Alex J. George Week 9
From OpenWetWare
Jump to navigationJump to search
Question
Do the amino acid residue differences within the variable regions of progressors and nonprogressors alter its structure significantly enough to affect its function?
- We obtained the amino acid sequences for our subjects from the Bedrock website
- Subjects 13,12,2,11,10,4
- After obtaining all of our protein sequences, we ran a multiple sequence alignment to find the differences and similarities
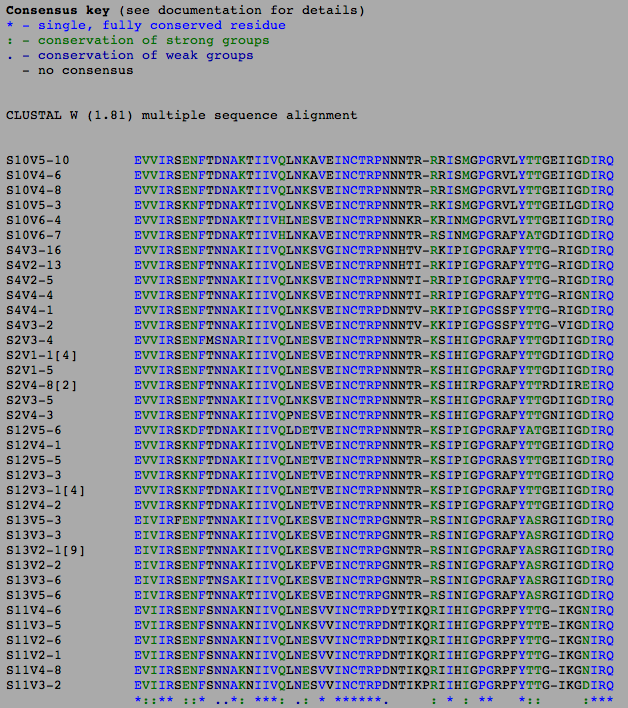
- There are fewer differences than the DNA sequences because some of the mutations in the DNA could have been nonsense mutations in which they didn't change the amino acid.
- From Chapter 6, a couple of the procedures are applicable. The primary structure analyses can help us determine the hydrophobic and hydrophilic regions within the protein. For example, in gp120. the V3 region should appear as hydrophilic since the loop sticks out from the protein and is essential to cell interaction.
- We have chosen to analyze two sequences in depth- one from a rapid progressor (S11, V4-6) and one from a nonprogressor (S12,V4-1). We decided on these two sequences because Subject 11 differed from the standard more often than the other groups. Conversely, Subject 12 seemed to have the most sequences that were a consensus for the protein.
Final Presentation: File:Protein Structure Final Powerpoint-Alex and Bobby.ppt
Individual Journals
- No Week 1 Journal
- Alex J. George Week 2
- Alex J. George Week 3
- Alex J. George Week 4
- Alex J. George Week 5
- Alex J. George Week 6
- Alex J. George Week 7
- Alex J. George Week 8
- Alex J. George Week 9
- Alex J. George Week 10
- Alex J. George Week 11
- Alex J. George Week 12
- Alex J. George Week 13