IGEM:IMPERIAL/2007/Projects/Biofilm Detector/Modelling
Infector Detector: Modelling
Overview of Modelling
Welcome to our Portal Page for the modelling of Infector Detector.
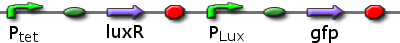
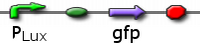
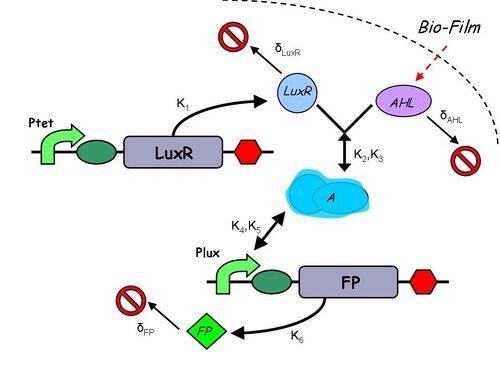
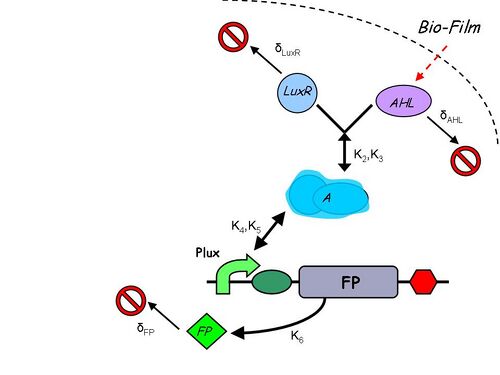
Infector Dectector (ID) is based on the Quorum Sensing Pathway and our aim in modelling of ID is to determine the concentration of AHL in biofilm we can detect such that we report a visible signal .We are looking at two constructs to emulate the quorum sensing pathway:
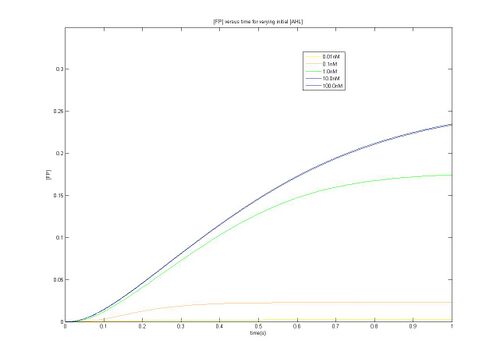
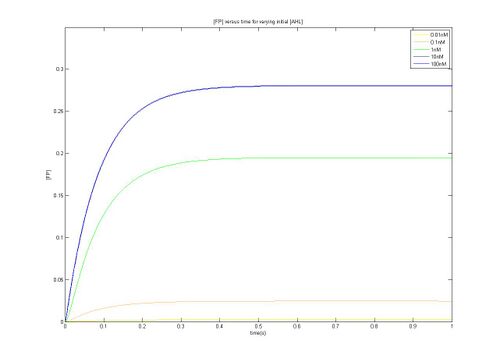
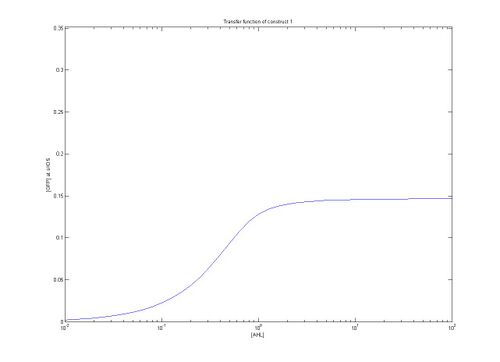
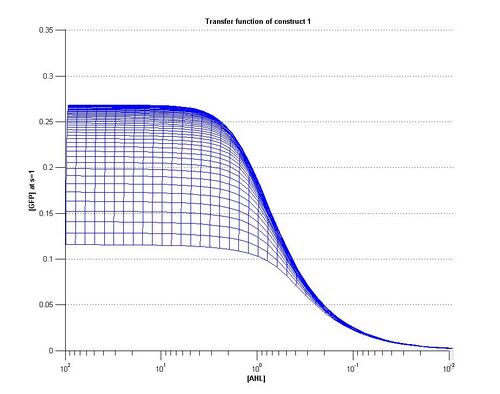
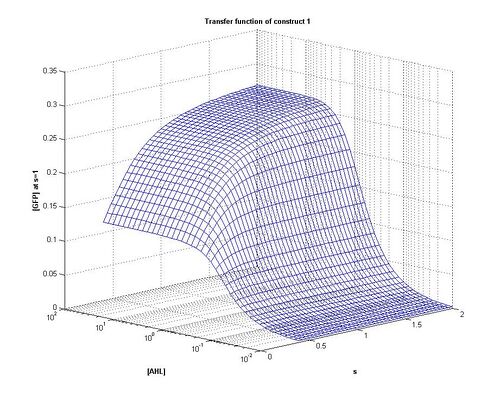
- As can be seen from the above plots, construct 1 takes longer to reach steady state [FP], meaning that over the same time period it reaches a lower maximum value
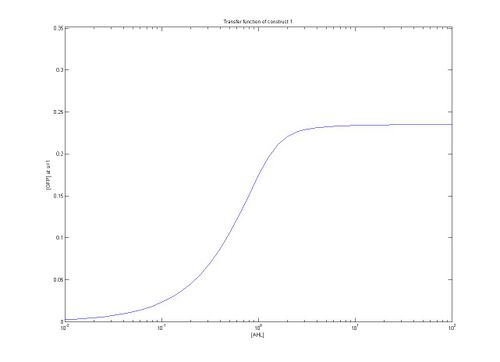
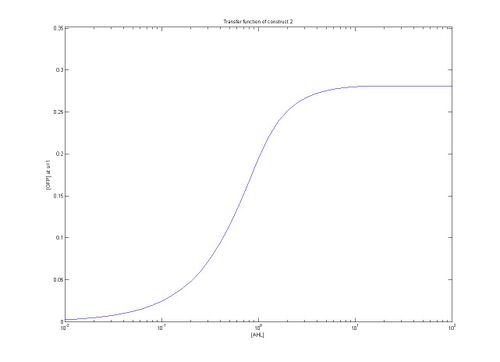
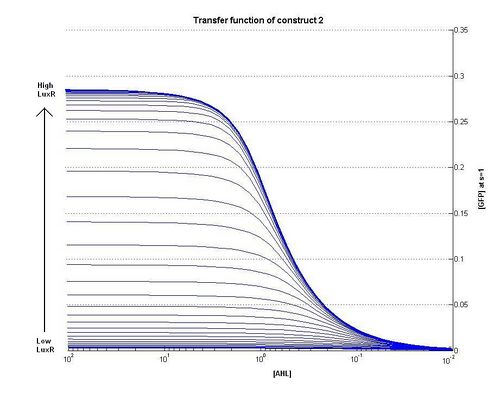
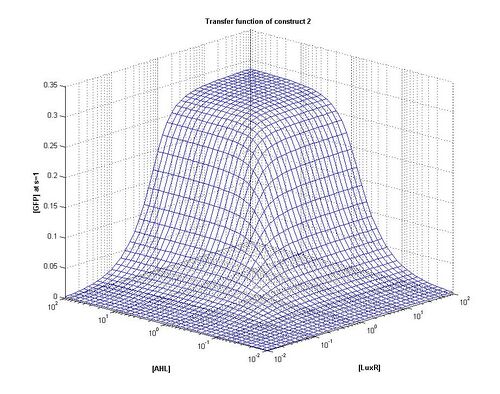
As can be seen from the plots above the the threshold moves according to the level of LuxR. Looking at our plots we can see that at time, s=0.5 the threshold is different for construct 1 to construct 2
17.08.07 Modelling General Concerns
For protocols to figure out :
- Degradation terms:GFP,LuxR,AHL - want expt to find delta as a function of chassis
