Chenlab: Difference between revisions
Kaifu Chen (talk | contribs) No edit summary |
Kaifu Chen (talk | contribs) No edit summary |
||
| Line 30: | Line 30: | ||
'''Recent news:''' | '''Recent news:''' | ||
March, | March, 2020: | ||
* Bo's software CECF to define cell identity regulators by machine learning of histone modifications was accepted in principle by '''Nature Communications'''. Cheers! | |||
March, 2020: | |||
* Guangyu's software TADsplimer for Hi-C analysis was accepted for publication by '''Genome Biology'''. TADsplimer revealed that there are always hundreds of TAD splits between cell types. We found these TAD splits are associated epigenetic regulation of cell identity gene expression. Cheers! | * Guangyu's software TADsplimer for Hi-C analysis was accepted for publication by '''Genome Biology'''. TADsplimer revealed that there are always hundreds of TAD splits between cell types. We found these TAD splits are associated epigenetic regulation of cell identity gene expression. Cheers! | ||
December, 2019: | December, 2019: | ||
Revision as of 19:09, 26 April 2020
Chen Laboratory
The Chen Lab is a system biology research group interested in computational modeling of cell identity regulation in development and diseases. Different cell types in a healthy body share the same genetic sequence. Difference in the identities of these cell types is determined by epigenetic mechanism. Meanwhile, many genetic variations in germ line or somatic cells were found to affect epigenetic factors, and can cause disease due to cell identity dysregulation. To study cell identity regulation in development, we are focused on endothelial lineage specification, and further work with collaborators to apply our knowledge to other cell lineages such as embryonic stem cell, cardiomyocyte, and neuron cells. To study cell identity dysregulation in diseases, the lab is focused on heart failure, and further works with collaborators to apply our knowledge to other diseases such as cancers, Rett Syndrome, and Hutchinson-Gilford Progeria Syndrome.
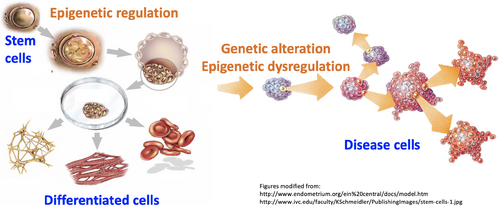
The most important discovery in the lab is that the cell identity genes as a unique group are distinct from other genes in the molecular mechanisms to regulate their expression (Nature Genetics 2015). Our findings further provide insights into the molecular regulation of normal cell identities in development (Science 2019), molecular regulation of genetic variations (Nature 2018), and dysregulation of cell identity in diseases, particularly in cancers (Molecular Cell 2018). These findings build a knowledge foundation for us to develop bioinformatics techniques for discovery of cell identity regulators (manuscript submitted) and cancer driver genes (manuscript submitted). These techniques have been proved to successfully recapture known cancer driver genes, and further, overcome critical constraint on current technologies by successfully identifying many cancer driver genes that were not detectable by conventional mutation analysis. These cancer driver genes led to new therapeutic targets and diagnostic markers as have been verified in cell, xenograft, and PDX models (Nature Communications 2018; IJC 2018; Oncogene 2019) and thus, will benefit numerous cancer patients.
Major research directions in the lab include:
- Develop novel bioinformatics techniques.
- Study epigenetic regulation of gene expression.
- Investigate molecular regulation of genetic variations.
Cell identity is regulated by a complex molecular network system that include many genetic, epigenetic, and environmental factors. The recent development of high throughput techniques to collect molecular and clinical data provided the unique opportunity for computational modeling of cell identity on the base of big data, of which the size is growing with an exponential speed.
Recent news:
March, 2020:
- Bo's software CECF to define cell identity regulators by machine learning of histone modifications was accepted in principle by Nature Communications. Cheers!
March, 2020:
- Guangyu's software TADsplimer for Hi-C analysis was accepted for publication by Genome Biology. TADsplimer revealed that there are always hundreds of TAD splits between cell types. We found these TAD splits are associated epigenetic regulation of cell identity gene expression. Cheers!
December, 2019:
- Kaifu served as a reviewer for the NIH TAG study section.
November, 2019:
- Dr. Ruli Gao joined our Center to start her independent lab for single-cell bioinformatics and genomics researches, welcome onboard!
October, 2019:
- Kaifu served as a reviewer for the NIH TAG study section.
September, 2019:
- Dr. Xinlei Gao who has just finished her B.S. and Ph.D from the University of Science and Technology of China will join us as a new postdoc very soon. Welcome Xinlei!
- Dr. Yanchun Zhang who has just finished her B.S. and Ph.D from the Nanjing University will join us as a new postdoc very soon. Welcome Yanchun!
- We will continue to recruit one additional postdoctoral Fellow with strong background in or extensive experiences with quantitative sciences (Statistics, Mathematics, Physices, Computer Sciences, or related area)
June, 2019:
- Kaifu served as a reviewer for the NIH GCAT study section.
May, 2019:
- Dr. Shuo Zhang from the University of Houston will join us as a new postdoc after receiving his PhD in September. Welcome Shuo!
January, 2019:
- Jie and Bo's paper in collaboration with the Fang and Bai labs is accepted to Science. Jie developed a new bioinformatics tool CAGREfor this project and successfully applied it to the genomic data interpretation. Congratulations!
December, 2018:
- Dr. Xin Wang joined the group as a new postdoc, welcome onboard!
- Dr. Trinh Tat joined the group as a new postdoc, she will receive joint postdoctoral training in the Kiss and Chen labs, welcome onboard!
