Non: Week 13
From OpenWetWare
Jump to navigationJump to search
Purpose
The purpose of this week's lab to use various tools to explore the protein structure of the SARS-CoV-2 spike protein.
Combined Methods/Results
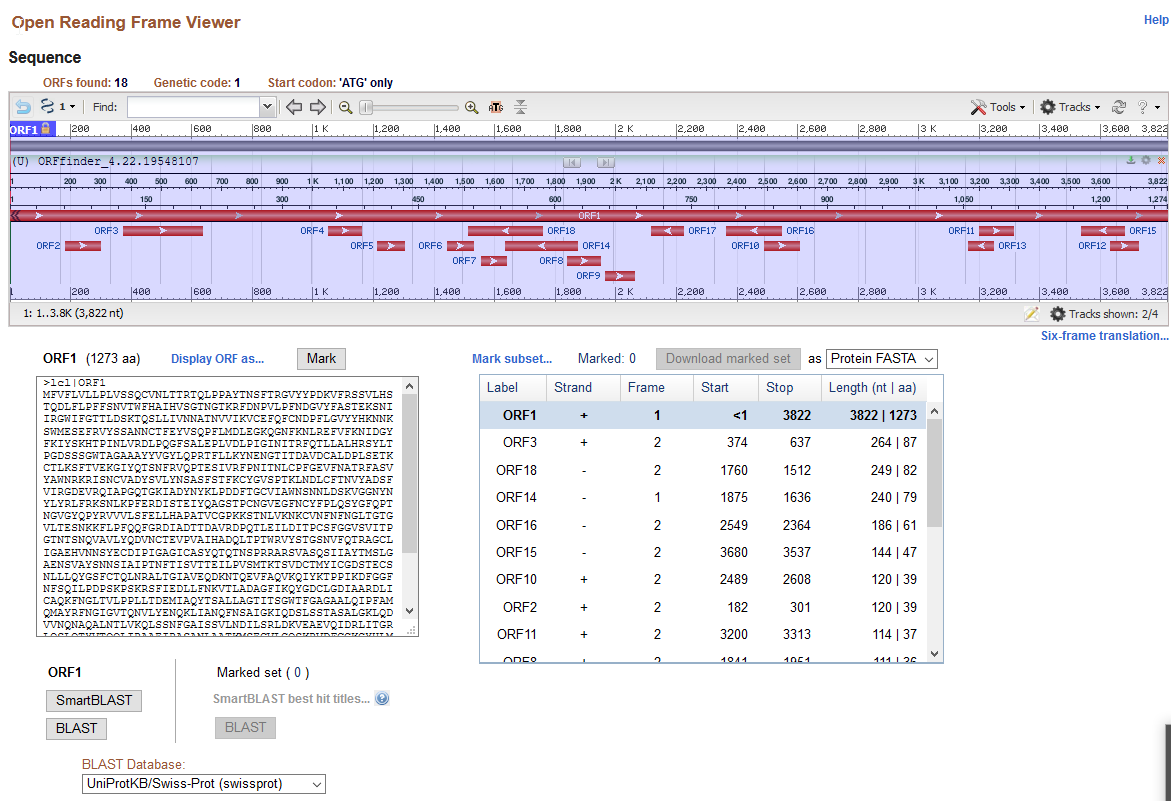
- First, I copied the FASTA sequence of the SARS-CoV-2 spike protein from the Week 13 Protocol into the NCBI Open Reading Frame Finder.
- Next I went to the UnitProt database to look up information on the SARS-CoV protein.
- Searching for "SARS-CoV" generated 833 results.
- I looked at P59594 as a reference to see what types of information could be accessed.
- For that database page, you could find info about the function, taxonomy, sub-cellular location, pathology, PTM processing, interactions, structure, family, domains, sequence, and similar proteins.
