Lauren M. Magee Week 12
From OpenWetWare
Jump to navigationJump to search
Powerpoint Slides
Using YEASTRACT to Infer which Transcription Factors Regulate a Cluster of Genes
In the previous analysis using STEM, we found a number of gene expression profiles (aka clusters) which grouped genes based on similarity of gene expression changes over time. The implication is that these genes share the same expression pattern because they are regulated by the same (or the same set) of transcription factors. We will explore this using the YEASTRACT database.
- Open the gene list in Excel for the profile/cluster that you analyzed for the Week 11 Assignment.
- Copy the list of gene IDs onto your clipboard.
- Launch a web browser and go to the YEASTRACT database.
- On the left panel of the window, click on the link to Rank by TF.
- Paste your list of genes from your cluster into the box labeled ORFs/Genes.
- Check the box for Check for all TFs.
- Accept the defaults for the Regulations Filter (Documented, DNA binding plus expression evidence)
- Do not apply a filter for "Filter Documented Regulations by environmental condition".
- Rank genes by TF using: The % of genes in the list and in YEASTRACT regulated by each TF.
- Click the Search button.
- Answer the following questions:
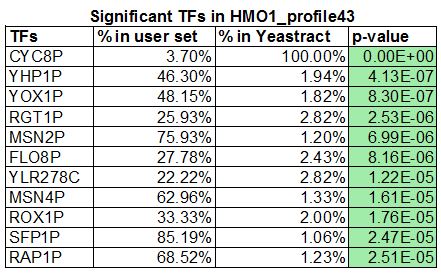
- In the results window that appears, the p values colored green are considered "significant", the ones colored yellow are considered "borderline significant" and the ones colored pink are considered "not significant". How many transcription factors are green or "significant"?
- 11 genes are significant
- List the "significant" transcription factors on your wiki page, along with the corresponding "% in user set", "% in YEASTRACT", and "p value".
- Are CIN5, GLN3, HMO1, and ZAP1 on the list?
- No the above genes are not included on the list.
- In the results window that appears, the p values colored green are considered "significant", the ones colored yellow are considered "borderline significant" and the ones colored pink are considered "not significant". How many transcription factors are green or "significant"?
- For the mathematical model that we will build in class, we need to define a gene regulatory network of transcription factors that regulate other transcription factors. We can use YEASTRACT to assist us with creating the network. We want to generate a network with approximately 15-30 transcription factors in it.
- You and your partner will need to analyze the same gene regulatory network for your modeling project. Compare the lists of "significant" factors that you and your partner generated.
- How many of the transcription factors appear in both of your lists?
- SFP1P, YHP1P, YOX1P, YLR278C, MSNP2, and MSN4P, a total of 6 common transcription factors.
- You will use these transcription factors and add CIN5, GLN3, HMO1, and ZAP1 if they are not in your list. If the overlap in the lists between you and your partner does not add up to the 15-30 factors required, use your discretion to add transcription factors from either of your lists (the non-overlapping ones) until you reach a list of 15-30 factors. Explain in your electronic notebook how you decided on which transcription factors to include. Record the list and your justification in your electronic lab notebook.
- Go back to the YEASTRACT database and follow the link to Generate Regulation Matrix.
- Copy and paste the list of transcription factors you identified (plus CIN5, GLN3, HMO1, and ZAP1) into both the "Transcription factors" field and the "Target ORF/Genes" field.
- We are going to generate several regulation matrices, with different "Regulations Filter" options.
- For the first one, accept the defaults: "Documented", "DNA binding plus expression evidence"
- Click the "Generate" button.
- In the results window that appears, click on the link to the "Regulation matrix (Semicolon Separated Values (CSV) file)" that appears and save it to your Desktop. Rename this file with a meaningful name so that you can distinguish it from the other files you will generate.
- Repeat these steps to generate a second regulation matrix, this time applying the Regulations Filter "Documented", "Only DNA binding evidence".
- Repeat these steps a third time to generate a third regulation matrix, this time applying the Regulations Filter "Documented", DNA binding and expression evidence".
Analyzing and Visualizing Your Gene Regulatory Networks
We will analyze the regulatory matrix files you generated above in Microsoft Excel and visualize them using GRNsight to determine which one will be appropriate to pursue further in the modeling.
- First we need to properly format the output files from YEASTRACT. You will repeat these steps for each of the three files you generated above.
- Open the file in Excel. It will not open properly in Excel because a semicolon was used as the column delimiter instead of a comma. To fix this, Select the entire Column A. Then go to the "Data" tab and select "Text to columns". In the Wizard that appears, select "Delimited" and click "Next". In the next window, select "Semicolon", and click "Next". In the next window, leave the data format at "General", and click "Finish". This should now look like a table with the names of the transcription factors across the top and down the first column and all of the zeros and ones distributed throughout the rows and columns. This is called an "adjacency matrix." If there is a "1" in the cell, that means there is a connection between the trancription factor in that row with that column.
- Save this file in Microsoft Excel workbook format (.xlsx).
