BioMicroCenter:Multiplex
| HOME -- | SEQUENCING -- | LIBRARY PREP -- | HIGH-THROUGHPUT -- | COMPUTING -- | DATA MANAGEMENT -- | OTHER TECHNOLOGY |
MULTIPLEX BARCODING
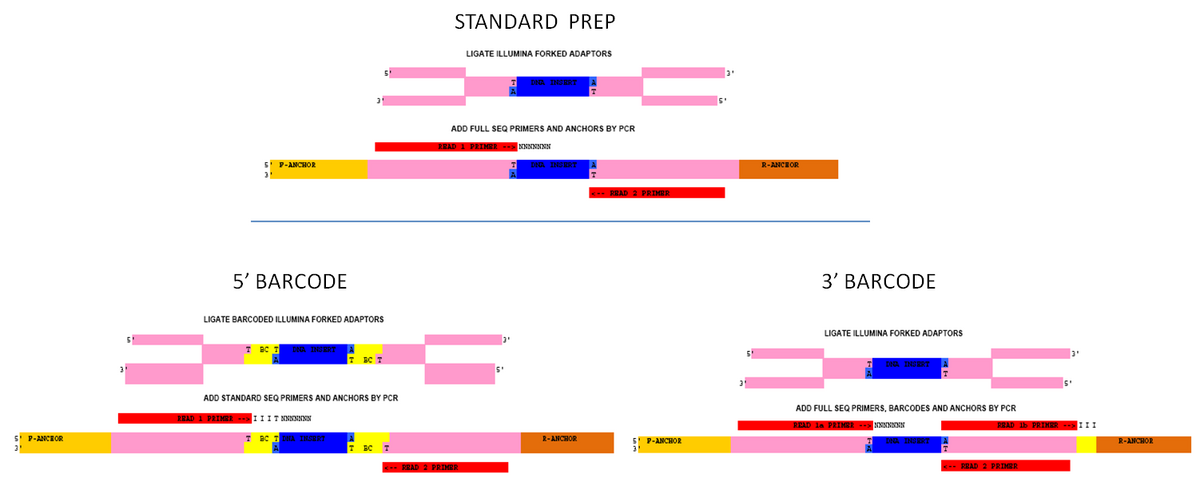
Multiplexing methods allow for running multiple samples in a single Illumina lane. There are multiple strategies for multiplexing Illumina samples but they can generally be broken down into two classes: 5' barcodes and 3' barcodes.
The main advantage of 5' barcodes is the fact that they are read as the first part of read 1, eliminating the need for a separate barcode reprime. A disadvantage is that sample prep will require two different primers (both halves of the Y-shaped adapter). Also, because the cluster-calling algorithms require a balanced base composition over the first few nucleotides, you must use a large number of barcodes in each run (16+). In addition, because the barcodes are present during the ligation reaction, care must be taken to prevent them from introducing biases in library construction.
3' Barcodes are more flexible as they are added during the PCR step, can have a number of different compositions, and can be run in any number. They do require a separate barcode read to sequence, which can add extra cost to the flowcell.
MULTIPLEXING AT BIOMICRO
The BioMicro Center uses 3' barcoding for our sample preparation. Barcoding is available for FREE as part of our BMC Fragmented DNA sample prep (SPRI and enrichment) service and is done during the amplification step. The barcodes we use are 6nt in length and each is differentiated by at least 2nt from any other barcode. We currently have a stock of 40 barcodes and have designs available for over 100.
Reading BMC barcodes requires a re-prime of the sequencer and there are some additional charges for multiplexing samples. A full list of our prices is here.