ISEEM Progress September 2009
| Home | Project | People | News | For Team | Calendar | Library |
I. Progress made toward completion of our goals
A. Personnel
Thomas Sharpton started as a post-doc in the Pollard lab.
B. Research
Eisen lab
MicrobeDB is a resource that was constructed by Morgan Langille while in his previous research lab. Morgan has updated the resource and installed it on the new "edhar" server for the iSEEM project.
MicrobeDB is a local storage repository of all RefSeq microbial genomes (Bacteria and Archaea) from NCBI. All genomes are downloaded monthly from NCBI, stored locally on the Edhar server. The files, including information about the sequenced strain and the genes within it, are parsed into a MySQL database for easier access and use. Users of the iSEEM project can either access the database directly using MySQL queries, or use the Perl API that has also been written to query the MySQL database more easily.
The downloading and parsing of the NCBI files is done automatically once a month, thereby having the most recently sequenced genomes available for use. Each monthly download is given a "version id" that represents a snapshot of the data at a given date. Each version can be saved by a user and easily referred to over time without the worry of the data changing.
MicrobeDB is split into three major groupings of information (Genome Projects, Replicons, and Genes):
- Genomeproject
- Contains information about the genome project and the organism that was sequenced
- E.g. taxon_id, org_name, lineage, gram_stain, genome_gc, patho_status, disease, genome_size, pathogenic_in, temp_range, habitat, shape, arrangement, endospore, motility, salinity, etc.
- Each genomeproject contains one or more replicons
- Replicon
- Chromosome or plasmids
- E.g. rep_accnum, definition, rep_type, rep_ginum, cds_num, gene_num, protein_num, genome_id, rep_size, rna_num, rep_seq (complete nucleotide sequence)
- Each replicon contains one or more genes
- Gene
- Contains gene annotations and also the DNA and protein sequences (if protein coding gene)
- E.g. gid, pid, protein_accnum, gene_type, gene_start, gene_end, gene_length, gene_strand, gene_name, locus_tag, gene_product, gene_seq, protein_seq
MicrobeDB allows centralized access to the previously finished genomes that are often one of the first resources needed in several of the iSEEM projects. In particular, the identification of novel families and sub-families in metagenomics data will first require the construction of gene families for finished genomes. In addition, it is planned that the gene families and other data produced by novel iSEEM projects may be added to MicrobeDB to allow centralized access and easier collaboration of the data.
BioTorrents
BioTorrents (www.biotorrents.net) was created by Morgan Langille to allow faster and easier exchange of biological data and software. BioTorrents acts as a central location for researchers to share their open-access data and software and uses the Bittorrent peer-to-peer technology.
BioTorrents can aid to the sharing of biological data in two separate ways:
1) Currently, researchers that want to make their own data or software accessibly by others will either host it on their own server or submit it to a large database (e.g. submitting genome data to NCBI). In both cases, the speed at which data can be downloaded from these servers depends solely on the bandwidth available to the host. Bittorrent technology increases these transfer speeds by allow the people who download the data to also share with each other thereby increasing the download speed for everyone. In addition, researchers that download the data can continue to help others download the data faster by continuing to share the resource on their own computer. This also allows the data to become decentralized so that if the main server is temporarily or permanently off-line the data is still available from the other researchers sharing the data. For example, the metagenomic datasets hosted at CAMERA are often very large and if they were to be added to BioTorrents they could be downloaded much faster if several users share the data at the same time.
2) Traditionally, most biological data that is made accessible to other researchers is only shared once the project is finished and often published. These data sets are then formally submitted to a central resource such as NCBI, or are hosted on a project or research lab website. However, many projects produce data sets in the intermediate steps of their analyzes that are never published or shared with other researchers. BioTorrents can provide easy and informal access to this data for those wanting to share their results. Researchers have started to see the benefits of sharing experiment thoughts, ideas, and results before publication via Blogs and other public web forums. BioTorrents adds on to this open-access movement by providing a central and easy method to share biological data.
Community profiling
Community profiling is a non-similarity based approach for the annotation of genes with unknown function and is being researched by Morgan Langille. It leverages the massive amount of sequence data from multiple samples in metagenomics data to identify families of genes that have similar profiles (absence/presence or relative numbers of genes) across various environmental samples. The ideas is that these families with similar profiles may have similar functions. It is currently not clear how well this methodology will work in practice, but if successful it could help identify possible functions for completely novel gene families.
Previously, a first attempt at community profiling was done by identifying PFAMs (11,000 HMMs) in metagenomic sequences from GOS (41 million peptide sequences across 80 samples). A more recent analysis was performed using "all metagenomics peptides" from CAMERA (node1055606926236189023.fasta), that included several other samples in addition to the GOS samples (43,240,119 sequences). PFam predictions by HMMER 3 (beta) were grouped into their corresponding samples and enumerated. This data was then clustered using microarry clustering software, Cluster 3.0.
