Stanford/BIOE44:Module 3:Day1
M2: Day 1 - What Pollutant to Detect?
Introduction
Your primary goal today is to identify an environmental pollutant that can be via a genetically encoded sensor. Your goal is to be able to specific the standardized DNA encoding your new sensor by 3p this Thursday, so that we can place the DNA synthesis orders at that time. Overall, you should design a device that receives as input an environmental pollutant and that produces a PoPS signal as output (so that your sensor device can be connected to one or more PoPS-based color generators). Please consider today's class notes for starting hints and materials regarding environmental pollutants that might be worth detecting.
Reprise From Last Thursday
What happened with our gel extractions?
What did Isis do?
I had a hunch that the isopropanol step was messing up our gel extractions so I decided to test this. I also prepared some color generators and vectors.
- I digested 10ul K274003, K274100, K274200 with XbaI/PstI and K274220 (pSB2k3 vector) with EcoRI/PstI. This is roughly 2ug of DNA for each.
- For K274003 (this is what I used to test the gel extractions) I separated the digestion into 3 wells on a gel. The other digests each got their own well. (0.8% agarose)
- I cut out each band and gel extracted with and without isopropanol for the K274003 pieces. The rest were extracted with a Zymo kit.
- After extraction I ran 0.5ul on a gel to determine concentration (this is the gel in the image below).
The Results
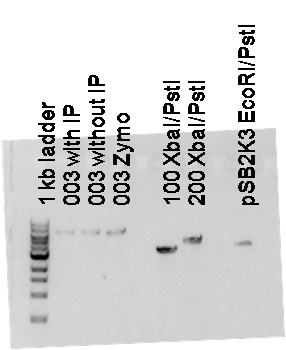
Legend
| Label | Meaning | Concentration |
| 1kb ladder | NEB 1kb ladder | |
| 003 with IP | Part K274003 (green generator) cut at XbaI/PstI and gel extracted using Qiagen kit with isopropanol step included. | 50 ng/ul |
| 003 without IP | Part K274003 (green generator) cut at XbaI/PstI and gel extracted using Qiagen kit with no isopropanol step. | 50 ng/ul |
| 003 Zymo | Part K274003 (green generator) cut at XbaI/PstI and gel extracted using Zymo kit. | ~75 ng/ul |
| 100 XbaI/PstI | Part K274100 (red generator) cut at XbaI/PstI, gel extracted with Zymo kit | 250 ng/ul |
| 200 XbaI/PstI | Part K274200 (orange generator) cut at XbaI/PstI, gel extracted with Zymo kit | 250 ng/ul |
| pSB2K3 EcoRI/PstI | Vector pSB2K3 cut at EcoRI/PstI, gel extracted with Zymo kit | 150 ng/ul |
Getting Started
Refine a part
To quickly give you a sense of what you'll ultimately need to be able to do (in order to complete your new sequence designs by Thursday), and also to get you working with some cool software tools, please start today's lab by turning a natural sequence of DNA into a standard biological part. The sequence of DNA that you'll be refining encodes a gas filled protein vesicle. Soil bacteria apparently control their buoyancy in order to maintain or change their position in a soil column (e.g., in response to rain). Follow these steps to show that you can find and refine a natural sequence of DNA encoding a protein balloon!
- Search the web for "protein vesicles soil bacteria"
- Scan your search results and look for any research paper that might have identified the relevant genes encoding such a biochemical function
- Search for the same article on the NIH PubMed literature database (see the "Useful Tools" section below).
- From your PubMed search results, see if there are any nucleotide sequences associated with the article that are stored in the Genbank database
- Download or copy the full Genbank entry for the natural sequence of DNA encoding the genetic / biochemical functions you want.
- Decide what physical assembly standard you want to use (this will determine what restriction sites you need to remove, if any).
- Check for any incompatible restriction sites within the natural sequence. Remove the sites via silent mutations (note any changes that you make).
- Check to make sure that your DNA meets any needed or relevant functional composition standards (e.g., PoPS input)?
- Bring your new sequence to Kosh or Isis for verification.
Choose a new pollutant to detect
We've heard from our colleagues in the Stanford Environmental Engineering Department that they would value having biodetectors for the following compounds (and other things):
- Nitrate (in seawater)
- E. coli (in seawater
- Radium (in seawater)
- Norovirus (in seawater)
- Emerging organic contaminants (e.g., Caffeine, Lipitor, Ethinylestradiol, Ibprophen)
- X-ray contrast media (e.g., Diatrizoate and Iopromide)
- Selenite
Quickly consider any or all from the above list, or add something to the list that you can justify / make an argument for investing your energy and course resources to pursue. Ask us questions. See if you can make any progress on finding a gene that might encode a detector for your target compound.
GOAL: BY THE END OF CLASS TODAY YOU MUST HAVE CHOSEN ONE TARGET AND HAVE AT LEAST ONE LEAD PAPER TO READ COVERING THE BIOLOGY OF THE GENETIC MATERIAL.
FOR NEXT TIME
Before leaving lab today, get approval from Drew regarding the research article you will skim that will help inform your sequence refinement work during Thursday's lab. Your "for next time" work for this Thursday will be to skim / read your specific article.
Useful tools
Literature
Sequence Viewing and Manipulation
- ApE (free download)
- NEBcutter
- BIOFAB sequence refiner
- GeneDesigner 3.0
