20.109(S15):Diagnostic primer design (Day2)
Introduction
Today you will begin the second research project for Module 1. Specifically, you will design primers to test bird fecal samples for avian influenza virus (AIV) using real-time reverse transcription-polymerase chain reaction (RRT-PCR). AIV is found in several bird taxa; however, it is most consistently associated with waterfowl, gulls, and shorebirds. These birds are considered to be natural maintenance hosts of AIV, meaning that they have adapted to carry and transmit influenza strains. AIV can spread to domestic poultry and cause widespread outbreaks that result in serious disease. Furthermore, some AIVs have crossed the species barrier and caused disease in humans and other mammals. An example of an AIV crossing species barriers is the H5N1 subtype, which first infected humans in 1997 during a poultry outbreak. In fact, over the last century AIVs carried by wild birds have been implicated in the three major influenza pandemics. A better understanding of AIV abundance and epizootiology in wild birds will aid public health officials in the development of models that better predict human influenza pandemics.
To screen wild birds for AIV, embryonating chicken eggs can be used to incubate virus present in samples collected from birds. Following incubation, the viral genetic material is isolated and screened to identify AIV infected birds. Though this method is very sensitive, results may not be obtained for 1-2 weeks. To quicken the identification and screening process RRT-PCR was developed as a rapid and less expensive alternative. We will discuss this method in further detail later in this Module. For today, your goal is to design the tools you will need for your RRT-PCR assay.
To examine the prevalence of AIV in wild birds in New England, you will screen genetic material isolated from fecal samples collected from a ‘scallop pile’ in Nantucket. In both projects for Module 1, you will use primers to amplify DNA. The primers you will use next time anneal to bacterial 16S sequences and will allow you to amplify this section of the genome for cloning. For this project, you will design primers that anneal to the matrix (M1) gene within the influenza A genome to enable the identification of AIV-infected birds.
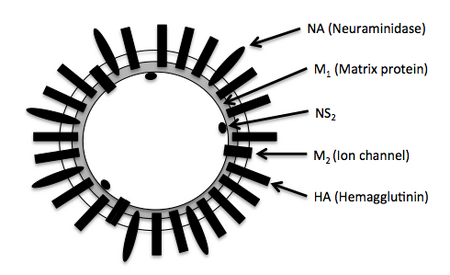
Influenza A viral genomes are composed of eight segments of negative-sense, single-stranded RNA that encodes 11 proteins. Segments 1, 2, and 3 encode subunits of RNA polymerase. Hemagluttinin (HA) is encoded on segment 4. A nucleoprotein (NP) is encoded on segment 5. Neuraminidase (NA) is encoded on segment 6. Two matrix proteins (M1 and M2) are encoded on RNA segment 7 from different reading frames. Segment 8 encodes two non-structural proteins (NS1 and NS2) from different reading frames. Subtypes of influenza A are categorized according to the HA and NA proteins, which are displayed on the surface of the viral envelope. There are 16 subtypes of HA and 9 subtypes of NA. In influenza A H5N1, the viral genome encodes HA subtype 5 and NA subtype 1. Because the M1 gene is less variable, it is a better target to use for your assay.
Today you will begin the process of developing an assay that can be used to identify AIV. The first step is to design primers that are specific for M1. Primers are short, single-stranded oligonucleotides that anneal to specific sequences of DNA or RNA. These small oligonucleotides can be generated to include any sequence and therefore can be designed to anneal to single location within the genome of an organism. Thus, primers are very useful tools for developing diagnostic assays that identify certain sequences of DNA or RNA within a sample.
In diagnostics, a two-stage identification system is often used. First, a quick yes/no screen is completed -- is the organism there? In this stage, sensitivity is prioritized: one would like as few false negatives as possible, even if that means some false positives come up, because the latter can always be excluded in the second stage. Next, the specific species or subtype is identified. In this stage, specificity is prioritized. The primers you design will screen for the presence/absence of AIV. The sensitivity of your primers will be compared to those currently used in the Runstadler Lab.
Protocols
Part 1: Explore existing AIV screening primers
The reference primer set that you will use to amplify M1 of avian influenza A comes from a 2007 paper by Jon Runstadler. (link).
| Primer name | Direction | Sequence (5'-->3') |
|---|---|---|
| Avian Influenza A Matrix Forward | Forward | ARATGAGTCTTGTRACCGAGGTCG |
| Avian Influenza A Matrix Reverse | Reverse | TGCAAAGACATCYTCAAGYYTCTG |
First, let's assess some basic properties of these primers using two online tools.
Primer properties
Integrated DNA Technologies, or IDT, sells DNA primers and offers free tools for analyzing them.
You'll also access the National Center for Biotechnology Information (NCBI), a resource that we have used in many different 20.109 modules.
- At the IDT website, under the heading Tools, choose OligoAnalyzer.
- Copy the Avian Influenza A Matrix Forward primer sequence into the “Sequence” box.
- Keeping the default settings, press the Analyze button and note the resulting length, G/C content, and melting temperature (Tm) of each primer.
- Based on our discussion so far of PCR, are the lengths of forward and reverse primers sufficient for a reasonable expectation of sequence uniqueness?
