BioMicroCenter:DataManagement
| HOME -- | SEQUENCING -- | LIBRARY PREP -- | HIGH-THROUGHPUT -- | COMPUTING -- | DATA MANAGEMENT -- | OTHER TECHNOLOGY |
Data Management
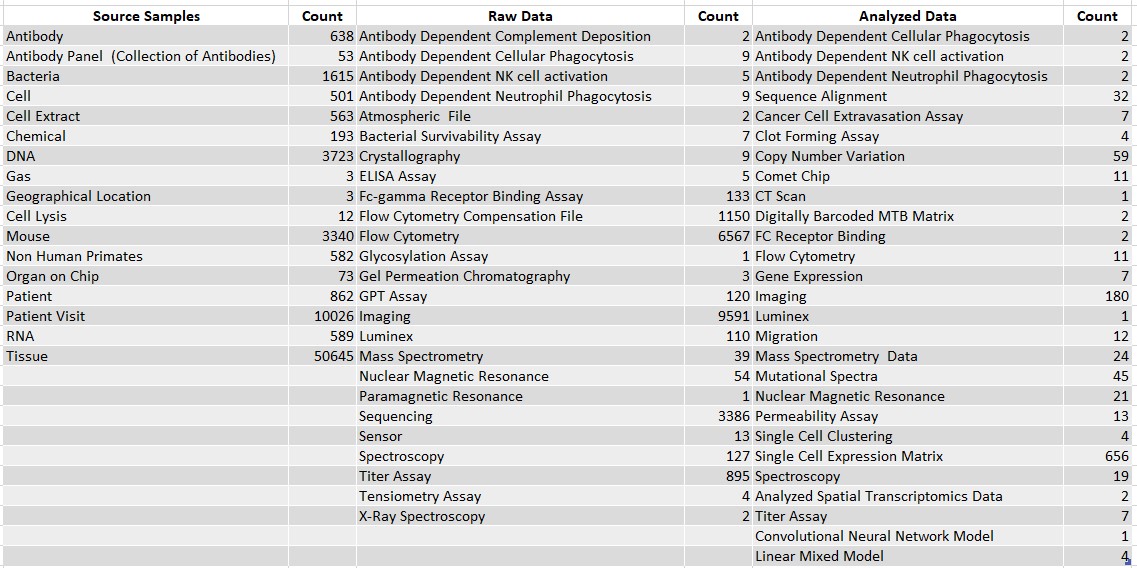
The data managers at the BioMicro center are responsible for the organization and storage of scientific data. We do this using NExtSEEK, a FAIR-compliant data management system, allowing for the data to be findable, accessible, interoperable, and reusable, ensuring the quality of the metadata being stored. NextSEEK is one of the only FAIR-compliant storage systems and is built to handle a variety of types of scientific data types, a contrast to other commercial databases that are built for one specific data type.
Currently, we are working on five different projects comprising of numerous different labs;
- The MIT SuperFund Research Program (SRP)
- Metastasis Network (MetNet)
- IMPAcTB
- Break Through Cancer (BTC)
- Cancer Systems Biology Consortium (CSBC)
The data managers at the BioMicro Center work with researchers to submit data in real time, making the data deposition and submission for publication process as simple as possible.
Published data is linked and available on FAIRDOMHub, at the links below:
More Information
To learn more about what we do and how we implement FAIR compliance, go here.
For those interested in learning more about NExtSEEK and FAIRDOMHub, navigate to this page.
For instructions on how to use NExtSEEK, go to Instructions and Tutorial.
| Project | Count |
|---|---|
| IMPAcTB | 57,622 |
| MetNet | 3,450 |
| SRP | 32,281 |
| BTC | 772 |
| CSBC | 240 |
| Statistic | Count |
|---|---|
| Samples | 97,082 |
| Raw Data Files | 22,528 |
| Analyzed Data Files | 1,126 |
| FAIRDOMHub Studies | 29 |
| Public FDH Studies | 18 |