Zrusso Biol 368 week 3
From OpenWetWare
Jump to navigationJump to search
Exploring HIV Evolution Electronic Notebook
- Found in the article the GenBank Accession numbers, but it only lists the first and last of the groups of sequences, so it only lists four sequences
- can navigate among the sequences by changing the URL, but as far as I can tell, there is no way to just go between the different sequences from the paper.
- I chose GenBank Accession number AF016821
- It says right in the title that it was taken from subject 2, visit 4, clone 5 from the USA
- Downloaded 5 FASTA sequences to a single word doc for use in the biology workbench
- created a bio workbench userID and attempted to input my sequences but using the browse and upload buttons would kick me back to the homepage, so instead I just manually copied and pasted the sequences in.
- performed a clustalW multiple sequence alignment on the 5 sequences to get an idea of how closely related they are
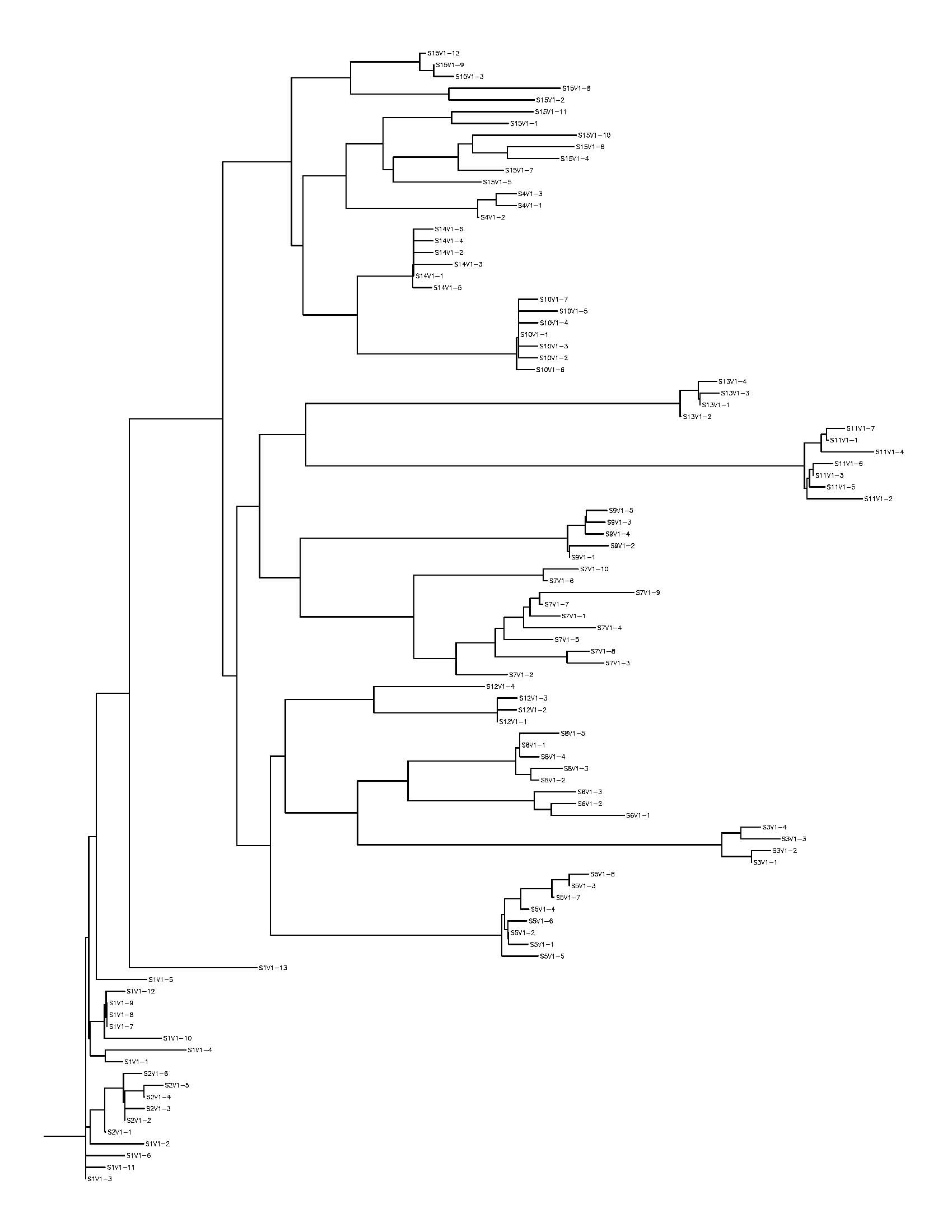
- rooted tree diagram of my 5 sequences
- multiple sequence alignment
- created new session for the subject HIV cross examination
- uploaded all the subject sequences first into .txt files and then into bio workbench
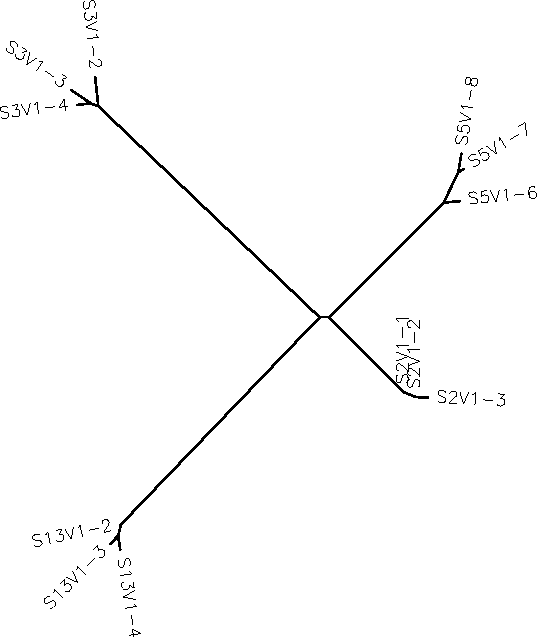
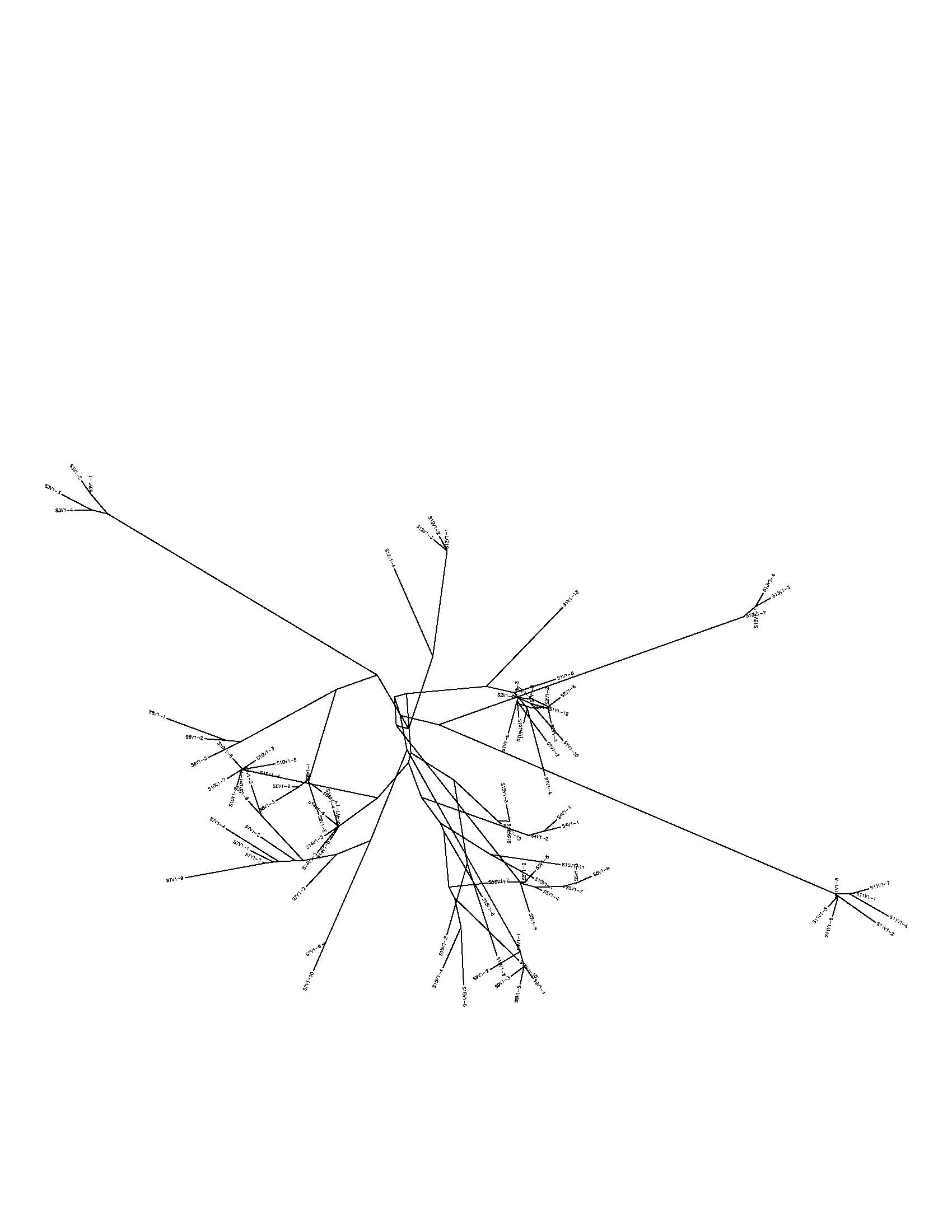