There are several ways to cluster and normalize the results as well as methods to try and minimize any bias between the environmental samples. Before fine tuning these variables, a measurement is needed to test how well the clustering is working. This measurement needs to be able to test how similar the functions are of the genes within each cluster. The following steps were conducted to perform this analysis:
- The hiearachal clusters were broken into 575 groups using a correlation cutoff of 0.90 or above.
- PFams were mapped to Gene Ontology terms using the pfam2GO mapping
- 1893 PFams had no associated GO term (695 of these were Domains of Unknown Funcion:DUFs)
- 3377 PFams did have one or more associated GO terms and could be used for further analysis
- Only 67 (of 575) clusters contained 4 or more PFams with at least one GO term
- G-SESAME was then used to measure the functional similarity between all GO terms in each cluster and each cluster was given a functional similarity average.
- The average of all cluster averages was 0.484
- 10 clusters had a score of 0.60 or greater.
- The data was then randomized by using the same GO terms but in different random clusters and a score of 0.412-0.420 over 4 iterations
- Each of the 4 iterations had only 1 or 0 clusters with a score of 0.60 or greater
In summary, the community profiling approach does seem to work, but at the current state not much better than random. However, the development of a metric for cluster functional similarity now allows for further improvement of the clustering and bias normalization.
Protein families
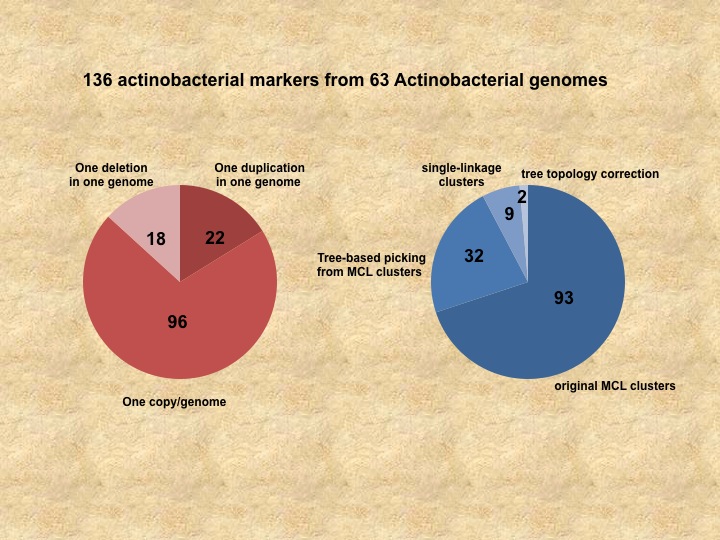
We've an automatic tree-based protocol to identify phylogenetic candidates for a given group of genomes created by Dongying Wu. Current phylogenetic markers are identified from MCL clusters. Such approach have two major issues: 1. MCL clustering can pool different marker families into one because of sequence similarities. 2.MCL clustering can break marker families apart because only limited numbers of inflation values can be used practically. To address the first issue, we build phylogenetic trees for all the mcl clusters, and use a newly developed program to parsing the trees to identify clades that include potential marker candidates. Universality and even distribution are evaluated for all the potential makers, HMM3 profiles are build and searched against the entire proteomes from the genomes to decide if the families are distinct from other peptides. To address the second issue, we perform a single linkage clustering after filter out the identified phylogenetic markers and families with large number of members in the MCL clustering step. The single linkage clusters are used for identify more makers using the same tree-base approach. We use 63 actinobacteria genomes as an example to test our new methods.We've identified 40% more markers as an results (See Figure), and the new approach is automated with no manual curation and readily to be applied for other phyla.
Green Lab
Metagenomic analysis of community phylogenetic structure
Phylogenetic inference for metagenomic data
Since community phylogenetic stucture can only be studied once a phylogeny has been inferred, we have focused on developing and evaluating different methods to infer phylogenetic relationships among metagenomic reads from different gene families.
As noted previously, using the HOT/ALOHA dataset hosted by CAMERA as an example data set, we identified metagenomic reads from 31 gene families using AMPHORA. To provide a sufficient number of reads to estimate phylogenetic diversity within each environmental sample in this data set, the metagenomic reads from all gene families (query sequences) were then combined with the 571 full-length sequences from AMPHORA (reference sequences) into a single concatenated/tiled alignment. Scripts to perform this concatenation/tiling are provided here.
We have used two methods to infer phylogenetic relationships based on metagenomic sequence reads from different gene families. Both take advantage of phylogenetic signal in the AMPHORA reference sequence database to improve placement of short, fragmentary metagenomic reads.
