Jennymchua Week 10 Assignment
Purpose
The purpose of this assignment is to use the questions assigned as a guideline on evlauating biological databases.
Database Evaluation
General information about the database
What is the name of the database?
The database is called ProteomicsDB.
What type (or types) of database is it?
- What biological information (type of data) does it contain? (sequence, structure, model organism, or specialty [what?])
- ProteomicsDB contains sequence coverage, a protease map, information on the peptides that make a given protein, proteotypicity (the likelihood of whetehr or not a peptide is uniquely identifiable in mass spectroscopy), reference peptides, false discovery rate (statistical analysis), expression in a given organism, biochemical assays, interaction networks, and current projects using that protein.
- Important to note that ProteomicsDB has information available for Homo sapiens and Arabidopsis thaliana.
- What type of data source does it have?
- It is a secondary data source.
- It is a community, human, and curated database.
What individual or organization maintains the database?
ProteomicsDB is maintained by a group of scientists and software developers from the Proteomics and Bioanalytics department at Technische Universität München (TUM) and Cellzome GmbH, which is part of a larger pharmaceutical company based in Great Britain. TUM is a public research institution and Cellzome GmbH is monitored by GlaxoSmithKline, Inc. which gets its money from investors, income, etc.
Scientific quality of the database
Does the content appear to completely cover its content domain?
The content appears to near-completely cover its content domain. Proteomics is the study of the set of proteins expressed in an organism, and the database (as previously answered) has sequences, maps, related content, and even further information available to search.
- How many records does the database contain?
- Two organisms, Homo sapiens and Arabidopsis thaliana.
- Homo sapiens
- 79% of total proteome
- Arabidopsis thaliana
- 71% of total proteome
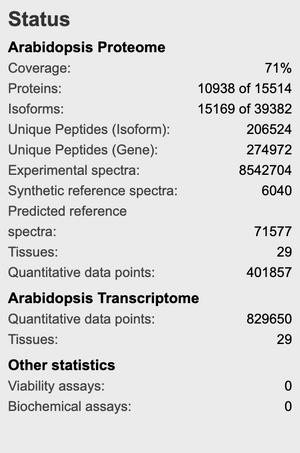
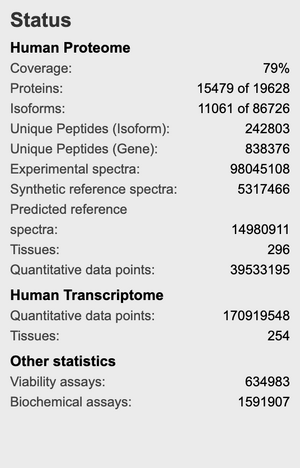
- What claims do the database owners make about coverage in the corresponding paper?
- The claims are based on attempting to "enrich the data" by adding additional datasets related to current proteome data and new services to allow for a community-based aspect in allowing user to upload and analyze their own datasets.
What species are covered in the database?
Homo sapiens and Arabidopsis thaliana.
Is the database content useful? I.e., what biological questions can it be used to answer?
The purpose of ProteomicsDB is to visualize large collections of proteomics data; for example, real-time exploration of protein abundance across different tissues in a human.
Is the database content timely?
- Is there a need in the scientific world for such a database at this time?
- I think so. Samaras et al. in Nucleic Acids Research claim "a wide range of drug-target interaction data can be visualized in ProteomicsDB as well, which enables the exploration of combination treatments in a dose-dependent protein-drug interaction graph in-silico," so this dataset could definitely be helpful in future pharmaceutical trials.
- Is the content covered by other databases already?
- There appears to be other proteomics databases, but perhaps not as comprehensive with the amount of categories ProteomicsDB has. Though, other databases have more species of mammals.
How current is the database?
- When did the database first go online?
- May 29, 2014.
- How often is the database updated?
- Unclear, mentioned in the Nucleic Acids Research article that it is "continuously updated."
- When was the last update?
- Last update in Nucleic Acid Research was in 2017.
General utility of the database to the scientific community
Are there links to other databases? Which ones?
No other databases linked.
Is it convenient to browse the data?
Not really. One has to search for a specific protein in either a human or Thale cress and the opening page when one attempts to search a protein or peptide says in the center of the page "No data" because nothing was previously searched. It would be better if all the options were listed and then narrowed down as the user typed or searched.
Is it convenient to download the data?
Yes, once one searches for a specific protein, for example hemoglobin, there is a one-click button to download all related data.
- In what file formats are the data provided?
- CSV files.
- Are they standard or non-standard formats?
Standard.
Evaluate the “user-friendliness” of the database: can a naive user quickly navigate the website and gather useful information?