- Check to see that all of the transcription factors in the matrix are connected to at least one of the other transcription factors by making sure that there is at least one "1" in a row or column for that transcription factor. If a factor is not connected to any other factor, delete its row and column from the matrix. Make sure that you still have somewhere between 15 and 30 transcription factors in your network after this pruning.
- For this adjacency matrix to be usable in GRNmap (the modeling software) and GRNsight (the visualization software), we need to transpose the matrix. Insert a new worksheet into your Excel file and name it "network". Go back to the previous sheet and select the entire matrix and copy it. Go to you new worksheet and click on the A1 cell in the upper left. Select "Paste special" from the "Home" tab. In the window that appears, check the box for "Transpose". This will paste your data with the columns transposed to rows and vice versa. This is necessary because we want the transcription factors that are the "regulatORS" across the top and the "regulatEES" along the side.
- The labels for the genes in the columns and rows need to match. Thus, delete the "p" from each of the gene names in the columns. Adjust the case of the labels to make them all upper case.
- In cell A1, copy and paste the text "rows genes affected/cols genes controlling".
- Now we will look at some of the network properties. Again, repeat these steps for each of the three gene regulatory matrices you generated above. See this file for an example of how to do the following instructions.
- Create a new worksheet and call it "degree". Copy and paste your adjacency matrix from the "network" sheet into this new worksheet.
- In the first empty cell in column A, type "Out-degree". In the cell to the right of that in Column B, type the equation
=SUM(and select the range of cells in column B that has 1's and 0's in it, close the parentheses, and press Enter. This quantity is the number of genes that the transcription factor in that column is controlling, or the out-degree. Copy and paste that equation across all of the columns. - In Cell 1 of the first empty column to the right of the adjacency matrix, type "In-degree". In Cell 2 of this column, type the equation
=SUM(and select the entire row of 1's and 0's, close the parentheses, and press Enter. This quantity is the number of transcription factors that regulate the gene in that row, or the in-degree. Copy and paste the equation down the entire column, including the row that contains the out-degree sums. - The number in the lower right-hand corner, the sum of sums, is the total number of edges in the adjacency matrix. We would like to see about 50 (40-60 or so) edges in the matrix. If the matrix is too dense, it will slow down the modeling program because it will be difficult to estimate the parameters in the model.
- We want to plot the degree distributions for each of your gene regulatory networks. In the "degree" worksheet, create three columns to the right called "Frequency", "In-degree total", and "Out-degree total". In the "Frequency" column, number sequentially from 1 to the largest degree number in your calculations above. In the "In-degree total" column, type the number of genes with that in-degree for each of the frequencies. In the "Out-degree total" column, type the number of genes with that out-degree for each of the frequencies.
- Select the "Frequency", "In-degree total", and "Out-degree total" columns. Go to the "Insert" tab and select the column chart type to insert a plot of the degree distribution. Copy and paste the charts for each gene regulatory matrix into your PowerPoint presentation.
- Now we will visualize what these gene regulatory networks look like with the GRNsight software.
- Go to the GRNsight home page (you can either use the version on the home page or the beta version, which has slightly different visualization properties).
- Select the menu item File > Open and select one of the regulation matrix .xlsx file that has the "network" worksheet in it that you formatted above. If the file has been formatted properly, GRNsight should automatically create a graph of your network. Move the nodes (genes) around until you get a layout that you like and take a screenshot of the results. Paste it into your PowerPoint presentation. Repeat with the other two regulation matrix files. You will want to arrange the genes in the same order for each screenshot so that the graphs can be easily compared.
Guiding Questions
- Determining candidate transcription factors that regulate a cluster of genes from your dataset.
- Gene Profile #43 produced the most significant transcription gene factors when entered into Yeastract. With the addition of CIN5, GLN3, HMO1, and ZAP1 there were just fifteen genes for us to include in our gene regulatory network. The following genes were found to be present in both the HMO1 Profile #43 and the Wild-Type: SFP1P, YHP1P, YOX1P, YLR278C, MSNP2, and MSN4P, so we will be using these as a mode of comparison between the two.
- Creating three candidate gene regulatory networks.
- Using Yeastract we were able to create three candidate gene regulatory networks: DNA binding plus expression evidence, DNA binding and expression evidence, and DNA binding only.
- Determining the total number of edges and degree distribution of your three gene regulatory networks.
- Visualizing the networks.
- The networks for the HMO1 gene aren't as complex as I assume the other model in our class may be. There was only about 70, 30, and then 7 edges in our three networks. With the deletion of genes that had rows and columns of all 0's, we ended up with less contributors to our model.
- Choosing a particular gene regulatory network to pursue for the modeling.
- It was clear that the DNA binding plus expression evidence was the best model, because it featured the most transcription factors (15) and edges (about 70) out of all of our gene regulatory networks.
- Week 1
- Week 2
- Week 3
- Week 4
- Week 5
- Week 6
- Week 7
- Week 8
- Assignment Cancelled
- Week 9
- Week 10
- Week 11
- Week 12
- Week 13
- Week 14
- Week 15