The first approach has been to perform a phylogenetic analysis on the entire combined alignment using maximum likelihood methods. This approach works for data sets of moderate size such as the HOT/ALOHA data set but would not be computationally feasible for larger metagenomic data sets such as GOS due to the time consuming nature of the ML tree search and optimization. For large data sets such as GOS we are currently applying a similar method using the hybrid neighbor joining/likelihood tree inference methods implemented in FastTree 2.0 to infer phylogenetic relationships among the 154,102 sequences identified by AMPHORA in the GOS data set. This approach appears to be working well; most metagenomic reads are placed with fairly high bootstrap support.
The second approach uses a recently developed algorithm for placement of individual reads onto a reference phylogeny using likelihood optimization of query sequence placement and branch lengths. This method is implemented in RAxML 7.2 and speeds up the placement of reads relative to the previously described methods since individual reads can be placed without optimizing their relationships with all other reads. The method also gives an indication of how much unceratinty there is in the placement of each query sequence by performing bootstrap replicates of the analysis to estimate uncertainty in sequence placement. We implemented scripts that generate a posterior distribution of likely phylogenetic trees from the bootstrap replicate placements of metagenomic sequences on the phylogeny, and we are in consultation with the author of RAxML to modify the output of the software to provide this information automatically.
We are currently using the metagenomic simulator developed by the Pollard lab to quantify the relative performance of these methods.
Community phylogenetic structure
Using the phylogenetic trees linking all metagenomic sequences in the HOT/ALOHA data set described above, we analyzed bacterial community phylogenetic structure and turnover along the oceanic depth gradient sampled by this study.
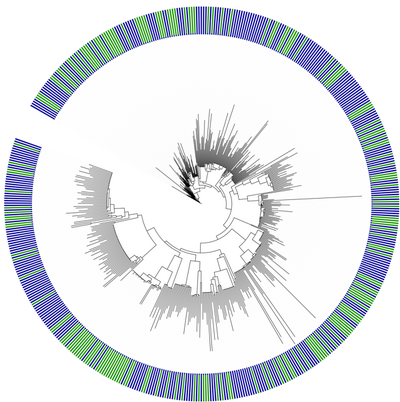
We compared taxonomic diversity and phylogenetic diversity along the ocean depth gradient. Taxonomic diversity, measured as the number of OTUs based on 16S sequencing in the samples at a 95% or 99% cutoff, was uncorrelated with depth. Phylogenetic diversity, measured as the total branch length in each sample, was strongly correlated with depth. Because phylogenetic diversity is correlated with the number of sequences in a sample, we standardized phylogenetic diversity by comparing it with the diversity expected from a random draw of the same number of sequences from the pool of all sequences observed at all depths. Standardized phylogenetic diversity showed a quadratic relatonship with depth, with samples in the epipelagic zone (<200m depth) phylogenetically clustered, while samples at intermediate depths were phylogenetically even. This result was robust to different methods of tree inference and was observed across bootstrap replicate estimates of the phylogeny.
Clustering of communities based on phylogenetic distances among samples was measured using hierarchical complete linkage clustering of UniFrac distance calculated using the maximum likelihood phylogeny. Phylogenetic clustering of communities grouped samples from similar depths together, but illustrated a pattern where bathypelagic samples (770m and 4000m depth) were highly phylogenetically similar to one another, and surprisingly were actually more phylogenetically similar to the epipelagic photic zone samples (10m and 70m depth) than to the lower epi- and mesopelagic samples (130m and 200m depth).
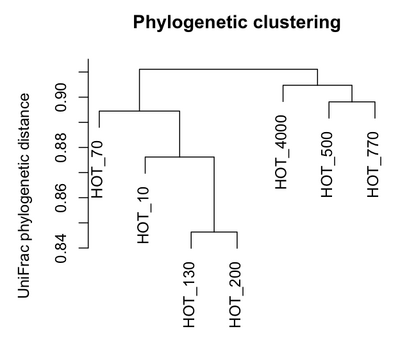
We are writing up these results and hope to submit the manuscript shortly. We have begun running similar analyses on the GOS data set, but the much larger size of this data set compared to the HOT/ALOHA data has required modification of existing methods to calculate phylogenetic diversity. We are exploring options including developing new software tools or using existing programs such as FastUnifrac and Mothur to deal with analysis of phylogenetic diversity data sets with hundreds of thousands of sequences.