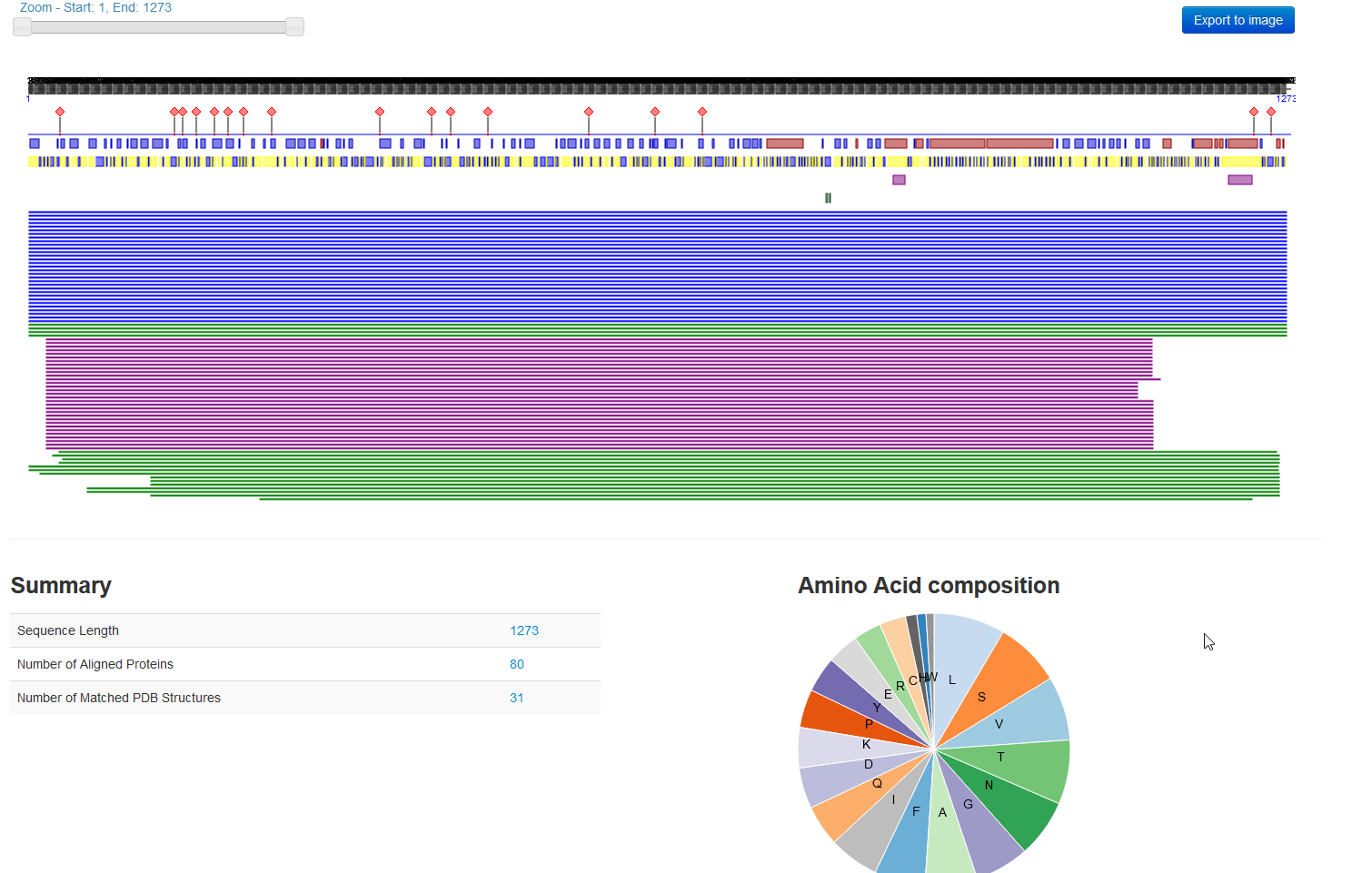
- Then, I used PredictProtein to predict the structure of the surface glycoprotein of SARS-CoV-2 mentioned in Data S1 of the Walls et al. article.
