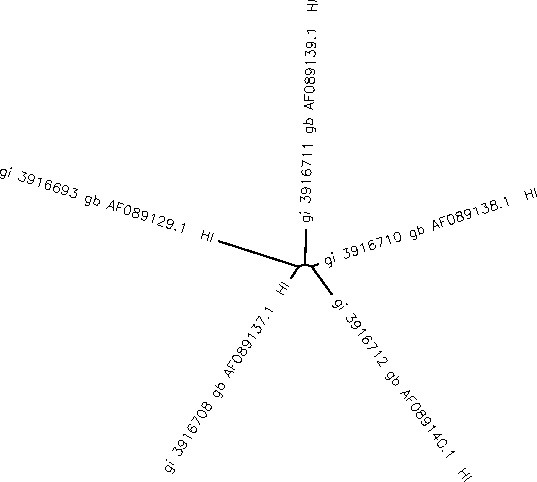Matthew R Allegretti Week 3
From OpenWetWare
Jump to navigationJump to search
Week 3 Assignment
Purpose
- The purpose of this lab is to become familiar with HIV sequence data using sequences from the envelope gene of HIV.
Methods and Results
Activity 1
Part 2
- What was the accession number of the sequence you chose?
- AF089129.1
- Which subject of the study was that HIV sequence from? Which section of the record contains information about who the HIV was collected from?
- Subject number 3, Visit 4, Clone 7. The definition section contains this information.
- Download several (4 to 6) sequences in FASTA format to your local hard drive by selecting several at the same time in the summary view so they are saved into a single text file. Be careful to remember where you put the file and what you name it so you can find it later.
- Open the file that you saved with a word processor to confirm that you have the sequences and that they are in the FASTA format. In the FATSA format each sequence is preceeded by a label which begins with the greater than sign.
- All sequences had a greater than symbol and label.
Part 3
- Using the Biology Workbench, follow the directions in part 3 of activity 1 for Exploring HIV Evolution: An Opportunity for Research in order to generate an unrooted tree for the sequences.
Data and Files
Conclusion
The purpose of this lab was to become familiar with HIV sequence data using sequences from the envelope gene of HIV. This was accomplished by accessing a number of online databases and gathering sequence data. This data was then used to generate an unrooted tree in order to interpret the sequences.
Preparation for Week 4 Journal Club
Part 1
- Seroconversion The change of a serologic test from negative to positive, indicating the development of antibodies in response to infection or immunisation.
- Divergence The evolutionary process wherein a population of species diverge into two or more descendant species, resulting in once similar or related species becoming more and more dissimilar.
- Nested PCR primers: A set of oligonucleotide primers used for the amplification of DNA by the polymerase chain reaction (PCR) in which the outermost 5′ and 3′ pair are used in the first phase of amplification and a second pair is designed to prime within that PCR product to produce a shorter amplified sequence. Greater specificity of amplification is expected from this use of two pairs of primers. (Cammack et al., 2008).
- Restriction sites The site in a polynucleotide chain as to where the restriction enzyme cleaves nucleotides by hydrolyzing the phosphodiester bond between them.
- Synonymous Having the character of a synonym; expressing the same thing; conveying the same, or approximately the same, idea.
- Plasma Viral Load The number of viral particles (usually hIV) in a sample of blood plasma. HIV viral load is increasingly employed as a surrogate marker for disease progression. It is measured by pCR and bDNA tests and is expressed in number of HIV copies or equivalents per millilitre.
- CD4 T-cells A form of T lymphocyte with CD4 receptor on the cell surface that recognizes antigens of a virus-infected cell.
- Epitope That part of an antigenic molecule to which the t-cell receptor responds, a site on a large molecule against which an antibody will be produced and to which it will bind.
- Frequency-dependent selection: A form of natural selection in which the fitness of a particular genotype depends on its frequency in the population. See also apostatic selection, stabilizing selection. (Cammack et al., 2008).
- PBMC: Peripheral blood mononuclear cells. The mononuclear cells of the blood: monocytes and lymphocytes. (Cammack et al., 2008).
Part 2
Abstract
- Studying evolution of HIV-1 envelope sequences in 15 seroconverting drug users
- Disease progression seems to be linked to strains whose evolution selected for nonsynonymous mutations
- In the nonprogression group, his evolution may have been selected against for higher replication
- Both quantity and scope of mutations have an effect on adaptation to a host environment
Introduction
- HIV has high replication and mutation rate
- Make it suitable for adjusting to a host environment
- In a stable host environment, scientists expect a "best fit" virus would be the dominant strain present initially, and all future strains will be less represented
- In an unstable environment, genetic diversity is expected to be substantially higher
- Immune response targets the most abundant viral variety
- May lead to a substantial drop in the viral load, but not the viral diversity, since less abundant strains contribute most to diversity
- Numerous small and diverse populations lead to further mutation, and cascade effect of variability
- Too many varieties has been proposed as the source of overwhelming the immune response
- Too many changes for the immune system to keep up with
- This is why the study focuses on diversity during HIV-1 evolution
- Why this study is different/significant
- Larger cohorts
- Direct examination of sequence patterns not done previously
- Analyzing more time points
- Different patterns of selection are shown in nonprogressing and progressing varieties
- Main point: High genetic diversity responsible for CD4 T-Cell decline
Methods
- Study population
- 15 injection drug users in the AIDS Linked to Intravenous Experiences study
- 6 month intervals
- Subsets of population
- Rapid progressors: fewer than 200 CD4 T cells within 2 years of seroconversion
- Moderate progressors: CD4 T cell levels that declined to 200–650 during the 4-year period of observation
- Nonprogressors: maintained CD4 T cell levels above 650 throughout the observation period
- Sequencing
- Nested PCR used to amplify 285 base pair region of envelope gene
- Obtained from PBMC infected cells
- Plasma viral load determined by reverse transcription PCR
- Phylogenetic trees were constructed
- Analysis performed to test for relationship between diversity and CD4 count
- Strain sequences analyzed to determine amount of difference between strains
- Figure 1 shows the diversity, divergence, and CD4 counts for all subjects throughout the study period
- Subject 9 and 15 were examined to see if two different viruses were contracted
- Table 1 summarizes progression of each subject throughout the study, shows trends for each group.