I don't think so. Perhaps because the website is based in Germany, it was a bit slow to load on my home WiFi, so that's one of the first setbacks. As mentioned earlier, no data (or even category options for data) is presented when one first clicks onto the "Proteins" tab, which feels incomplete for someone who is new to evaluating and navigating through databases.
- Is the website well-organized?
- Yes, however a good portion of the information is "unclickable," meaning they are just words and not hyperlinks to information because they are "Coming Soon." This can be a bit offputting.
- Does it have a help section or tutorial?
- No! And this is where I think ProteomicsDB can benefit from the most!
- Are the search options sensible?
- Yes, when one searches for a protein, almost all of the information (or where to find it) is presented.
- Run a sample query. Do the results make sense?
- I first clicked on the Homo sapiens tab on the left side, then I clicked on the "Protein" tab to search for hemoglobin. I clicked on the "Hemoglobin subunit gamma 2" link, and I was presented with tons of information about its expression, sequence, biochemical assays, reference peptides, etc. The results do make sense!
Is there a license agreement or any restrictions on access to the database?
There is a tab specifically for Terms of Use, and there doesn't seem to be any restrictions besides the usual redistribution, copying, or modification without permission violation.
Summary judgement
Would you direct a colleague unfamiliar with the field to use it? Yes and no. Yes because I think it is the most detailed, accessible, and organized proteomics database, but there is also a lot unfinished (yet still advertised) and finding the information can be tricky at first.
Is this a professional or "hobby" database? The "hobby" analogy means that it was that person's hobby to make the database. It could mean that it is limited in scope, done by one or a few persons, and seems amateur. This is a professional database, though I'm sure the researchers who manage it definitely consider it a passion project, as I had never heard of proteomics (or much of the other subcategories they offer on the database). While it does not seem amateur, it does seem niche in the sense that it's not a field that gets much mainstream attention.
Conclusion
ProteomicsDB claims to "expedite the identification of various proteomes and their use across the scientific community." While it offers a lot of information with regards to protein subunits, there are only two organisms as a whole, and I'm not sure what the benefit of having a whole database for just those two are. With that being said, once one searches a protein, it's fairly easy to access the information, but figuring out how to get there can be a bit confusing and off-putting. There is a lot of "Coming Soon" information which is promising, but what's the point of advertising it on a main page if it's not there? Overall, I think in the coming years, ProteomicsDB has the potential to be usable in a university class, but it still seems rather new and unfinished.
Acknowledgements
- Except for what is noted above, this individual journal entry was completed by me and not copied from another source.
Jennymchua (talk) 22:25, 31 March 2020 (PDT)
References
- OpenWetWare. (2020). BIOL368/S20:Week 10. Retrieved March 31, 2020, from https://openwetware.org/wiki/BIOL368/S20:Week_10.
- ProteomicsDB. (2019). ProteomicsDB. Retrieved March 31, 2020, from https://www.proteomicsdb.org/proteomicsdb/#overview.
- Samaras, P., Schmidt, T., Frejno, M., Gessulat, S., Reinecke, M., Jarzab, A., ... & Aiche, S. (2020). ProteomicsDB: a multi-omics and multi-organism resource for life science research. Nucleic acids research, 48(D1), D1153-D1163.
My user page
Template link
Class Assignments
- BIOL368/S20:Week 1
- BIOL368/S20:Week 2
- BIOL368/S20:Week 3
- BIOL368/S20:Week 4
- BIOL368/S20:Week 5
- BIOL368/S20:Week 6
- BIOL368/S20:Week 8
- BIOL368/S20:Week 9
- BIOL368/S20:Week 10
- BIOL368/S20:Week 11
- BIOL368/S20:Week 13
- BIOL368/S20:Week 14
Weekly Assignments
- jennymchua Week 2 Assignment
- jennymchua Week 3 Assignment
- jennymchua Week 4 Assignment
- jennymchua Week 5 Assignment
- jennymchua Week 6 Assignment
- jennymchua Week 7 Assignment
- jennymchua Week 8 Assignment
- jennymchua Week 10 Assignment
- jennymchua Week 11 Assignment
- jennymchua Week 13 Assignment
- jennymchua Week 14 Assignment
Class Journals
- BIOL368/S20:Class Journal Week 1
- BIOL368/S20:Class Journal Week 2
- BIOL368/S20:Class Journal Week 3
- BIOL368/S20:Class Journal Week 4
- BIOL368/S20:Class Journal Week 5
- BIOL368/S20:Class Journal Week 6
- BIOL368/S20:Class Journal Week 8
- BIOL368/S20:Class Journal Week 9
- BIOL368/S20:Class Journal Week 10
- BIOL368/S20:Class Journal Week 11
- BIOL368/S20:Class Journal Week 13
- BIOL368/S20:Class Journal Week 14