>YP_009724390.1 surface glycoprotein [Severe acute respiratory syndrome coronavirus 2] MFVFLVLLPLVSSQCVNLTTRTQLPPAYTNSFTRGVYYPDKVFRSSVLHSTQDLFLPFFSNVTWFHAIHV SGTNGTKRFDNPVLPFNDGVYFASTEKSNIIRGWIFGTTLDSKTQSLLIVNNATNVVIKVCEFQFCNDPF LGVYYHKNNKSWMESEFRVYSSANNCTFEYVSQPFLMDLEGKQGNFKNLREFVFKNIDGYFKIYSKHTPI NLVRDLPQGFSALEPLVDLPIGINITRFQTLLALHRSYLTPGDSSSGWTAGAAAYYVGYLQPRTFLLKYN ENGTITDAVDCALDPLSETKCTLKSFTVEKGIYQTSNFRVQPTESIVRFPNITNLCPFGEVFNATRFASV YAWNRKRISNCVADYSVLYNSASFSTFKCYGVSPTKLNDLCFTNVYADSFVIRGDEVRQIAPGQTGKIAD YNYKLPDDFTGCVIAWNSNNLDSKVGGNYNYLYRLFRKSNLKPFERDISTEIYQAGSTPCNGVEGFNCYF PLQSYGFQPTNGVGYQPYRVVVLSFELLHAPATVCGPKKSTNLVKNKCVNFNFNGLTGTGVLTESNKKFL PFQQFGRDIADTTDAVRDPQTLEILDITPCSFGGVSVITPGTNTSNQVAVLYQDVNCTEVPVAIHADQLT PTWRVYSTGSNVFQTRAGCLIGAEHVNNSYECDIPIGAGICASYQTQTNSPRRARSVASQSIIAYTMSLG