User:Thorntca
Casey Thornton's Discussion Questions: Taking a closer look
Week 9: Cap independent translation
Paper:
Discussion Questions:
1. This paper proposes a mechanism where m6A markers are utilized to promote cap-independent translation. This may confer increased fitness for translation of some necessary genes. My question is: Based on this paper, what can be hypothesized about how genes are chosen for this process, and what subset of genes might utilize this mechanism?
2. In figure 2, Meyer et al demonstrate that cap-independent translation was dependent on position of the m6A element in the 5' UTR region. Why might that be? And could the differential expressivity and different m6A positions correlate with secondary structural elements of the mRNA transcript?
3. On that same note - how does considering the secondary structural elements of RNA contribute to this cap-independent translation story? The authors tested cell stress as a function of heat shock response. Heat, and other cell stress environments such as pH, salinity, and detergent variability can alter secondary structures. Is it feasible to hypothesize that the m6A modification could correlates to a unique mRNA secondary structure, which is susceptible to cap-independent translation?
Citations:
- Meyer KD, Patil DP, Zhou J, Zinoviev A, Skabkin MA, Elemento O, et al. 5’ UTR m6A promotes Cap-independent translation. Cell. 2015;163:999–1010. doi: 10.1016/j.cell.2015.10.012.
Week 8: Predestination in the cell cycle
John Calvin contemplates how cells decide between proliferation and quiescence.
Paper:
Cell predeterminism in the news:
Discussion Questions:
1. Cell lines require authentication, in order to provide evidence that your measurements and observations correlate with a standard cell line that can be reproduced. I have worked in mouse research, and I know that mouse breeding schemes vary widely between laboratories. My question is: Is there a authentication standard for reporting the genetic background of the mice used in a publication?
- We talked with Dr. Amanda McCullough, and found that there are not the same standards for mouse authentication upon study publication. The authentication standards that have become common-place for cell lines have become standard only in the past few years. There are not standards for mice yet, but that may be developed in the future to improve reproducibility.
2. This paper highlighted my concerns about single cell approaches in the molecular sciences. I am extremely interested in parsing apart the unique genomes of the somatic tissues in oreganisms, but this paper has raised the concern that single-cell studies produce low sample sizes. My question is: How should the scientific community weigh the value of high resolution single-cell data versus the inherent low population sampling of single-cell measurement?
3. How does this finding inform therapeutic approaches to cell-cycle control? The mother cell may determine daughter cell fate, but how can this information be utilized to alter the signals that the mother cell passes on, or the daughter cell memory?
Citations:
- Yang H.W., Chung M., Kudo T., Meyer T. Competing memories of mitogen and p53 signalling control cell-cycle entry. Nature. 2017 Sep 21;549(7672):404-408. doi: 10.1038/nature23880.
- Purvis J.E., Kedziora K.M., Cell biology: The persistence of memory. Nature. 2017 Sep 21;549(7672):343-344. doi: 10.1038/nature23549.
Week 7: Hsp90 and its role in human disease
Paper:
Hsp90 in the news:
Discussion Questions:
1. This paper discusses how severe phenotypes associate with Hsp70 and relatively moderate phenotypes associate with Hsp90. Can this conclusion be simplified to: Hsp70 associates with unfolded proteins, which is a severe result of a gene mutation, and Hsp90 associates with partially folded proteins, which corresponds with a less severe mutation. What conclusion does this paper come to that is more than: severely mutated proteins cause severe phenotypes?
- This paper confirms that "simple" paradigm - and uses it to show that in the case where proteins are not so mutated as to be Hsp70 associated and severely disfunctional, then they bind Hsp90 and are more regulatable by Hsp90 in the less severe disease state. Hsp90 specifically can alter the presentation of a genetic disease - providing a mechanism for how environmental stress on Hsp90 can result in phenotypic variation in identical genetic diseases.
2. What therapeutic approaches can be taken now that we understand more about how Hsp90 buffering contributes to genetic disease phenotype presentation?
3.What is the LUMIER assay and how is it used to elucidate protein:protein interactions?
- The LUMIER assay is a luminescence-based technique for mammalian interactome mapping and quantification. Simply, one protein is enriched in one cell extract, and then that extract is tested for enrichment of a second protein - in order to indicate whether these proteins form a complex. Useful resources for understanding this assay are as follows:
Citations:
- Karras G.I., Yi S., Sahni N., Fischer M., Xie J., Vidal M., D'Andrea A.D., Whitesell L., Lindquist S. HSP90 Shapes the Consequences of Human Genetic Variation. Cell. 2017;168:856-866.
- Researchers uncover a role for HSP90 in gene-environment interactions in humans. Phys.org - News and Articles on Science and Technology, 23 Feb. 2017, phys.org/news/2017-02-uncover-role-hsp90-gene-environment-interactions.html.