Novel Biodiversity Measures
Field theory for biogeography: a spatially-explicit model for predicting patterns of biodiversity
We have completed the project outlined in our June Progress Report, building on the foundation of neutral community ecology to derive the first spatially-explicit neutral prediction for the Species- or Taxa-Area Relationship (SAR) (O'Dwyer and Green, Ecology Letters, in press, pdf linked here: [1]). The SAR is one of the most fundamental patterns in spatial ecology, and our work provides both a null prediction for its shape, and a quantitative connection between the SAR and beta-diversity. Our approach is to model a community from the bottom-up, specifying simple mechanistic rules for the dispersal of individuals across their environment, and combining these rules with the stochastic processes of birth, death and speciation. Despite the conceptual simplicity, the combination of explicit spatial processes with demographic stochasticity presents a formidable challenge. To take on this challenge, we used the methods of statistical and quantum field theory to set up and solve a set of functional differential equations, using which we make a number of ecological predictions.
A crucial feature of our framework is that we combine and link our prediction for the SAR with a relatively novel measure of taxonomic beta-diversity. This measure is the probability that two individuals picked at random, and separated by a distance r, will belong to the same taxon, and is denoted by F(r), a function of r. This function has been unexplored in microbial communities, but we believe the connection between these two distinct measures of spatial diversity will be crucial in making predictions for microbial diversity. The undersampling of microbial life makes SARs extremely difficult to measure directly, but in contrast to the SAR, F(r) does not strongly depend on rare taxa. The structure of our model provides the link between these two quantities, and we believe that the connection will be a powerful tool in uncovering microbial diversity at the taxonomic level.
In terms of the details of our prediction for the SAR, we find three distinct phases. At intermediate scales our SAR is closely approximated by a power law, so that on log-log axes a plot of species number as a function of area is linear, with a slope determined by the underlying evolutionary processes. This tri-phasic pattern has been identified across decades of empirical studies, but it has often been thought that the pattern must arise from the effects of spatial heterogeneity: as one samples increasingly large regions, more environmental niches are uncovered, allowing increasing numbers of species to occupy these niches. Our work demonstrates that neutral processes and dispersal limitation alone give rise to an extremely realistic prediction for the SAR, without invoking spatial heterogeneity and environmental selection, and shows that a power law SAR at intermediate scales arises naturally from the combination of speciation with local dispersal. Of course, this does not rule out environmental heterogeneity as an important, or even the primary driver of the SAR for marine microbes, and the next phase of our project will test this hypothesis.
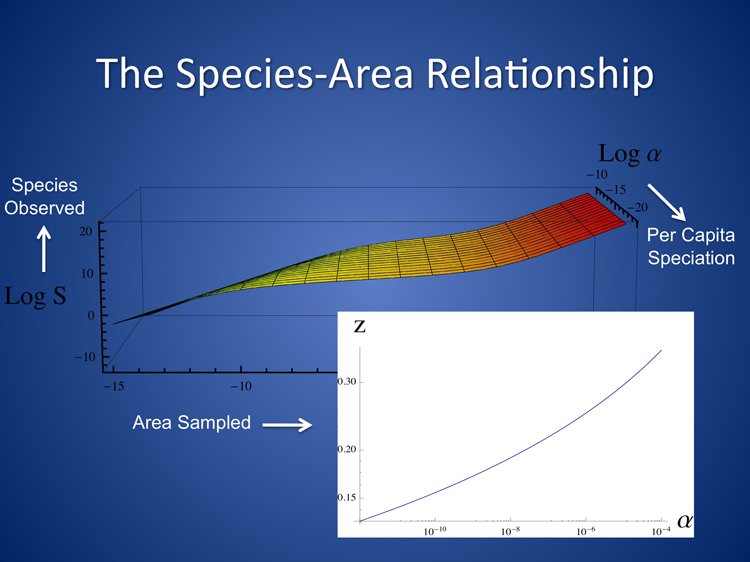
Taxa from Metagenomic Data: who is where? (in collaboration w/Pollard lab)
The next phase of our project involves significant synergy with the Pollard lab, and so we have been working closely together to determine the abundance and location of different Operational Taxonomic units (OTUs) in the Global Ocean Survey (GOS) data-set. The aim of the Green lab is to compare this data with the predictions of our spatially-explicit neutral model, described above and currently in press. This differs from the approach in the Pollard lab (description linked here), so that both labs are tackling questions related to taxonomic diversity, from different directions.
The overall objectives of the Green lab in this phase of the project are two-fold. First, we will test how closely our neutral predictions reflect the patterns in the GOS data. In particular, we will be looking at a measure of beta-diversity, F(r), which is the probability that two individuals separated by a distance r belong to the same OTU. The second stage will depend on the results of the first. If we find that our neutral model is capturing the structure of F(r) across different spatial scales, we will use the theoretical link we have derived between F(r) and the SAR to make a prediction for the marine microbial Taxa-area relationship. On the other hand, if we find that the neutral F(r) doesn't reflect the spatial structure we find in the GOS, we will use this data as a guideline to adding complexity to our model. An outstanding question in ecology is the balance between neutral, stochastic processes, and deterministic, niche-based processes, and we believe the combination of our flexible theoretical framework with microbial data will begin to unravel this question.