- Are the G/C contents in the appropriate range?
- Complete a self-dimer analysis on each primer and a hetero-dimer analysis of the two primers together.
- It should not be energetically favorable for the primer to dimerize. The delta G for dimers should at least be greater than -9 , and ideally be greater than -5 kcal/mol.
- Annealing of one primer to the 3' end of another primer (or itself) is particularly bad. Why?
- Having a G or C (a "GC clamp") at the 3' end of each primer helps promote complete binding due to the relatively greater hydrogen bonding for GC versus AT. Note that 1-2 bases is best, while 3 or more consecutive GC bases might promote non-specific binding. Do the primers above have such clamps? Why is effective binding at the 3' end (versus 5' end) particularly important?
- It's best to avoid long stretches of a single base, or even a single type of base (purine or pyrimidine). Are the primers above relatively random or do they break this rule?
- Finally, you should to test whether these primers bind to a unique site within the avian influenza A genome or instead are promiscuous binders.
- Go to nucleotide BLAST and copy your first primer into the "Query" box.
- Under Algorithm Parameters, be sure that "Automatically adjust parameters for short input sequences" is checked (the Algorithm Parameters link is located at the bottom of the page).
- Click BLAST and wait for your results.
- What are the top three matches for the Avian Influenza A Matrix Forward primer? Are the matches to matrix genes? From what animals were the sequences obtained? What are the AIV subtypes for the top three matches?
- Repeat the above steps for the reverse primer.
- What nucleotides are represented by R in the forward primer sequence? By Y in the reverse primer? Why might you want to include these in the primers you design for your AIV screening assay?
- When you design your own primers below, you will perform all these same checks.
Part 2: Design novel AIV matrix gene-specific primers
As discussed above, diagnostic assays often use a two-step process to screen samples. Professor Runstadler takes a very similar approach to flu screening in his lab. All bird samples are broadly checked for flu infection using primers that target the highly conserved matrix gene of the virus. Samples that come up positive are screened for human-pathogenic strains such as H2.
To design your primers you will first need to identify highly conserved regions within the matrix gene using ClustalW. Clustal programs are widely used to align multiple sequences.
- Go to ClustalW and copy the sequences (and label information) from this document into the “Input sequences” box.
- Click Run ClustalW and wait for your results.
- Select “clustalw (aln)” to see your alignment.
- Qualitatively, how similar are the sequences? What do the asterisks below the DNA sequences signify?
- Search the alignment to find the target sequences for the Avian Influenza A Matrix Forward and Reverse primers.
- Note that the reverse primer sequence is the reverse complement of the gene sequence. Why?
- How similar are the aligned sequences at these sites?
- Determine the length of the sequence between the primers.
- Is the amplicon in the appropriate size range?
- Using the information concerning primer design that we discussed in lecture, design primers for your AIV screen assay.
- As a reminder, use the following guidelines to design your primers: 1. Tm = 58C +/- 3C, 2. GC% = 50-60% (preferably ending in a G or C), and 3. amplicon length = 100-200bp.
- Your primers should not overlap the current Runstadler primers by more that 5bp.
- Complete the assessments from Part 1 and record the properties of your primers.
- Important note: As you work, remember to keep in mind that the forward and reverse primer bind to different strands of DNA. You may find it easiest to work with only the coding strand sequence at first, and then take the complement of the reverse primer during the very last step of your design.
- When you are happy with your primers, please add them to the table on the "Talk" page, with each primer written from 5' to 3' -- this direction is the standard way that primers are written on an order form. Be sure to include a unique name for your primers (avoid 'forward' or 'primer 1') You must post your primer designs to the M1D2 talk page by M1D3 to facilitate ordering them from IDT, Inc. Note: New due date because of snow storm: M1D4
Homework
Due on M1D3 Note: Q1 and Q2 are due on M1D4 due to snow storm
- For the AIV screening part of this module, you will complete a primer design memo. While the information is fresh in your mind, use your M1D2 notes to generate a table appropriate for submission and write a paragraph explaining your design choices and hypothesis. See the Motivation section of memo assignment for details. Note that you are not yet required to write up the "overall strategy" paragraph.
- Post your final primer design sequences on the M1D2 Talk page so these may be ordered in time. Please add the word "FINAL" next to/below your team color when you are satisfied with the design.
- Include the basepair number at which each primer 'starts' (where the 3' end anneals).
- Read the paper and guiding questions for our upcoming discussion on M1D3. Spend just 10-15 minutes preparing a single slide for your part of the discussion. For now, aim for two things: (1) You should copy the figure you will be presenting into your slide, and (2) You should try to come up with a concise title that states the main conclusion to be drawn from that figure.
- Decide if you will present a Journal Club I article for M1D6 (focus is on microbe/disease detection), or a Journal Club II article for M1D9 (focus is on characterizing microbial communities). Follow the instructions to sign up.
- Review the guidelines about oral presentations for this class (and in general).
- Download the grading rubric that will be used to evaluate this assignment.
Navigation Links
Next Day: 16S PCR and paper discussion
Previous Day: Microbial DNA extraction