User:Janet B. Matsen
|
Janet B. Matsen Department of Chemical Engineering |
UPDATE: I'm now a Data Scientist at Zymergen. My current info is on LinkedIn and Twitter.
I am a Chemical Engineering PhD candidate graduating in Winter 2016. My major project has been implementation of a novel carbon-fixation pathway, which included a computationally designed enzyme and three enzyme reactions not found in nature. For my last year I have transitioned to fully computational work. My new project involves metagenomics and metatranscriptomics of a methane oxidizing community using collaborative programming, remote and cloud computing, machine learning, and visualization of large multivariate data sets. In addition, I will graduate with an Advanced Data Science certificate for coursework in statistics, machine learning, data management, and data visualization.
Please see my resume or LinkedIn for more professional information and GitHub for some of my code.
I work with Mary Lidstrom, David Baker, and David Beck.
OpenWetWare Contributions
I started the Lidstrom Lab OWW wiki in 2011 and love posting what I learn! Wet lab biology is full of dogmas that I enjoy challenging. When I learn that dogmas are false, or the importance of particular variables in methods, I post in The Lidstrom lab's wiki. I have over 27,000 contributions over dozens of pages. Two popular ones are my Guide to Gibson Assembly (58,000 views 4/2016), and SDS-PAGE (46,000 views 4/2016).
In addition, I posted some protocols specific to my PhD work to a GitHub driven web page for use in working with Helen Chan, a fabulous Chemical Engineering undergraduate .
Research Interests
- production of chemicals using microbes
- protein engineering
- metabolic engineering
- synthetic biology
- transcriptomics
- chemical engineering applied to biology
Education
PhD (in progress) University of Washington, Seattle
- Chemical Engineering, expected graduation: 2015-16
- Lidstrom Lab
B.S. University of California, Berkeley
- Chemical Engineering, 2010
Publications
- Matsen, Yang, Stein, Beck, & Kalyuzhnaya. Global molecular analyses of methane metabolism in methanotrophic alphaproteobacterium, Methylosinus trichosporium OB3b. Part I: transcriptomic study. Frontiers in Microbiology (open access), 2013
- Yang, Matsen, Konopka, Green-Saxena, Clubb, Sadilek, Orphan, Beck, & Kalyuzhnaya. Global molecular analyses of methane metabolism in methanotrophic Alphaproteobacterium, Methylosinus trichosporium OB3b. Part II. metabolomics and 13C-labeling study. Frontiers in Microbiology (open access), 2013
Awards & Activities
- 2012 honorable mention for the National Science Foundation's Graduate Research Fellowship Program
Outreach:
- 2011-present Outreach Coordinator for the Puget Sound chapter of the American Institute of Chemical Engineers
- Leading a mentoring project with 8 chemical engineering mentors and 8 students from the Technology Access Foundation Academy in Kent, WA.
- 2010-2011 Outreach Coordinator for the University of Washington chapter of the American Chemical Engineering Society
- Organized two half-day and one all-day events for students from MESA, the Math, Engineering, Science Achievement organization of Washington, involving 60 volunteer- hours and resulting in 660 student-hours of outreach to disadvantaged minority students.
- Misc. outreach:
- Gave a presentation to high school students describing statistical challenges associated with transcriptomics research.
- Hosted a booth at Engineering Discovery Days at University of Washington, engaging and educating the public about chemical engineering.
My Personal Pages
- GitHub
- Protocols specific to my project, as of 3/2015. Hosted on GitHub.
- Guide to Gibson Assembly
- Lab Tips & Tricks
- Useful Links
- Books I like
- Personal Notes for Thesis Project
Not maintained any longer:
Tools to Share
GitHub Repository
- Janet on GitHub
- All of my plasmid files are found in a version controlled repository
- Protocols for my personal use and collaboration with Helen Chan (Chemical Engineering Undergrad) are here: GitHub Pages: Janet Matsen.
- I'm beginning to contribute code to the LidLab GitHub repository. My sub-folder is here.