- Compared rate of change for divergence and diversity
- Figure 2 provides a mean lope per year comparison among progressor groups
- Figure 3 is the phylogenetic tree generated by the study
Results
- Variable patterns of CD4 decline
- Viral load was also highly variable for viral genomic RNA/mL
- Nonprogressors tended to have lower viral loads
- 873 clones were sequenced and analyzed
- Changes quantified by genetic diversity and divergence at each visit
- Change in diversity: 22.94 to 5.10 nt per clone per year
- Change in divergence: 0.13% to 2.09% of the nucleotides per clone per year
- Viruses were initially homogenous, which was consistent with previous reports
- Diversity and divergence increased over time in all three categories
- Diversity and divergence showed strong negative correlation to CD4 cell count
- Strong relationship between selection for nonsynonymous changes and decline in CD4 count
- Phylogenetic trees showed no predominance of a single strain for an extended period of time
- Figure 4 shows phylogenetic trees for a number of individuals from the cohorts
- Demonstrates differences in evolution of the virus in each subject
Discussion
- Higher level of diversity and divergence attributed to lower CD4 cell count
- subjects with similar CD4 counts and higher diversity showed a greater decline after 12 months
- Nonsynonymous substitution rates three times higher in progressors than in nonprogressors
- One previous study supported that higher diversity was linked to greater decline in CD4, but with fewer time points
- Another study showed inconclusive results when linking diversity to decline rates
- Subjects may have had already compromised immune systems in some cases
- Nonsynonymous substitutions might be selected for in strains that are already difficult for the host immune system to target
Evaluation
- The results of the study support the reality that HIV is incredibly difficult to combat, in part because of its massive variability and ability to evolve rapidly
- This study may be limited by its age. It is almost 20 years old, and many developments in both technology and understanding of HIV may have been capable of further developing, supporting or refuting, the information put forth by this study.
- I would suggest future studies to use modern sequencing technology to gather substantially more frequent sequencing information from a larger population of subjects. This would serve to observe more closely how rapidly evolution is occurring, as well as being able to establish a more consistent correlation between diversity and CD4 decline rates.
Acknowledgments
While I worked with the people noted above, this individual journal entry was completed by me and not copied from another source.
References
- Week 3 Instructions
- National Center for Biotechnology Information
- Donovan, S., & Weisstein, A. E. (2003). Exploring HIV evolution: An opportunity for research. Stanley (Eds.), Microbes Count, 137-148.
- Biology Workbench
- Biology Online Dictionary
- Markham, R. B., Wang, W. C., Weisstein, A. E., Wang, Z., Munoz, A., Templeton, A., ... & Yu, X. F. (1998). Patterns of HIV-1 evolution in individuals with differing rates of CD4 T cell decline. Proceedings of the National Academy of Sciences, 95(21), 12568-12573. Retrieved from http://bioquest.org/bedrock/problem_spaces/hiv/Markham_1998.pdf
- Cammack, R., Atwood, T., Campbell, P., Parish, H., Smith, A., Vella, F., & Stirling, J.(Eds.), (2006). Oxford Dictionary of Biochemistry and Molecular Biology. : Oxford University Press. Retrieved 19 Sep. 2016, from http://www.oxfordreference.com/view/10.1093/acref/9780198529170.001.0001/acref-9780198529170.
Useful links
- Stem Cell Overview
- Template
- Student Page
- Week 1 Instructions
- Journal Week 1
- Week 2 Instructions
- Week 2
- Journal Week 2
- Week 3 Instructions
- Week 3
- Journal Week 3
- Week 4 Instructions
- Week 4
- Journal Week 4
- Week 5 Instructions
- Week 5
- Journal Week 5
- Week 6 Instructions
- Week 6
- Journal Week 6
- Week 7 Instructions
- Journal Week 7
- Week 8 Instructions
- Week 8
- Journal Week 8
- Week 9 Instructions
- Week 9
- Journal Week 9
- Week 10 Instructions
- Week 10
- Journal Week 10
- Week 11 Instructions
- Week 11
- Journal Week 11
- Week 12 Instructions
- Week 14 Instructions
- Week 14 Assignment Part 1 and 2
- Journal Week 14
- Week 15 Instructions
- Week 15
- Journal Week 15
- My user page