Biomod/2015/Kansai/Sources

| HOME | Team | Project | Design | Sources | Experiment | protocol |
Protocol
・Materials Staples DNA were purchased from IDT ( U.S.A )
Each staple DNA were dissolved by sterilized water that the concentration of each solution might be 100 μM.

These solutions were taken 2 μL each and mixed.

The volume of the solution was diluted with sterilized water to 500μL. we prepared staple DNA mixture.The sample (50 μM) was prepared mixing M13mp18 (4nM, Takara, Japan), staple DNA mixture (20nM) and 1×TAE/Mg2+ (12.5 mM). This mixture was kept at 90˚C for 10 minutes and cooled from 90˚C to 25˚C at a rate of -1.0˚C/min to anneal the strands.

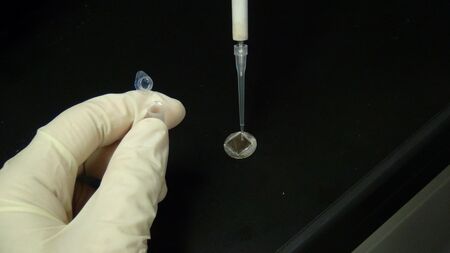
The annealed mixture (1μL) was deposited on freshly cleaved mica, additional 1X TAE/Mg2+ buffer (40 µL) was added.
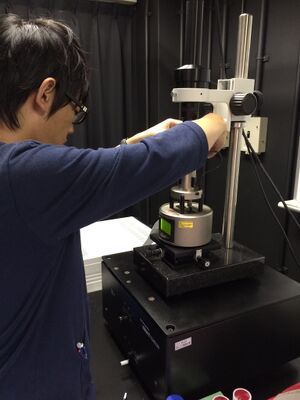
We observed this product by AFM. the imaging was performed in the fluid Tapping mode with a BL-AC40TS tip (Olympus, Japan) and performed on a Multimode 8/ Nanoscope system (Bruker AVS).

<html>
<a href="#top">↑ Back to top</a>
</html>