So far, in collaboration with the Pollard lab, we have isolated and aligned the 16S reads from the GOS, discussed in more detail below, and begun the process of constructing an accurate phylogeny from this alignment. The matrix of branch lengths between every pair of reads on this tree will provide the input for the bioinformatics tool, MOTHUR, which we will use to determine the abundance and location of OTUs across the different sample sites in the GOS.
Pollard Lab
Distance-Decay and Taxa-Area Relationships
A basic question in microbial ecology is the extent to which habitat heterogeneity and dispersal limitation affect the large-scale distribution of taxa. At one extreme, it is possible that most taxa occur everywhere, while on the other extreme, it is possible that the distributions of taxa are highly constrained by environmental conditions and inability to move between locations. Addressing this question has implications for understanding microbial evolution -– whether microbes are primarily adapted to specific niches or cosmopolitan –- and estimating regional to global microbial diversity.
Two characteristics of the distributions of taxa particularly bear on this question, taxa-area relationships and distance-decay relationships. Taxa-area relationships describe how the number of taxa occurring in a region increases with the area of that region. Thus, they can also allow regional or global taxa richness to be estimated. As for distance-decay relationships, naturally, distant communities tend to share fewer taxa than proximate ones. Distance-decay relationships describe how the similarity between between pairs of communities (or samples) decays as a function of the distance between them.
Unfortunately, it is challenging to construct microbial taxa-area relationships because censusing microbes in large regions is difficult. For instance, it is impractical to census all microbes inhabiting large volumes of sea water. However, distance-decay relationships can easily be constructed for microbes, and in addition to allowing direct ecological inferences, they can allow taxa-area relationships to be derived (see the June 2009 progress report). To construct a distance-decay relationship, the taxa in numerous small samples must first be censused. For various subsets of the samples, a measure of similarity is then calculated -- for instance, the number of taxa that the samples share divided by the total number of taxa in the samples -- and this is regressed on the distance between the samples to give the distance-decay relationship. A rapidly decreasing distance-decay relationship suggests strong effects of habitat heterogeneity and dispersal limitation, while a slowly decreasing distance-decay relationship suggests cosmopolitanism.
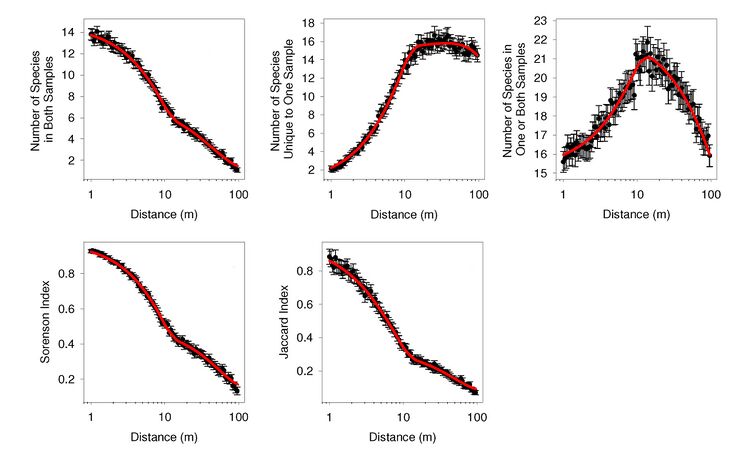
Both to make ecological inferences from distance-decay relationships and infer taxa-area scaling from them, a quantitative understanding of how distance-decay relationships arise is necessary. We have been developing and validating a theory to address this matter. Our theory assumes that the distributions of taxa can be approximated by polygons and disjoint points, and that samples are collected at random locations. Under these assumptions, we have proven that the average number of taxa (i) shared between a pair of samples, (ii) unique one sample in a pair of samples, and (iii) occurring in at least one sample vary as a the sum of a quadratic function of distance and a non-quadratic function, all divided by a function of the shape of the region that was sampled. We have also validated our theory using six large data sets. For these data sets, we have found that the observed distance-decay relationships conform remarkably well to the predictions of our theory (Distance-Decay Figure 1; details provided in the manuscript linked here).
The next steps in this project are twofold. First, we plan to apply our theory to metagenomic data, which is now possible using the recently-constructed operational taxonomic units (see Metagenomic Sequence Classifier, below). From these applications, it should be possible to make inferences about the effects of habitat heterogeneity and dispersal limitation on the distributions of microbes. Second, we plan to use the models from the distance-decay theory to construct estimators of richness-area scaling for microbes, a project that is ongoing.