- Did a preliminary analysis of three sequences from 4 different subjects as an unrooted tree
- obviously the clones from each of the subjects were more closely related to each other than to clones from other subjects, but subjects 2 and 5 were more closely related to each other than to subject 3 or 13.
- I picked subject 2 clones 1,2,3; subject 3 clones 2,3,4: subject 5 clones 6,7,8.; and subject 13 clones 2,3,4
- subject 5 seems to show the greatest variation in his/her clonal variants with clone 6 deviating much higher up-branch than the other two who sit at the end.
- I picked subjects 7, 2, and 11 to determine S and Theta as well as the min and max difference.
- figuring out how to determine theta is turning out to be difficult since it is a harmonic sum
- found and linked a website that will use sigma notation
- imported the alignments and used clustaldist to create a distance matrix
- subject 2 had a minimum score of .004 and a max score of .011. After multiplying by the number of bases present, this is 1 min and 3 max
- subject 7 had a min score of .007 and max score of .063. After multiplying by the number of bases present, this is 2 min and 18 max subject 11 had a min score of .003 and a max score of .024. After multiplying by the number of bases present, this is 1 min and 7 max
- compared subjects 2 and 7 and min score was .081 and max score was .116. After multiplying by the number of bases present this is 23 min and 33 max.
- compared subjects 2 and 11 and min score was .121 and max score was .145. After multiplying by number of bases present this is 34 min and 41 max
- compared subjects 7 and 11 and min score was .131 and max score was .167. After multiplying by number of bases present this is 37 min and 48 max.
- excel spreadsheet of data can be found here
Week 4 Journal Club Prep
Word List
- seroconverting - the development of detectable specific antibodies to microorganisms in the blood serum as a result of infection or immunization.
- cohort - a group of subjects who have shared a particular time together during a particular time span
- epidemiology - the branch of medical science concerned with the occurrence, transmission, and control of epidemic diseases
- seropositive - showing positive results on serological examination
- monophyletic - of or relating to a single stock
- progressor - someone who has an increase in the count of whatever is being studied, in this case, HIV
- synonymous - equivalent in meaning
- epitopes - A localized region on the surface of an antigen that is capable of eliciting an immune response and of combining with a specific antibody to counter that response.