Week 6: One Ring to Rule Them All - Cohesin mediates TAD formation
Paper:
Cohesin in the news:
Discussion Questions:
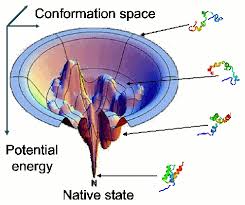
1. Recent developments in protein folding predication have centered around the "multi-state energy lanscape funnel" that centers around the idea that proteins traverse down the tunnel through a variety of energetically/enthalipcally favorable intermediate states where it can get trapped - or continue to the most favorable state - the folded folded state. How does DNA folding follow this same paradigm? Could compartmentalization of histone markers in DNA be likened to secondary structure in proteins, and TADs correspond to teritary structure? How does this comparison change what we think about DNA occupying "one state"?
2. The authors describe how removal of cohesin leads to loss of loop domains, and an increase of the "plaid" shown in Hi-C, which corresponds to increased compartmentalization of loci with similar histone marks. The native loops domains are proposed to promote gene-gene interactions that mediate gene expression in a cell, but no phenotypic data is provided for cells with cohesin loss. What phenotypic changes would you hypothesize occur in the high-plaid genome?
- The authors address this topic later on in the paper. Before and after cohesin repression: 1% of inactive genes became active, 13% of active genes expression changed by ~2 fold. From these findings, the phenotypic contribution of cohesin-mediated loop domains is not immediately clear.
3. The plaid pattern in association to compartmentalization of loci with similar histone marks is natively present in native chromatin, and is increased in the absence of cohesin. Is this coagulation of similar histone regions mediated by the cell in order to promote favorable interactions, or are they conversely energetically favorable interactions that are inhibited by cohesin-mediated loop domain formation?
Citations:
- Cohesin Loss Eliminates All Loop Domains Cell, Vol. 171, No. 2. (8 October 2017), pp. 305-320.e24, doi:10.1016/j.cell.2017.09.026 by Suhas S. P. Rao, Su-Chen Huang, Brian Glenn St Hilaire, et al.
- Researchers Map Human Genome in 4-D as It Folds. Phys.org - News and Articles on Science and Technology, 6 Oct. 2017, phys.org/news/2017-10-human-genome-d.html.
Week 5: Chromosome topology as a key player in epigenetic regulation
Papers:
- Disruptions of Topological Chromatin Domains Cause Pathogenic Rewiring of Gene-Enhancer Interactions
- A CRISPR Connection between Chromatin Topology and Genetic Disorders
Topologically associated domains in the news:
Discussion Questions:
1. Diseases associated with limb malformations occur around the EPHA4 locus - but when the EPHA4 gene is knocked-out, there is no limb malformation phenotype. Why, then, has this locus been focused on to study?
- We took on this topic in the pre-journal club discussion. We learned that papers are not always presented in the same way or order that the experiments were done or designed. In this case, through their study, they found that the disrupted EPHA4 locus is associated with mis-regulation of genes near the locus, and that is associated with limb malformations. They present EPHA4 as a way of introducing the region of the genome that is the focus of the study, despite it not having been the catalyst for the project.
2. In they authors' introduction of TADs, they state that TADs are observed to exist regardless of transcriptional status. What is meant by this statement?
- Dr. Carbone took this to mean that various TADs in the genome exhibit variable expressivity of the genes in the associated chromatin region. The TAD structure encompass genes with high expressivity and genes with low expressivity; thus TADs are shown to exist regardless of the transcriptional status of the genes they contain, and this raises questions for how TAD structure informs gene expression.
3. How do you think haplotype variation contributes to studying TADs? What are possible experimental options for resolving haplotype variance in TAD measurement techniques?
Citations:
- Lupiáñez D.G., Kraft K., Heinrich V., Krawitz P., Brancati F., Klopocki E., Horn D., Kayserili H., Opitz J.M., Laxova R. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell. 2015;161:1012–1025.
- Bing Ren, Jesse R. Dixon, 2015, 'A CRISPR Connection between Chromatin Topology and Genetic Disorders', Cell, vol. 161, no. 5, pp. 955-957
Week 4: Revealing transcription in heterochromatin via PIWI RNA
Papers:
- A heterochromatin-dependent transcription machinery drives piRNA expression
- Molecular biology: Rhino gives voice to silent chromatin
Press Coverage:
This week, I presented!
- Follow this link to Revealing transcription in heterochromatin via PIWI RNA to see what I had to say about chromosome structure, transposons, PIWI RNAs, and the protein complexes that allow for their transcription
Citations:
- Peter Refsing Andersen, Laszlo Tirian, Milica Vunjak, Julius Brennecke.A heterochromatin-dependent transcription machinery drives piRNA expression.Nature, 2017; DOI:10.1038/nature23482
- Zamore, P. (2017). Molecular biology: Rhino gives voice to silent chromatin.Nature, 549(7670), pp.38-39.