Update on the metagenomic simulator
In the previous report, we describe our metagenomic simulator, which is being developed by Samantha Riesenfeld under the direction of Katie Pollard. The simulator has undergone major revisions, based on discussions with iSEEM members about what types of simulated data would be most useful. One of the major changes concerns the Reference Database, which is the set of complete gene sequences used to mimic the universe of "known", complete gene sequences. For each simulation, these sequences are aligned (as peptides) with the peptide reads simulated during that run. Originally, the Reference Database was chosen randomly from the set of AMPHORA Reference Sequences for the chosen gene. The Reference Database now consists of 100 gene sequences, 75 of which are selected (in decreasing order) according to the amount of phylogenetic diversity that the corresponding taxa contribute to the entire set of AMPHORA taxa. The rest of the sequences in the Reference Database are selected uniformly at random from the AMPHORA sequences. Another major change relates to the sequences that are seeds for the simulated reads. Originally, these sequences were chosen from the Reference Database; now they are chosen uniformly at random from all AMPHORA Reference Sequences for the gene. An additional step has also been added in the processing of the generated reads: before being aligned, they are scanned by an AMPHORA script to pick up only those reads that appear to come from the desired gene. Finally, the set of reads is filtered so that at most one read for each original, unfragmented sequence appears in the output alignment. This last step is optional and may be skipped in order to create alignments of larger numbers of reads. Additional information about the updated parameters and the iSEEM discussion of them appear on the simulation parameters wiki page.
Forty simulated data sets are currently available here to iSEEM members. This data contains twenty simulations for the rpoB gene and twenty simulations for the smpB gene. We plan on creating simulations for additional AMPHORA Marker genes as needed. In each available simulation, 20 gene sequences were chosen uniformly at random from AMPHORA to be fragmented into 500 simulated reads. The reads were then scanned and filtered as described above, in order to provide simple testing data. Alignments with more simulated reads can easily be created, either from the current data sets or by running additional simulations. Steve Kembel of the Green Lab and Tom Sharpton of the Pollard Lab are currently using the simulated data sets to test phylogenetic inference methods, in order to judge their suitability as tools in the identification of OTUs in metagenomic data. As phylogenies are the starting point for many other types of metagenomic analyses as well, we plan on using the simulation data to do more extensive testing of phylogenetic inference methods.
Metagenomic Sequence Classifier
Our general goal with this project is to develop a phylogenomic pipeline that will classify, or annotate, metagenomic sequences into protein families. In the final product, the classified sequences will be used to explore biodiversity, discovery novelty, and describe trait-based community assembly across gene family space. Development and subsequent exploration will be primarily undertaken by Thomas Sharpton under the guidance of Katie Pollard and Jonathan Eisen.
1. Generalized workflow

Phylogenomic annotation is a well documented process, requiring the classification of sequences into homologous families, sensitive and specific family member sequence alignment, and characterization of the evolutionary relationships between members. Unfortunately, all aspects of this generalized process are frustrated by the short and fragmentary nature of metagenomic reads. To circumvent these limitations, we have documented a bioinformatic strategy (see right) that first references well-annotated whole genome data to characterize known protein families as probabilistic models (Hidden Markov Models or HMMs) and then uses these models to guide the classification, alignment and phylogenetic analysis of metagenomic reads.
The endpoint of the workflow is a database cataloging phylogenies of gene families (inclusive of reference and metagenomic data) which will be leveraged to investigate questions pertinent to the discovery of novel gene family diversity and trait based community assembly. The workflow is currently in development using the well-defined SSU-rRNA gene family as a test case (see below)
2. Methodological Classification Tests using SSU-rRNA (w/ Green lab)
A working group composed of Thomas Sharpton and Josh Ladau of the Pollard lab and Steve Kembel and James O'Dwyer of the Green lab is interested in developing a generalized methodology to identify operational taxonomic units (OTUs) from metagenomic sequence data for a variety of purposes (See above sections for discussion on application). This process provided a test case for gene family classification, specifically the SSU-rRNA family, which is commonly leveraged for taxonomic classification.
We tested several methods of classification and alignment using a subset of the complete global ocean survey (GOS) metagenomic library as a trial dataset and high quality SSU-rRNA sequences made available from the Ribosomal Database Project. Our work finds that leveraging statistical profiles that define gene family evolutionary diversity (e.g., Context Free Grammars and Hidden Markov Models) provides stronger inferences of classification and alignment than other methods considered (progressive and iterative sequence alignment). Specifically, we've demonstrated the following:
- the higher the quality of the reference sequence data used to construct profiles, the better the classification sensitivity
- reads are aligned to profiles independently, ensuring that a noisy sequence does affect not majority of the alignment signal
- short reads can be locally aligned to profiles in a high-confidence fashion
- at least for SSU-rRNA, the profiles must be assembled independently for each major life domain (structural variation is high across domains)
In short, probabilistic family profile based methods provide a robust means of classifying and aligning short, fragmented metagenomic reads into well defined families. We have also constructed quality control tools of the resulting alignments, which excise particularly noisy sequences and columns to ensure strong signal for subsequent analysis. The working group is currently testing various methods of phylogenetic analysis on the resulting alignments. Ultimately, these methods will be applied to other gene families to phylogenomically annotate as many reads as possible.
