20.109(S15):Module 1
Module 1: DNA Foundations and Diagnostic Engineering
Lecturer: Jonathan Runstadler
Instructors: Shannon Hughes, Noreen Lyell, and Leslie McClain
TA: [[User: |Djenet Bousbaine]]
Overview
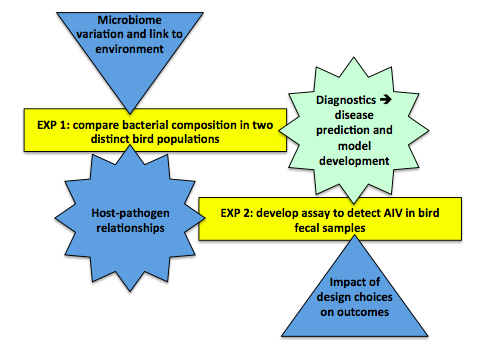
In this module, you'll complete two mini-investigations while gaining foundational skills – in laboratory techniques, data analysis, and both written and oral communication – that will serve you well in the remaining two modules. Throughout, host–microbe interactions and their implications for human health will be a unifying theme. You will study two classes of microbes that can be present in the bird gut, whether by ingestion or infection. Some of these microbes are pathogenic (disease-causing), and some may have the potential for zoonotic (inter-species) transfer to humans.
In the primary experiment, you will pool data with your peers to perform a phylogenetic analysis of the bacteria found in two distinct populations of migratory birds. You will look for similarities and differences and speculate about the mechanisms that brought these about. This study has parallels to recent investigations of human microbiome diversity that shed light on variations in metabolism, susceptibility to infection, and other measures of health. More directly, your results may suggest a link between a particular factor in a bird's environment, such as climate or diet, and characteristics of the resulting gut bacterial population, such as the dominant species or degree of variety. And hey, we'll also learn what bacteria you might be exposed to the next time you accidentally encounter bird feces!
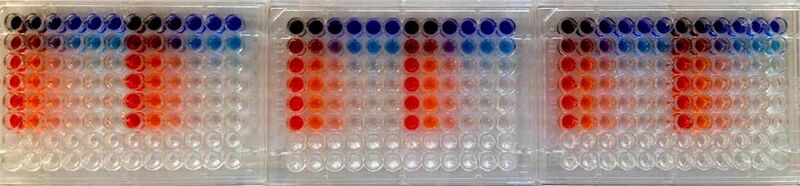
In the secondary experiment, you will develop an assay to screen bird cloacal samples for avian influenza virus (AIV). You will examine the sequence of a highly conserved gene in the AIV genome and design primers that target this gene. Your primers will be used in a real-time reverse transcription-polymerase chain reaction (RRT-PCR) to determine the presence or absence of AIV in three bird samples. The sensitivity of your primers will be compared to that of primers currently used in the Rundstadler Lab for AIV screening.
We thank the Runstadler lab for access to bird samples (pre-screened for flu and everything!), and especially Wendy Puryear for helpful technical support and discussions as this module was developed.
Lab Links
Day 1: Microbial DNA extraction
Day 2: Diagnostic primer design
Day 3: 16S PCR and paper discussion
Note: 1 week between Day 3 and Day 4.
Day 4: DNA cloning
Day 5: DNA sequencing
Day 6: Journal club I
Day 7: Phylogenetic analyses
Day 8: Diagnostic primer analysis
Day 9: Journal club II
Assignments
Microbiome data summary
Primer design memo
Journal club Presentation
20.109 Blog summary