BISC209/S12: Antibiotics
Antibiotic production
Many microbes secrete antimicrobial compounds to help them compete with other microorganisms for habitat. Some of the bacteria that are common antibiotic producers are the Actinomycetes (including Streptomycetes species), many of the Bacillus species, and the fruiting myxobacteria, to name just a few among many, many antibiotic producing bacteria. You can also test for the opposite: the sensitivity of your soil organisms (or known stock bacteria) to manufactured or secreted antibiotics.
(This testing will take 3 weeks.)
Week 1:
Identify how many potential antibiotic producers you might have. Definitely test any isolates that are likely to be Actinomycetes, Myxobacteria, or Bacillus. It might be wise to test all 4 of your isolates since the soil is the main source of microbes that supply the world's antibiotics. It's possible that you might discover the next great antimicrobial drug and get very rich by selling the patent for your discovery to a drug company. Remember that the discovery of penicillin was completely accidental.
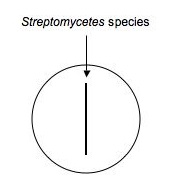
Using aseptic technique, transfer an isolated colony (possible Streptomyces, etc.) that's likely to be an antibiotic producer to a microfuge tube containing 500μL of sterile water. Vortex to mix. Using a sterile swab, dip the swab in the diluted bacteria and make an inoculation (as shown below) down the middle of a plate of nutrient agar. Make a second plate exactly like the first for each isolate to be tested. Label them carefully and incubate the plates for ~1 week at RT.
Week 2
TO BE PROVIDED in LAB 7:
3 fresh cultures of Eschericia coli (Gram negative), Staphylococcus epidermidis (Gram positive) and Micrococcus luteus (Gram positive) grown in nutrient broth.
NEED TO MAKE (Optional): If you want to test any of your isolates for sensitivity to an antibiotic producer, start a log phase broth culture in appropriate media so that it will be ready to use in the next lab.
PROTOCOL
- Use a sterile swab and aseptically apply a line of inoculation of each of the provided broth cultures of : E. coli, Micrococcus, and S. epidermidis. as shown below. Draw a line perpendicular to the antibiotic producer's (Streptomyces) inoculation. Be careful not to touch the antibiotic producer's growth. Using the same swabs, inoculate a new NA plate (one plate for all three cultures)by making a line across the plate for E. coli, Micrococcus and S. epidermidis. Incubate this plate along with your test plate. It will serve as a control to make sure that lack of growth is due to antibiotic sensitivity and not to no living cells in the inoculum.
-
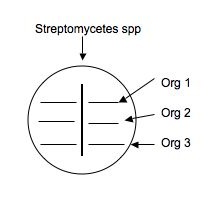
- Repeat this procedure on another plate using broth cultures made of any of your isolates that you want to test for sensitivity to your antibiotic producers,
- Label a template in your notebook with which organism is where on each plate.
- Incubate for another week.
Week 3
- Examine the plate and look for evidence of inhibition of growth of "test" organisms near the antibiotic producer's midline streak.
- Draw the results and evaluate whether or not there was evidence that an antibiotic was produced by the organism and, if so, which of the bacteria tested were sensitive to it and to what degree. If you found no inhibition of growth, does that mean that your potential antibiotic producer does not secrete an any antimicrobial compounds? Why or why not?