Wu Lab
Zorro: Probability-Based Multiple Sequence Alignment Masking
The quality of multiple sequence alignment may have a large impact on the final phylogenetic tree. Marking and removing the ambiguous regions of the alignment, a step known as masking and trimming in phylogenetics, can be beneficial, especially for divergent sequences that are difficult to align. Traditionally, this was done by manual curaton, which has simply become infeasible for genome-level phylogenetic analysis. In AMPHORA, we used masks embedded with the Hidden Markov Models of the protein families to automate the masking and trimming process. The mask, although needs to be generated only once for each protein family, still relies on skilled but sometimes arbitrary manual curation. Recently, we developed a probability-based algorithm named Zorro to assess the quality of the alignment and use it to mask the regions of uncertainty. Previously we have shown that Zorro outperforms Glbocks in both specificity and sensitivity (Gblocks is a program that masks alignments based on the conservation score of each column in the alignment and a set of ad hoc rules).
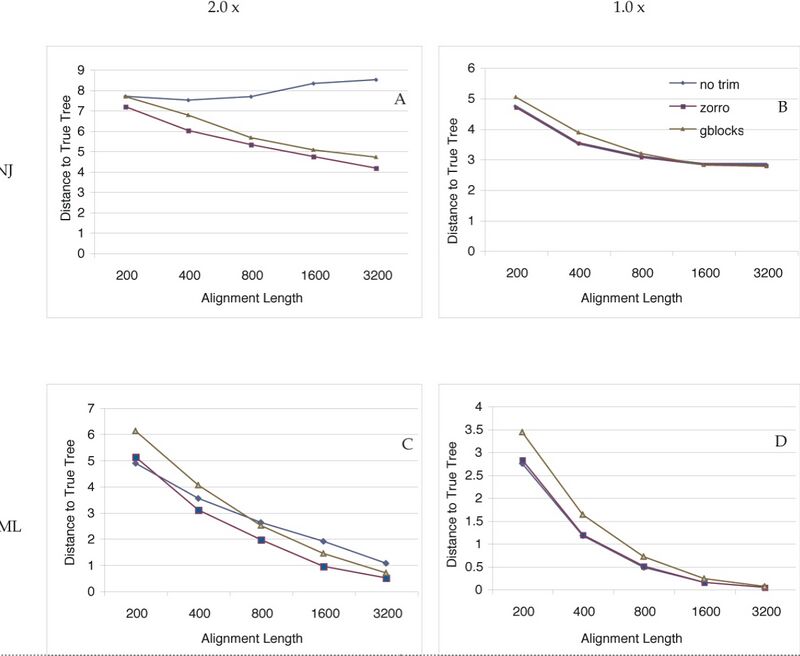
Since trimming could in theory remove phylogenetically informative sites, we tested whether trimming by Zorro actually led to better phylogenetic trees. In a simulation study, we subjected multiple sequence alignments to three different treatments: no trimming, trimming by Zorro or trimming by Gblocks. We then reconstructed neighbor-joining and maximum likelihood trees and measured the accuracy of the trees by calculating their Robinson Foulds topological distances to the true tree. To test the relative performance of trimming, we also permuted the alignment length, tree shape (symmetric vs. asymmetric) and the degree of divergence of the protein sequences. For each unique permutation, 200 simulations were run and the average Robinson Foulds distance was calculated. A total of 8,000 trees were estimated and compared. The results are shown in the figure.
Our simulation study shows that trimming by Zorro or Gblocks significantly improves the phylogenetic trees when the sequences to be compared are relatively divergent (Figure panel A and C). Trimming decreases the topological distances to the true tree with respect to the complete alignment. This is most likely due to increased signal-to-noise ratio after elimination of problematic regions. For the complete alignments, the signal-to-noise ratio stays the same and the neighbor-joining trees estimated from them do not get better with the increasing alignment length. The improvement in phylogenetic accuracy is most pronounced in the neighbor-joining trees of the relatively divergent sequences. Maximum likelihood trees also benefited from trimming, but the impact was smaller. This is because the maximum likelihood method can accommodate to a certain degree the evolution rate heterogeneity and extract some useful phylogenetic information from the difficult to align regions that are removed from the alignments. Alignment length is also an important factor to consider. Short alignments in general benefit less from trimming than longer alignments, because the loss of informative sites offsets the gain of the signal-to-noise ratio to a larger extent in the shorter alignments. This suggests that for phylogenomic studies, when sequences from multiple genes are concatenated to generate "mega-alignment", trimming should always be carried out to obtain the best phylogeny possible.
