PersonalGenomes@Home
|
PurposeThe Personal Genome Project (PGP) will publicly sequence the DNA of 100.000 volunteers and make the results freely available. The analysis of the raw data entails a huge computational effort and will require much computational power. The PGP is an open source project. In this spirit we would like the data analysis to become an open source community effort through distributed computing with 'PersonalGenomes@Home'. The part of the data analysis that will be initially distributed to the community is the analysis of raw Illumina images to produce reads and alignment against a reference. We are currently setting up the open source software packages Swift and BOINC as well as Tranche and Free Factories. CollaboratorsRunning SwiftSwift is designed to process a single tile of Illumina data at a time. It can be run in two modes:1) base-caller only (processes intensity files produced by the Solexa pipeline) and 2) image analysis. The output files are purity-filtered and non purity-filtered reads in fastq format, a run report, an intensity file and files listing all processed images.
Processing PGP DataAL - After verifying the functionality of the code with example data (available through Swift), we have run Swift on tile 87 of one PGP2 data set. Images are 7.1MB in size and the processing resources required exceed the abilities of AL's personal computer (1.5MB RAM). Swift is designed to process one full tile of Illumina data at a time. Attempts to process less than 36 cycles remain unsuccessful. Keeping in mind that the goal is to perform the data analysis on a 'regular' computer, we did not move to larger memory capacities. Instead, the large images were cropped into three horizontal thirds of 2.4MB each using ImageJ, an open source image processing tool. Images overlapped by 10 pixels to avoid errors caused by edge effects. The one tile was then individually processed in three sub-tiles using Swift in approximately 20 minutes.
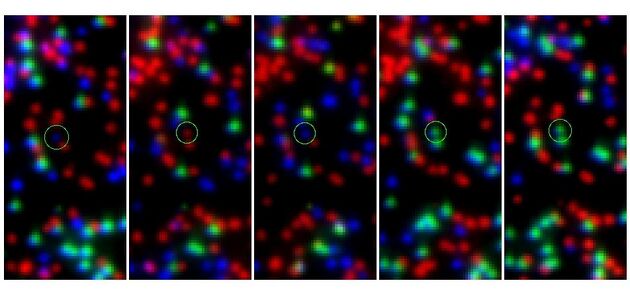
Finding the MFN2_R364W Variant and ControlTo prove that Swift produces correct results, the stack 87 data were searched for the MFN2_R364W variant and stack 27 for the control. The last numbers are the x and y coordinates on the images. Differences in aligning techniques between the Illumina pipeline and Swift result in minor discrepancies in coordinate assignment. Previous analysis had shown a T in the 11th cycle: The Swift analysis confirms this result: Analogous, the control sequence was extracted from stack 27. Previous analysis had shown a C in the 13th cycle: The Swift analysis confirms this result:
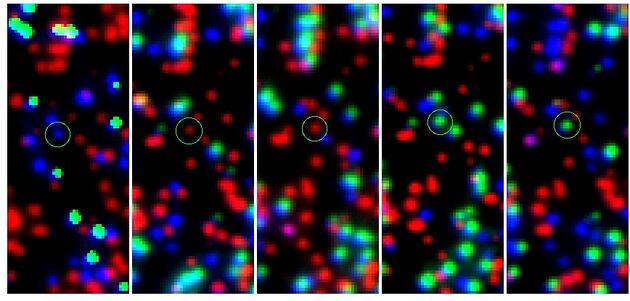
Swift on Free FactoriesWe have installed and run the Swift software on two virtual machines running on the http://bio.freelogy.org clusters at the Church lab. These nodes are intended as a production/development environment for coordinating the distributed computing effort. We plan to use the BOINC software. For now, we have automated the data analysis using a shell script, that downloads all images from PGP2 data set using wget, crops them into thirds using ImageMagick and runs Swift on the entire data set. This analysis has been run with version 140 of Swift. With a new quality score recalibration in version 149, we reprocessed the entire data set. The reads produced by Swift for PGP2 data set can be found below. PGP2 Sample DataWe currently have raw images, two sets of processed reads, and draft assemblies (from each read-set against both the whole HG18 reference assembly and against just the annotated capture regions) for about 10% of the exome of PGP2. We also have Affymetrix 500K data (binary) w/ two sets of calls for this participant. These are summarized below:
The above information is preliminary. The current release is here.
PGP2 Sample Data AnalysisAfter using Swift to produce reads, AWZ ran Mega BLAST to get alignments. Gapped AlignmentsFirst, we were interested in finding all gapped alignments including their read ID. AL wrote a perl script which reads the original Mega BLAST output and inserts the read ID before each line beginning with a '>', which marks an alignment. Gapped alignments are marked 'Gaps', and a simple grep -B5 -A7 'Gaps' can extract all gapped alignments including the read ID they are associated with. The results are available here: PGP2 Gapped Alignments. This is an older data product from Swift and reads are only 35 base pairs in length. Unique AlignmentsWe are also interested in finding all reads with a unique alignment. Query= pgp2_bot_stack_100:121:0
Query= pgp2_bot_stack_100:121:0
(35 letters)
The order of relevant information is: read ID read ID n chromosome numbers (i.e. n lines starting with a number) read ID >chromosome number 13 lines of information per chromosome. For a unique alignment there will only be one line starting with a number, and we need to extract the 13 lines following it. To extract the unique alignments, AL processes the file line by line with a perl script: read and save ID
The Script (oh boy!)#!/usr/bin/perl -w use strict; use Getopt::Long; my $input="reads_plusid"; my $output="reads_plusid_unique_aligns"; my $id="foo"; my $suche="bar"; my $counter=0;
ProblemsSpeed: grep on the chromosome number is not unique; a chromosome can be aligned to several reads; therefore the grep is piped into another grep for the ID associated with the unique read as found before. The input file has 1,602,962,364 lines, the double grep does not help with speed. In tests, 10^7 lines are processed in 10 minutes, which extrapolates to 26 hours for 1,602,962,364 lines ... if line starts with '>' && previous line contains ID
grep -B1 -A12 line > output
False alarms: I have created a subset of the data consisting of 10^7 first lines of the input data (using 'head'). And while the script does find all the unique alignments, there are false alarms: one read is flagged as 'unique', but the output contains several alignments associated with it. In this subset of data, this happened for only one read. Stats37GB of input data contain 1,602,962,364 lines.
Alignment with BowtieBowtie is an ultrafast, memory-efficient short read aligner. It aligns short DNA sequences to the human genome at a rate of over 25 million 35-bp reads per hour. Bowtie indexes the genome with a Burrows-Wheeler index to keep its memory footprint small: typically about 2.2 GB for the human genome. See also: Bowtie paper in Genome Biology. We have used bowtie with its default settings to align the PGP2 data from the Swift analysis described above. Each 1GB stack was processed in 50 seconds. Results can be found here: PGP2 Bowtie Alignments |