AENSVAYSNNSIAIPTNFTISVTTEILPVSMTKTSVDCTMYICGDSTECSNLLLQYGSFCTQLNRALTGI AVEQDKNTQEVFAQVKQIYKTPPIKDFGGFNFSQILPDPSKPSKRSFIEDLLFNKVTLADAGFIKQYGDC LGDIAARDLICAQKFNGLTVLPPLLTDEMIAQYTSALLAGTITSGWTFGAGAALQIPFAMQMAYRFNGIG VTQNVLYENQKLIANQFNSAIGKIQDSLSSTASALGKLQDVVNQNAQALNTLVKQLSSNFGAISSVLNDI LSRLDKVEAEVQIDRLITGRLQSLQTYVTQQLIRAAEIRASANLAATKMSECVLGQSKRVDFCGKGYHLM SFPQSAPHGVVFLHVTYVPAQEKNFTTAPAICHDGKAHFPREGVFVSNGTHWFVTQRNFYEPQIITTDNT FVSGNCDVVIGIVNNTVYDPLQPELDSFKEELDKYFKNHTSPDVDLGDISGINASVVNIQKEIDRLNEVA KNLNESLIDLQELGKYEQYIKWPWYIWLGFIAGLIAIVMVTIMLCCMTSCCSCLKGCCSCGSCCKFDEDD SEPVLKGVKLHYT
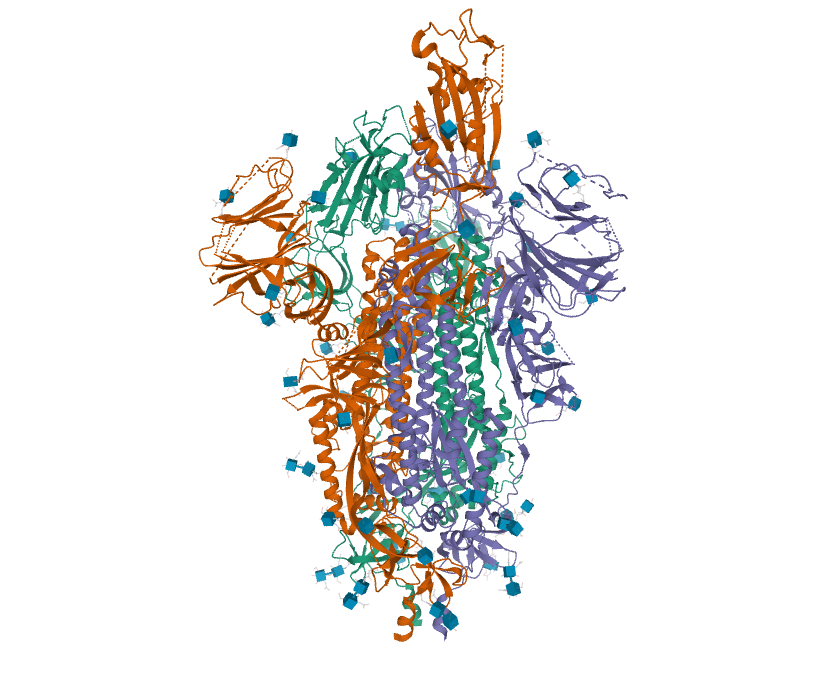
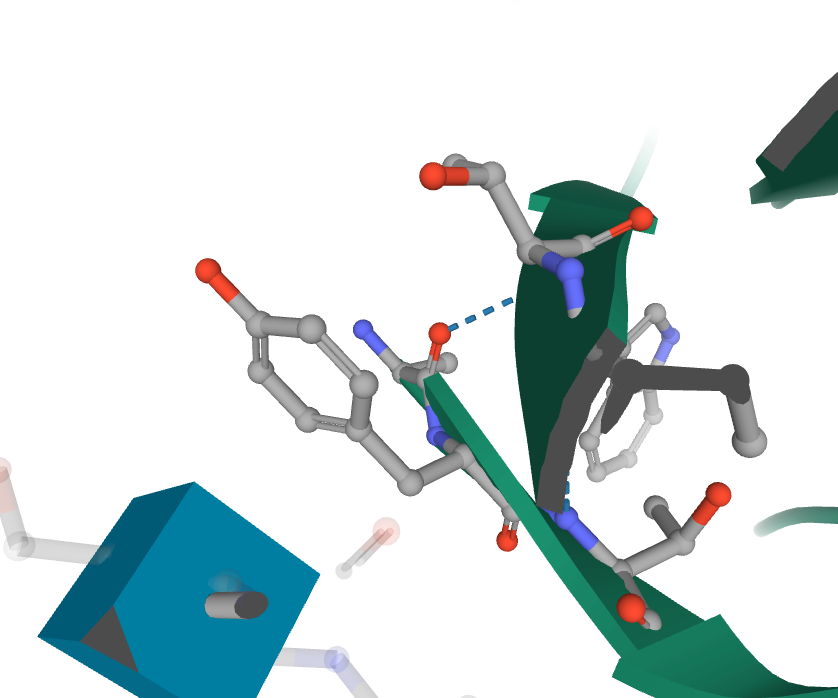
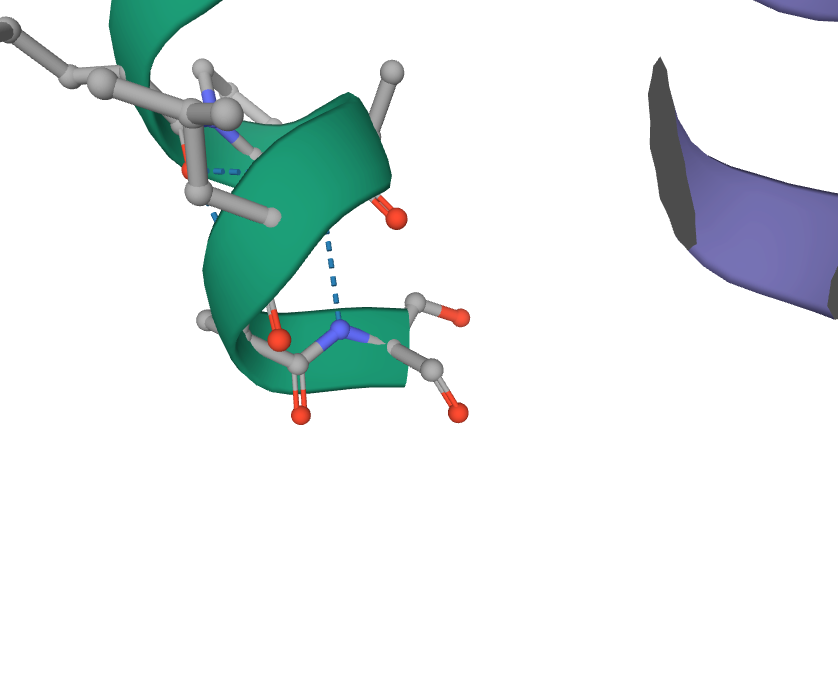
- Finally, I viewed the structure of the glycoprotein from the article, specifically 6VYB.
while the C terminus is S1146.
