BIO254:Phototransduction
Definition
Phototransduction is the process through which photons, elementary particles of light, are converted into electrical signals. Visual phototransduction occurs in the retina through photoreceptors, cells that are sensitive to light. The membrane potential of a photoreceptor hyperpolarizes in response to light, causing a reduction in the amount of neurotransmitter released by the photoreceptor onto downstream neurons. Phototransduction thus enables the photoreceptor to encode a light stimulus as a chemical output.
Photoreceptor Cells
Two types of photoreceptors: rods and cones
There are two types of photoreceptors distributed unevenly across the retina: rods and cones. Rods are very sensitive cells specialized for night vision. In bright light conditions the response of the rods is saturated and cones, faster but less sensitive photoreceptors, mediate day vision.
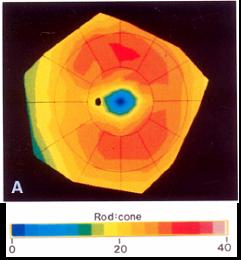
Rod-cone distribution in the human retina (Curcio, CA et al. 1990)
The rod:cone ratio is lowest in the foveal region and higher in the periphery. The black dot to the left of the fovea represents the optic disk, the site where axons exit the retina.
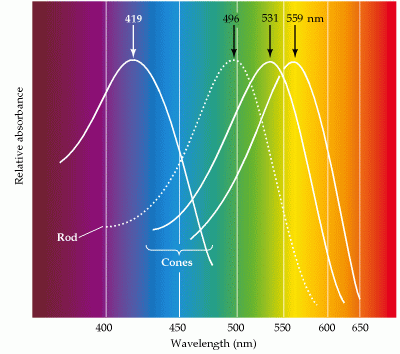
There are three types of cones, each one of them responding best to different wavelengths (short, middle, and long). Their combined responses generate color vision.
Absorbance spectra of rod and cone photoreceptors (Purves, p.239, 2001)
Opsin, the key molecule for phototransduction
Both rods and cones contain opsin, a G protein-coupled receptor. Opsin is bound to a light-absorbing chromophore, 11-cis-retinal (an aldehyde of vitamin A). Different types of opsins are involved in transducing light of different intensities and wavelengths. Each photoreceptor expresses only one type of opsin. Rhodopsin is present in rods and transduces dim light while photopsins are present in cones and generate color vision.
Phototransduction step by step
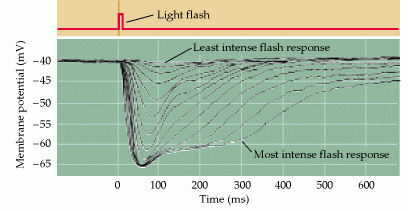
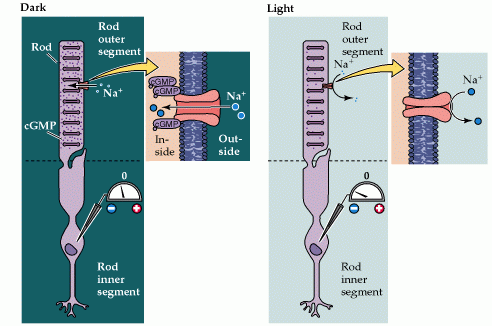
In the absence of light, the photoreceptors are depolarized to a membrane resting potential of -40mV. Light will hyperpolarize the plasma membrane of the photoreceptor to -70mV (Figure 1). This stimulus-induced hyperpolarization is a distinctive characteristic of the photoreceptor response, as many other neuronal types depolarize when stimulated.
Figure 1: An intracellular recording from a single cone stimulated with different amounts of light. Each trace represents the response to a brief flash that was varied in intensity. At the highest light levels, the response amplitude saturates. (Neuroscience, Purves et al., 2001)
A key second messenger molecule responsible for maintaining a depolarized rest state in photoreceptors is the nucleotide cyclic guanosine 3’-5’ monophospate (cGMP). High cGMP levels keep cGMP-gated ion channels in the open state and allow them to pass an inward Na+ current (Figure 2).