- Institute of Molecular Biotechnology (IMBA). "How cells hack their own genes." ScienceDaily. ScienceDaily, 23 August 2017. <www.sciencedaily.com/releases/2017/08/170823134850.htm>.
Week 3: TZAP and its role in telomere length
Papers:
- TZAP: A telomere-associated protein involved in telomere length control
- TZAP or not to zap telomeres
Press Coverage:
Discussion Topics: Getting a handle on the jargon: This paper characterized TZAP in a number of specialized assays. Below, is clarifying information about key features of a few of these assays.
1. What is a Myc tag?
- It is a polypeptide strand of residues [Amino terminus]-EQKLISEEDL-[Carboxy terminus], or N-Glu-Gln-Lys-Leu-Ile-Ser-Glu-Glu-Asp-Leu-C, which has highly charged residues, which allows for isolation of the protein by affinity chromatography.

2. How does 2D gel electropheresis work?
- This experiment can help resolve replication unique non-linear DNA structures within your sample. The two dimensions separate DNA moieties by 1) size and 2) shape. Each arc represents a unique DNA moiety in your sample.

3. Reflecting on the paper: Pop science has regarded telomere regeneration as the "cure" to ageing. How does this paper inform that theory?
Citations:
- Li J. S., Miralles Fuste J., Simavorian T., Bartocci C., Tsai J., et al., 2017 TZAP: a telomere-associated protein involved in telomere length control. Science 355: 638–641.
- Lossaint G., Lingner J., 2017 TZAP or not to zap telomeres. Science. 2017 Feb 10;355(6325):578-579. doi: 10.1126/science.aam7015.
- Modano, Dan. "TZAP Protein Controls Telomere Trimming Process." ReliaWire, 13 Jan. 2017. Web. 12 Oct. 2017.
Week 2: Identifiers for the 21st century
Paper:
Discussion Questions:
1. The article states "It is incumbent on metaresolvers to be vigilant about detecting & updating their redirection rules in the face of provider changes." This implies that metaresolvers are responsible for maintaining the system of persistent identifier continuity. My question is: What insentives do they have to maintain this very important part of current science? Metaresolvers are independently formed companies who have corporate interests, so what prevents them from entering another market?
2. The paper discusses versioning as a means of recording project, data, and identifier evolution over time. Despite the importance of versioning, storing digital records of databases requires enormous server storage space, and therefore older versions are discarded. The paper lists the version expiration time as 5 years. Do you think that information vital to experimental research will be lost over time? Will this impede research efforts in 20, 50 or 100 years?
3. The abstract refers to a "Wringing value" which isn't discussed later in the paper. Is that a common term in this field and how is it characterized?
Critique: For examples in Table 1, it seems like incorrect identifiers, such as CURIEs, would lead science professionals to incorrect content which carries its own information which would alert the user. For instance, the example for the "Unambigous" characterisitic...
Citation:
- McMurry J, Juty N, et al. Identifiers for the 21st century: How to design, provision, and reuse persistent identifiers to maximize utility and impact of life science data. PLoS Biol. 2017 Jun 29;15(6):e2001414. doi:10.1371/journal.pbio.2001414. eCollection 2017 Jun.
Papers:
- Variation in cancer risk among tissues can be explained by the number of stem cell divisions
- Stem cell divisions, somatic mutations, cancer etiology, and cancer prevention
Discussion Questions:
1. The original paper stated "...only 5 to 10% of cancers have a heritable component..." (Tomasetti et al, 78). How do you think the proposed model of cancer risk, where 65% of the differences in cancer risk can be attributed to stem cell division rates, would change if the known contribution of heritable mutations to cancer was increased to 20, 30 or 40%?
2. Based on the debate that has arisen from the publication of this paper, many have expressed that it is irresponsible for these authors to publish such a dramatic model without clarifying the implications to cancer research or to cancer patients. The question I pose is: What do you think the responsibilities are for those who generate theoretical models?
3. If we were to accept the assumptions of this model and accept that 66% of cancer risk may be attributed to stem cell division: How how do you think cancer funding allocation should change? Do some cancer research approaches become less important in an earl-detection model of cancer risk?
Citations:
- Tomasetti C, Vogelstein, B. Cancer etiology. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science. 2015; 347: 78-81.
- Tomasetti C, Li L, Vogelstein B. Stem cell divisions, somatic mutations, cancer etiology, and cancer prevention. Science. 2017;355:1330–4. doi: 10.1126/science.aaf9011.