- Favorite function for exporting data from the SpectraMax 190 plate reader: SpectraMax_190_plate_reader_data_importer
APE annotation library generator & list of primers to share with our lab
- This is the first script I ever wrote, and remains important in my research. Feel free to download it and enjoy it yourself.
- Ape Annotation Feature Library Creator
- This is an R script that converts the info in my list of primers into a file that I can use to annotate DNA files in APE with.
- It:
- trims out sequences not intended for sequencing such as Gibson assembly primers
- makes a label that combines the unique primer number, the melting temperature, and the letter F or R for forward or reverse, and an asterisk if you should consult the primer spreadsheet comments before using it
- assigns colors in APE that communicate whether it primers in the forward direction or the reverse direction.
- saves the info in the format APE needs, with the date it was generated in the title.
- This allows me to instantly see where all of the primers I own bind to a DNA sequence for a given project I am working on. It also allows me to share these primers very easily; by sharing the file it outputs allows my lab mates to instantly see if I have any primers that can be used in their project. It has been very handy for them!
- I am happy to help friends modify this script to be useful with their own primer libraries! No R experience is necessary.
- Anyone can access my most current primer "Annotation Feature Library" here. You can also see the files used to generate it there.
Use notes
- If the primer binds in the forward direction, the primer will be light gray
- If the primer binds in the reverse direction, it will be dark gray
- If the primer binds in the opposite direction stated in my primer table, it will appear red. (If it says F in the primer name, it is a reverse primer & vice versa.)
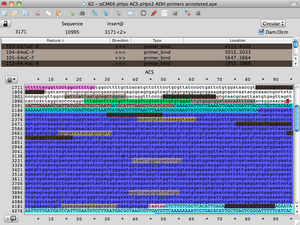
- Examples:
- Primer 7 is VF2 in BioBricks. Primer 60 is its reverse compliment. In a biobrick vector, it appears light gray for 7 and dark gray for 60. pCM66 happens to have this same sequence in the region upstream from the multiple cloning site, except it is REVERSED. Both primers will appear red as they bind in the opposite direction expected.
- I designed some primers for a Kan cassette. The Kan cassette in pCM66 is read in the reverse direction, so all the primers built for a forward Kan cassette appear red.
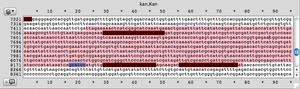

Kan primers binding in the opposite direction relative to my database appear red
Skills
- Metabolic engineering, molecular biology, enzyme assays, enzyme evolution, high-throughput screening, metabolomics, Gibson cloning
- R & ggplot2, Python, Git/GitHub, LaTeX, Inkscape, Linux
Why I love ggplot2 (and R)
Data is beautiful. Interacting and communicating data elegantly makes me happy.
R is a relatively easy language to pick up, whether or not you have prior programming experience. It is one of the best languages for noodling with tabular data and doing statistics, though Python's emulations of R's strengths are growing more appealing. I use R because I am in love with the ggplot2 plotting package in R. To get a sense of its power, just type "ggplot2" into google images. The book that introduces the fundamentals is freely available online.
Two cool features of ggplot2:
(1) Layers. Imagine you have defined a plot called p in a program. If you want to add some layer to the plot, you just say p + layer. You can just layer in data, aesthetics, statistics, etc. You can also make one base plot, then make a bunch of variants of it by adding different layers of interest. It is hard to imagine going back once you have this freedom. Layers have different types of geometries you can apply.
(2) Facets. Biological data and experimental data are complex! ggplot2 can help you plot complex data by spatially separating out variables, mapping multiple aesthetics (color, size, shape, outline, etc.) to one point, communicating more information than an excel plot can.
Having scripts that I can recycle when doing similar experiments allows me to do in-depth quality checks and make summary statistics that would be impractical to do with Excel. The quality check plots are automatically generated for each data set. Then I add plots that are specific to the questions I was investigating with my experiment. This series of plots paint a story about how the experiment was performed, what variables were important, and what the key findings are. I can compare elements of different experiments by comparing similar plots that are generated (almost) automatically for each experiment with the (almost) identical scripts. Here is a sample folder sample folder] from an experiment I did recently to get a small sense.