Figure 2: Cyclic GMP-gated channels in the outer segment membrane are responsible for the light-induced changes in the electrical activity of photoreceptors. (Neuroscience, Purves et al., 2001)
Phototransduction involves three main biochemical events:
Light entering the eye activates the opsin molecules in the photoreceptors
Upon photon absorption, 11-cis-retinal undergoes an isomerization to the all-trans form, causing a conformational change in the rhodopsin. The activated rhodopsin is called metarhodopsin II.
The precursor for 11-cis-retinal is all-trans-retinol (vitamin A). A diet rich in vitamin A is crucial for vision, since vitamin A cannot be synthesized by humans.
Activated rhodopsin causes a reduction in the cGMP intracellular concentration
The cytoplasmic cGMP levels are controlled by cGMP phosphodiesterase, an enzyme that breaks down cGMP. In the dark, the activity of this enzyme is relatively weak. When the photoreceptor is exposed to light, metarhodopsin II stimulates the activity of cGMP phosphodiesterase via transducin, a G protein. GDP-bound inactive transducin will exchange GDP for GTP following interaction with activated rhodopsin. GTP-bound active transducin will increase the activity of cGMP phosphodiesterase. The result is decreased levels of cGMP in the cytoplasm.
The photoreceptor is hyperpolarized following exposure to light
Decreased levels of cGMP cause the closing of cGMP-gated ion channels which will lead to membrane hyperpolarization.
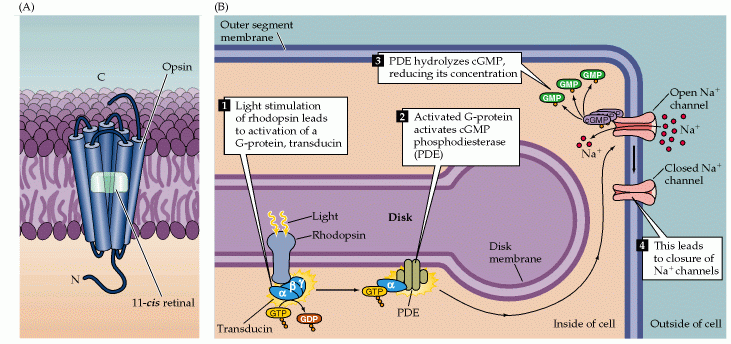
Figure 3 summarizes the phototransduction cascade.
Figure 3: Details of phototransduction in rod photoreceptors. (A) The molecular structure of rhodopsin, the pigment in rods. Rhodopsin is a G-protein coupled receptor consisting of opsin (a seven transmembrane domain protein) and 11-cis-retinal (a covalently bound chromophore). (B) The second messenger cascade of phototransduction. 1. Light stimulation of rhodopsin in the receptor disks leads to the activation of a G-protein (transducin). 2. The GTP-bound alpha subunit of transducin activates a phosphodiesterase (PDE). 3. The activated phosphodiesterase hydrolyzes cGMP into GMP, reducing its concentration in the outer segment and leading to the closure of sodium channels in the outer segment membrane. (Neuroscience, Purves et al., 2001)
Termination of the phototransduction cascade
The light response is terminated by several mechanisms.
(1) Inactivation of rhodopsin occurs through phosphorylation by the opsin kinase, followed by the binding of arrestin to phosphorylated rhodopsin.
(2) Inactivation of transducin occurs through the hydrolysis of bound GTP to GDP (Tα-GTP to Tα-GDP) via an intrinsic GTPase activity that is accelerated by the GTPase activating protein RGS9 (regulator of G-protein signaling).
(3) Inactivation of phosphodiesterase (PDE) is coupled to the inactivation of transducin. Inactivated transducin (Tα-GDP) dissociates from PDE, resulting in a cessation of PDE-mediated cGMP hydrolysis.
(4) Activation of guanylate cyclase by guanylate cyclase activating protein (GCAP) restores cGMP levels and thus promotes the re-opening of cGMP-gated channels.
Amplification in the phototransduction cascade
The activation of a single rhodopsin by a single photon is sufficient to cause a significant change in the membrane conductance. This is possible due to amplification steps present in the transduction cascade.
A single photoactivated rhodopsin catalyses the activation of 500 transducin molecules. Each transducing can stimulate one cGMP phosphodiesterase molecule and each cGMP phosphodiesterase molecule can break down 10^3 molecules of cGMP per second. Therefore, a single activated rhodopsin can cause the hydrolysis of more than 10^5 molecules of cGMP per second.