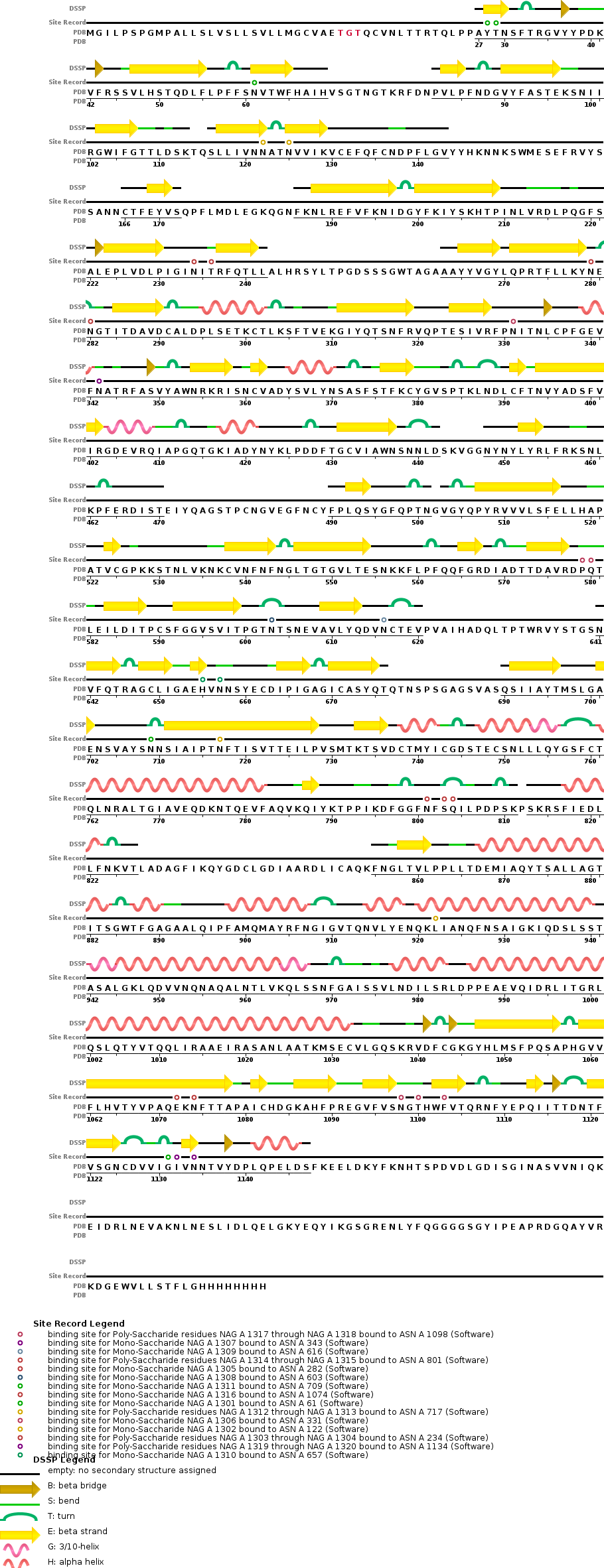
- The sequence chain image from RCSB shows all of the secondary structure features. They do not really match up with the prediction from PredictProtein.
- The article mentions 14 amino acids that are important for ACE2 binding: T402, R426, Y436, Y440, Y442, L472, N473, Y475, N479, Y484, T486, T487, G488, and Y491.
Research Question
- What question will you answer about sequence-->structure-->function relationships in a SARS-CoV-2 protein?
- What makes ACE2 the most optimal receptor for SARS-CoV-2/SARS-CoV binding?
- What sequences will you use? I want you to take advantage of sequence data available to perform a multiple sequence alignment as part of your project.
- SARS-CoV-2 spike protein; SARS-CoV spike protein;
- different ACE2 sequences, from different organisms?
- other common receptors utilized by viruses
- What protein tools will you use for analysis and answering your question?
- RCSB, UniProt, Phylogeny.fr
Data/Files
Scientific Conclusion
There are a variety of free online tools that allow for indepth analysis of protein structure and sequence. It is very hard to predict the structure of a protein.
Acknowledgements
- I worked with my partners Jenny and Carolyn in figuring out what our topic of analysis should be.
- I used the Week 13 Protocol for this assignment.
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
Non (talk) 22:35, 22 April 2020 (PDT)
References
- Walls, A. C., Park, Y. J., Tortorici, M. A., Wall, A., McGuire, A. T., & Veesler, D. (2020). Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. DOI: 10.1016/j.cell.2020.02.058
- OpenWetWare. (2020). BIOL368/S20:Week 13. Retrieved April 20, 2020, from https://openwetware.org/wiki/BIOL368/S20:Week_13
- ExPASy(2020). SIB Swiss Institute of Bioinformatics. Retrieved April 22, 2020, from https://web.expasy.org/translate/
- Cn3D. Bethesda (MD): National Library of Medicine (US), National Center for Biotechnology Information. Retrieved April 22, 2020, from http://www.ncbi.nlm.nih.gov/Structure/CN3D/cn3d.shtml
- PredictProtein (2020). RostLab. Retrieved April 22, 2020, from https://open.predictprotein.org/
- UniProtKB - P59594 (2020). National Center for Biotechnology Information (NCBI). Retrieved April 22, 2020, from https://www.uniprot.org/uniprot/P59594
- 6VYB. RSCB. Retrieved April 22, 2020, from https://www.rcsb.org/structure/6VYB