Stanford/BIOE44:Module 1:Day1
M1: Day 1 - Plasmid Extraction and Restriction Digest
Introduction
Today we are going to extract plasmids containing the color generator gene cassettes from E. coli cultures. Then we will cut the gene cassette out of the plasmid with a restriction digest. Next week we will be putting the gene cassette into a different plasmid.
Standard Assembly Overview
Background on Methods
Plasmid Extractions
Plasmids are extra-chromosomal self replicating pieces of DNA that are, in many cases, circular. In genetic engineering they are used to move information (genes, etc) from one host to another. When used in this context they are called vectors.
Restriction Digests
Restriction endonucleases, also called restriction enzymes, cut (“digest”) DNA at specific sequences of bases. The restriction enzymes are named for the prokaryotic organism from which they were isolated. For example, the restriction endonuclease EcoRI (pronounced “echo-are-one”) was originally isolated from E. coli giving it the “Eco” part of the name. “RI” indicates the particular version on the E. coli strain (RY13) and the fact that it was the first restriction enzyme isolated from this strain.
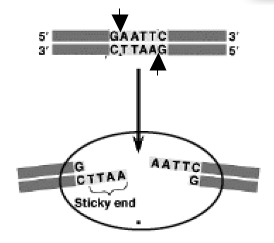
The sequence of DNA that is bound and cleaved by an endonuclease is called the recognition sequence or restriction site. These sequences are usually four or six base pairs long and palindromic, that is, they read the same 5’ to 3’ on the top and bottom strand of DNA. For example the recognition sequence for EcoRI is
5’ GAATTC 3’
3’ CTTAAG 5’
Other restriction enzymes, for example HaeIII, cut in the middle of the palindrome leaving no DNA overhang, called a “blunt end.” One of the most useful resources for restriction enzyme information is the website from New England Biolabs NEB homepage. Use their search engine to retrieve information about the recognition enzymes XbaI and PstI. Be sure you are clear on how they differ before you move on to the experiment. For example, do the enzymes have the same recognition sites? do they leave the same overhang? will they work in the same buffer? at the same temperature? These are some of the preliminary questions you'll have to ask yourself whenever you set up a restriction digest.
There are normally 4-5 components in a given reaction:
- DNA which will be digested: Must have specific site recognition site for enzyme. Also needs to be free of contaminants such as phenol, alcohol, excessive salts, which interfere with enzyme activity.
- Restriction enzymes: Restriction enzymes are fragile! They need to be treated carefully. ALWAYS on ice. Use a new pipette tip each time you go into tube; hold by top of tube. Enzyme is stored in 50% glycerol, so that it does not freeze at -20 degrees Celsius and the enzyme retains its function. However, more than 5% glycerol in a digest reaction will disrupt the proper reaction- either there will not be digestion, or the enzyme will exhibit star activity (which is term for cleavage at sites other than the defined recognition sequence of the enzyme). Some enzymes are more likely to exhibit star activity than others, for instance Eco RI; thus it is important to not add too much enzyme- it is better to avoid over-digestion and keep glycerol concentration lower. Also low ionic strength and high pH also contribute to star activity, so important to use proper buffer (see below). One important point about the enzyme- you should add it last. All reaction components should be mixed gently but thoroughly, and then the enzyme added and mixed again. Enzyme should never exceed 10% of total reaction volume.
- Buffer: The restriction enzyme requires a particular ionic strength and pH. Buffers contain various salts and other components which the enzyme needs in order to function optimally. Tris – buffer; MgCl2 – Mg is cofactor for RE. NaCl to maintain ionic strength, mimics cellular conditions. 0.025% Triton X-100 (detergent)- keeps proteins and DNA from sticking to sides of tube.
- BSA: Some enzymes require bovine serum albumin (BSA) for optimal activity, because it’s good to have other proteins around to take up space. None of the enzymes we are using require it though (adding it never harms the reaction).
- Water: to achieve proper ratios of each component.
The recommended incubation temperature for most RE is 37 degrees Celsius.
Extra Reading/References
- Standard Assembly Info
- Cambridge iGEM project 2009 - E. Chromi
- How are plasmids separated from chromosomal DNA?
- "A rapid alkaline extraction procedure" <-- (1979 paper from Nucleic Acids Research)
In Class
Part 1: Plasmid extraction (aka miniprep)
At the beginning of class you will be given a 5mL culture of E. coli harboring a plasmid that contains a BioBrick part. We will extract the plasmid from the bacteria using a kit (QIAPrep Spin Miniprep kit). Then we'll measure the concentration of the plasmid DNA extracted using a nanodrop spectrophotometer.
Mini-Prep Protocol
Materials
- 5mL E. coli culture
- microcentrifuge tubes
- Buffers P1, P2, N3, and PE
- Qiaprep column
- sterile water
Collect the Cells
- Label three microcentrifuge tubes.
- Pour approx. 1.5 mL of culture into each tube.
- Centrifuge at 10000xg for 5 minutes.
- Aspirate off supernatant. Be careful not to suck up your cell pellet.
Break the cells open
- Resuspend the cells in tube #1 with 250uL of Buffer P1.
- Transfer all liquid from tube #1 to tube #2 and resuspend the pellet.
- Repeat from tube #2 to tube #3. (Should end up with all the cells in tube #3. Throw away tubes 1 and 2)
- Add 250uL of Buffer P2 to cells. Invert the tube 4-6 times. Solution should become slightly clear and viscous. (This is the step where the cells are lysed and all the the proteins are denatured)
- Add 350uL of Buffer N3 to the solution. Mix immediately by inverting the tube 4-6 times. (Thick white goo will form - this is all the proteins precipitating out)
- Centrifuge for 10 minutes at max speed (approx 17000xg). While centrifuging, take out a column and label it.
Isolate the plasmids
- Apply the supernatant from the tube to the column by pipetting (approx 800ul). Be careful not to pipette any of the white stuff - this will clog your column and reduce your yield.
- Centrifuge for 1 minute at max speed. Discard the flow through.
- Apply 750uL of Buffer PE and centrifuge for 1 minute at max speed.
- Discard the flow through. Centrifuge the column for an additional 1 minute to remove residual wash buffer.
- Place the column in a clean 1.5ml micro-centrifuge tube. Apply 50uL sterile water to the column. Let it stand for 1 minute. Centrifuge at max speed for 1 minute.
Measure the concentration
- Measure the concentration of your newly prepped plasmid with the nano-drop. Record the concentration and 260/280 in your lab notebook.
Part 2: Restriction Enzyme Digest
We will take the plasmid we just extracted and set up a restriction digest to cut out the color generator genes.
- What restriction enzymes should we use?
For Next Time
Please read "Idempotent Vector Design for Standard Assembly of Biobricks" by Tom Knight.
The formal reference is here.
A direct link to the PDF is here.