- repertoire - a list or supply of capabilities
- recombinant - produced by the combining of genetic material from more than one origin
Outline
Intro
- HIV-1 viruses have a high mutation rate, which allows it to adapt to its environment and avoid antibody response
- It is beneficial to record the genetic diversity and specific mutations of HIV-1 viruses during their evolution in order to understand their ability to evolve to an ever changing ecosystem (us).
- Previous studies involved too few people with too few data points and did not take into account the specific genetic sequences of each virus
- Different selective pressures are associated with the different rates of CD4 cell decline
- The greater number of genetic variants in a system, the more rapid the CD4 cell decline.
Methods
- 15 subjects who were placed in groups (Rapid, Moderate, and Nonprogressors) based on rate of CD4 cell decline.
- The genetic sequence of each virus variant from each subject for each visit was determined through nested PCR and each sequence was uploaded onto GenBank
- Phylogenetic trees both rooted and unrooted for the all the variants of the various subjects were created using the Mega computer package
- The rates of synonymous and nonsynonymous mutations were calculated from the genetic sequences and the data was corrected for bias by adjusting for the number of possible mutation sites for each mutation type.
- Diversity was calculated as the mean of the number of nucleotide differences between clones
- Divergence was calculated as the mean of the percent difference of the nucleotides of the clones from a master sequence taken from one of the first visit clones.
- Rates of diversity and divergence were calculated using regression lines tracing all the diversity and divergence data points for each subject across the entire experiment.
Figures & Results
- Figure 1 shows the progression of T cell count, diversity, and divergence for all 15 subjects as a function of time since their first visit
- Rapid progressors show fastest rate of T cell count decline, along with an increase in both diversity and divergence expect for subject 15 who experienced a net drop in diversity.
- Moderate progressors all show net decline in T cell count except for subject 6 who shows a small increase, diversity and divergence increased for all moderate progressors.
- Nonprogressors all show net increases in T cell count, diversity, and divergence though change in diversity and divergence is low.
- Table 1 shows all the data points collected and calculated throughout the experiment.
- Figure 2 shows the mean rate of change of diversity and divergence for the 3 different types of progressors.
- The rate of change of diversity increases from nonprogressor to moderate progressor to rapid progressor
- there are large error bars for the rapid progressor group that may intersect with the error bars for the moderate progressors. Difference between the two groups was shown to not achieve significance because p value was not low enough.
- Figure 3 shows the phylogenetic tree of all of subject 9’s cataloged clones with the distance between the clones of each visit showing increasingly larger divergence over time from the original clones.
- Figure 4 shows multiple phylogenetic trees including subjects 5, 7, 8, and 14. Each tree shows the genetic evolution of each subject’s virus clone and subsequent clones vary greater and greater over time.
Discussion
- Higher diversity and divergence directly relates to a greater rate of CD4 decline.
- Synonymous mutations were not favored in any progressor group, but nonsynonymous mutations showed greater abundance in rapid and moderate progressors than in nonprgoressors. Indication that there is selection for change in the env sequence of rapid and moderate progressors and selection against change in the nonprogressors which contradicts established model for virus proliferation
- This report agrees with the Mcdonald study that viral strains in rapid progressors show greater genetic divergence, but disagrees with McDonald’s finding that rapid progressors had less net genetic diversity than slow progressors. Most likely due to insufficient data points in Mcdonald study.
- This report agrees with the Nowak study stating that increasing genetic diversity is associated to increasing CD4 cell decline rates.
- Hypothesis: main difference between rapid progressors and nonprogessors is the specificity of immune systems. Rapid progressors have specifically targeted immune responses to specific viral variants whereas nonprogressors attack the variants more broadly thus cutting down genetic diversity.