 p. 120: To ensure that genes for enzymes in a common pathway are induced or repressed simultaneously we could use the guild by association method explained on pg. 118-119. Using this method we can predict possible gene functions. Genes with similar expression profiles might have similar/the same promoters. If they do not contain the same promoter they can not be related. However, if the genes have the same promoters they have similar profiles of expression, we can use this information to test predictions. Using this, we can figure out what is being induced or repressed when we cluster the genes by using this method.
- Number 13 pg 121: If the repressor gene is deleted than glucose repressed genes would be induced. This would mean we would see Red.
- Number 14 pg. 121: When the transcription factor is overexpressed we would see red on the DNA chip
- Number 15 pg 121: I would assume that because these genes are induced we would not expect the repression of a particular gene.
- Number 16 pg 121: Control spots will not be affected by the gene that has been deleted or overexpressed. Running a microarray chip, we can verify that we have deleted or overexpressed a gene. We can verify this by making sure that the gene did not appear on the chip for deletion.
Finding a Journal Club Article/Microarray Dataset
My Partner is Isaiah M. Castaneda and we will be researching the microarray dataset from species Helicobacter Pylori. Our paper is Media:Douillard Microbiology 09 Hpylori microarray.txt