Figure shows that Zorro outperforms Gblocks in terms of improving the phylogenetic accuracy. Gblocks uses the conservation score of each position and a set of rules (e.g, the maximum number of contiguous nonconserved position is 8) to distinguish conserved blocks from highly divergent regions. However, different protein families have different functional or structural constraints and no single set of rules will fit all the proteins equally well. In addition, these rules are ad hoc and there seems to be little theoretical basis behind them. Therefore, Gblocks might remove too much information from the alignments. As a result, in some cases trimming by Gblocks actually shows detrimental effects on the final trees.
For sequences of less divergence, trimming does not seem to help (Figure panel B and D). This is expected because aligning these sequences is no longer an issue and the percentage of alignment that are trimmed is relatively small. We note that under no circumstances we have tested here, trimming by Zorro ever led to significant worse phylogenetic trees. Therefore, trimming by Zorro seems to be justified for phylogenetic inferences, especially of deep relationships such as these among the major bacterial lineages. Because of its speed, neighbor-joining is often chosen over the maximum likelihood method for large-scale phylogenetic analyses. Our study suggests that trimming leads to significant better neighbor-joining trees and therefore should be used in these phylogenomic studies.
We are in the process of finishing up writing the manuscript and we expect to publish the results soon.
C. Communications, Collaboration, Outreach and Education
Publications supported by this grant
- J.P. O'Dwyer and J.L. Green, "Field theory for biogeography: a spatially-explicit model for predicting patterns of biodiversity", Ecology Letters, In Press
- pdf linked here: [2]
- Wu D, Hugenholtz P, Mavromatis K, Pukall, Dalin E, Ivanova NN, Kunin V, Goodwin L, Wu M, Tindall BJ, Hooper SD, Pati A, Lykidis A, Spring S, Anderson IJ, D’haeseleer P, Zemla A. Singer M, Kerfeld C, Lang E, Gronow S, Chain P, Bruce D, Rubin EM, Kyrpides NC, Klenk H-P, Eisen JA. phylogeny-driven genomic encyclopedia of Bacteria and Archaea. Revision submitted.
Talks
- Kembel, S.W. and Green, J.L. "Ecology without species". Invited talk, Organized Oral Session on ‘Species interactions and community ecology in the context of relatedness’, Ecological Society of America Annual Meeting, Albuquerque NM, August 2009.
- Green, J.L. "Lady Lumps's Mouthguard". Invited talk, Rocky Mountain Biological Laboratory, Gothic, Colorado.
Outreach
- Katie Pollard led a journal club and gave two talks about comparative genomics to the National Student Leadership Conference (summer program for high school students): http://www.nslcleaders.org/
- Jessica Green is external faculty at the Santa Fe Institute and spent the month of July there discussing iSEEM related research with resident and visiting faculty.
- Jessica Green spent two days at the University of Colorado in Boulder. During this time she met with Cathy Lozupone and Rob Knight to discuss iSEEM related research. Knight felt that our approach to building phylogenies (as described in this progress report) was excellent.
- Jessica Green was the moderator of a symposium at the Ecological Society of America titled "The Rainforest Within: Biodiversity of the Human Body and its Relationship to Health and Disease" which was covered by Nature News (http://www.nature.com/news/2009/090807/full/news.2009.808.html) and their blog (http://blogs.nature.com/news/blog/2009/08/esa_2009_sweater_by_grandma_gu.html). Metagenomic approaches to understanding community assembly was a centerpiece of the session.
Working Groups
- S.Kembel led a workshop on ‘Ecological approaches to analyzing complex community datasets’ for 80 FESIN (Fungal Environmental Sampling and Informatics Network) members, BSA/MSA annual meeting, Snowbird UT, July 2009.
II. Group meetings
Weekly meetings
- Notes 7_1_09
- Notes 7_8_09
- Notes 7_15_09
- Notes 7_22_09
- Notes 7_29_09
- Notes 8_5_09
- Notes 8_12_09
- Notes 8_18_09
- Notes 8_19_09
- Notes 8_26_09
- Notes 9_2_09
- Notes 9_9_09
- Notes 9_16_09
- Notes 9_23_09
- Notes 9_30_09
Other meetings
- Andrey Kislyuk from the Weitz at Georgia Tech came to visit the Davis and UCSF groups to discuss metagenomic binning methods.
- Joshua Ladau and James O'Dwyer met at the University of Oregon to discuss models of microbial spatial distributions and methods for inferring microbial diversity.
III. Any unexpected challenges that imperil successful completion of the Outcome
CAMERA interactions
CAMERA provided login (shell) accounts for a few members of each iSEEM lab. They created accounts for the users and added groups for each associated lab. Each lab also has a mount point created on the CAMERA filestore. One point person per lab was given database access.