Human disorders of phototransduction
Bradyopsia (or ‘slow vision’) is a condition that results from mutations in genes encoding the transducin-inactivating protein RGS9 or the RGS9 anchor protein (R9AP) (Nishiguchi et al., 2004). By accelerating the inactivation of transducin, RGS9 and R9AP play an important role in the termination of phototransduction and recovery of the light response. Patients with bradyopsia have trouble adjusting to changing light conditions, experiencing a temporary blindness when first exposed to bright light. This disorder also manifests as an inability to sense the movement of objects in a bright environment.
Retinitis pigmentosa is an inherited disorder characterized by degeneration of photoreceptor cells and accumulation of retinal pigments. This disorder, which often leads to blindness, can result from mutations in a variety of genes expressed in photoreceptors (Hamel, C. 2006). Early genetic studies of autosomal dominant retinitis pigmentosa revealed a series of point mutations in the rhodopsin gene that were associated with the disease (Dryja, TP et al. 1991). The integrity of the light-sensing molecule rhodopsin is thus crucial for the proper activation of the phototransduction cascade.
Retinitis pigmentosa
Congenital stationary night blindness is an inherited disorder that affects rod photoreceptors and impairs vision under low-light conditions. This disorder may result from missense mutations in the rhodopsin gene that cause the mutated rhodopsin protein to constitutively activate transducin (Dryja, TP et al. 1993; Rao, VR et al. 1994). Persistent activation of the phototransduction cascade limits the fidelity of the light response by rod photoreceptors.
References
Principles of Neural Science, by Kandel, Schwartz, & Jessell, 4th edition, 2001.
Neuroscience, by Purves, Augustine, Fitzpatrick, Katz, LaMantia, McNamara, & Williams, 2nd edition, 2001.
The Quest for Consciousness, by Christof Koch, 2004.
Bio254 Lecture notes, by Tom Clandinin, October 23, 2006.
Curcio, CA et al. Human photoreceptor topography. J Comp Neurol. Feb 22;292(4):497-523 (1990)
Dryja, TP et al. Mutation spectrum of the rhodopsin gene among patients with autosomal dominant retinitis pigmentosa. Proc Natl Acad Sci U S A. 88(20):9370-4 (1991)
Dryja, TP et al. Heterozygous missense mutation in the rhodopsin gene as a cause of congenital stationary night blindness Nature Genetics 4: 280 - 283 (1993)
Hamel, C. Retinitis pigmentosa. Orphanet J Rare Dis. 11:1-40 (2006)
Nishiguchi, KM et al. Defects in RGS9 or its anchor protein R9AP in patients with slow photoreceptor deactivation. Nature 427:75-78 (2004)
Rao, VR et al. Rhodopsin mutation G90D and a molecular mechanism for congenital night blindness. Nature 367: 639 - 642 (1994)
Recent updates to the site:
List of abbreviations:
- N
- This edit created a new page (also see list of new pages)
- m
- This is a minor edit
- b
- This edit was performed by a bot
- (±123)
- The page size changed by this number of bytes
21 May 2026
|
|
21:46 | Hu:Publications 3 changes history +405 [Hugangqing (3×)] | |||
|
|
21:46 (cur | prev) +1 Hugangqing talk contribs | ||||
|
|
21:46 (cur | prev) +56 Hugangqing talk contribs | ||||
|
|
21:42 (cur | prev) +348 Hugangqing talk contribs | ||||
|
|
21:40 | Hu 2 changes history +195 [Hugangqing (2×)] | |||
|
|
21:40 (cur | prev) +105 Hugangqing talk contribs | ||||
|
|
03:26 (cur | prev) +90 Hugangqing talk contribs | ||||
19 May 2026
|
|
02:49 | Hu:Members 2 changes history +178 [Hugangqing (2×)] | |||
|
|
02:49 (cur | prev) −563 Hugangqing talk contribs | ||||
|
|
02:47 (cur | prev) +741 Hugangqing talk contribs | ||||
| 02:45 | Upload log Hugangqing talk contribs uploaded File:Naved.png (Naved) | ||||