- Guangyu's paper in collaboration with Qipeng and Qiaqia is accepted to the International Journal of Cancer. Cheers
October, 2018:
- Bo's paper in collaboration with Yang under supervision of Greg was accepted to Nature. Bo developed a new bioinformatics tool FAID for this project and successfully applied it to the genomic data interpretation. Bo is a co-first author, Guangyu is a co-author, and Kaifu is the co-coressponding author, Cheers!
- Dongyu received the Moran Foundation Award, a $2,000 addition to paycheck. Congratulations!
September, 2018:
- Kaifu received the Houston Methodist Career Cornerstone Award. Cheers!
August, 2018:
- Bo's paper for aging research based on Ribo-Seq data in collaboration with Zheng was accepted to Elife. Cheers!
- Dr. Min Zhang joined the group as a new postdoc, welcome onboard!
July, 2018:
- Kaifu was assigned as the founding Director to develop a Center for Bioinformatics and Computational Biology. The center will thrive to a size of 4-6 faculties in the following 5 years.
- Dr. Yanqiang Li joined the group as a new postdoc, welcome onboard!
June, 2018:
- Kaifu was promoted to Associate Professor, many thanks to hard workings of the fantastic team in the past 3 years!
- Kaifu received an Houston Methodist Award For Excellence in Peer-reviewed Publication for the discovery of the key role for MLL4 in establishing Broad H3K4me3 and Super enhancers at tumor suppressor genes (Zhao, et al, Molecular Cell 2018)
May, 2018:
- Dr. Yiwen Bu, who published a paper in Nature Cell Biology very recently, joined the group as a new postdoc, welcome onboard!
April, 2018:
- Dongyu's paper with Shilpa et al got accepted to Molecular Cell, the 19th paper published by the new lab, cheers!
- Kaifu received an Houston Methodist Award For Excellence in Peer-reviewed Publication for the discovery of the a novel role for BMI1 in regulating AR signaling in prostate cancer (Zhao, et al, Nature Communications 2018)
Dec, 2017:
- Dongyu's paper in collaboration with Sen et al was accepted to Nature Communications, the 16th paper published by the new lab, Cheers!
September, 2017:
- Dr. Guangyu Wang joined the group as a new postdoc, welcome onboard!
May, 2017:
- Houston Methodist is named one of “America’s Best Employers” by Forbes magazine for the second year in a row.
- Dr. Lei La will be a visiting scholar in the lab for 1 year. Dr. La is an Associate Professor at the Southern Medical University Nanfang Hospital ( a top hospital in China). Welcome!
April, 2017:
- Kaifu provided a talk in the EB / ASBMB annual meeting 2017.
- Dongyu's first-authorship manuscript for prostate cancer research is submitted, fingers crossed!
- Dongyu's first-authorship manuscript for brain cancer research is submitted, fingers crossed!
March, 2017:
- Kaifu become a member of the NCI-Funded Interdisciplinary Approaches to Cancer Metastasis Workshop, and will work together with the workshop to develop highly innovative and collaborative projects on cancer metastasis.
- A manuscript is accepted by Elife. Alin and Kaifu help the collaborators interpret the 17 sets of RNA-Seq data. Cheers!
February, 2017:
- A manuscript is accepted by the Nature Chemical Biology journal, Jie conducted the RNA-Seq analysis for this project. Cheers!
January, 2017:
- Dr. Guangyu Wang from Beijing Institute of Genomiocs will join the lab as a postdoc soon, welcome!
- Ms. Karen Lai from UT Austin will be an undergraduate intern in the lab this summer, welcome!
December, 2016:
- A manuscript is accepted by the Science journal, Kaifu helps supervising the MNase-Seq and ChIP-Seq analysis and is a co-author. Cheers!
November, 2016:
- Alin will start his tenure-track faculty position (assistant professor of statistics) at the Manhattan College. Congratulations!
- Bo has joined the lab. Welcome onboard!
September, 2016
- A manuscript is accepted to the Journal of the American Heart Association, Jie and Kaifu are co-authors, cheers!
- A manuscript is accepted to the Nucleic Acids Research, Kaifu is a co-author, cheers!
- Dr. Qingshu Meng will be visiting the lab for 1 year. Dr. Meng has outstanding background in Genomics research, and is a group leader at Beijing Institute of Genomics. Welcome!
August, 2016
- Kaifu will be an adjunt assistant professor of Texas A&M.
- Dr. Bo Xia from Georgia Tech will join the lab as a fresh postdoc soon. Dr. Xia is a PhD of biology and Master of computer science. Welcome!
April, 2016
- Jie's paper was accepted by Molecular Cell, congratulations!
April, 2016
- New postdoc position opening.
- Jie's co-first authorship manuscript is under revision at Molecular Cell (IF 14), fingers crossed!
December, 2015
- Kaifu's new manuscript was accepted to the Nucleic Acids Research, thank Alin for his help with the revision work!
- A manuscript was accepted to the Molecular and Cellular Biology, thank Dongyu for his help with the revision work!
November, 2015
- Dr. Jie Lu has joined the lab as postdoctoral Fellow, welcome onboard!
October, 2015
- Dr. Dongyu Zhao has joined the lab as postdoctoral Fellow, welcome onboard!
September, 2015
- The Broad H3K4me3 research work is highlighted by the Cancer Discovery journal (IF 20.26)
- Dr. Alin Tomonaga has joined the lab as a postdoctoral Fellow, welcome onboard!
August, 2015
- Kaifu's data-revisiting paper is accepted to Nature Genetics.
- Kaifu's MeCP2 mCH-binding paper (with Dr. Zoghbi lab) has been accepted to PNAS.