- If we put protease inhibitors in the mixture and can we assume negligible degradation for GFP and luxR, not sure about AHL have to look in literature for that.
- What is the visual threshold of [GFP] - want expt to find this
- Do you need to do it for GFP or just for the final reporter used, which is dsRED
- What is the concentration of promoters - is this chassis dependant ?
- You know - the weight of DNA added, and the mass of each plasmid. Knowing that there is only one promoter on each plasmid, you can calculate the concentration of promoters.
- Activation/Response Time of Plux (F2620)?
- You might want to check the part F2620, not sure if the information is useful, or valid for in vitro.
(Protocols) Construct 1 specific:
- What is the lifespan of the whole system?
- Can we get steady-stateof LuxR? - When will this happen ? - Before cell dies ?
- Having reached steady state is there enough E left to express [GFP]
(Protocols) Construct 2 specific:
- Can we obtain purified LuxR to be injected into System - protocol for prep of LuxR
- Protease inhibitors available?
- Yes, in homemade extract. For commercial extract, still waiting for reply from promega.
- How long will it take to get construct 2 ready ?
- Week 8, earliest (unfortunately)
For modelling to figure out :
- K-values for rxn pathway
- k5/k4 = 5*10-10M [source: pmid=17400743]
- Response of Biofilm : Is [AHL] constant ?
- Simulation of [LuxR] vs. Time
Concentration of LuxR for construct 2
At equilibrium:
[math]\displaystyle{ [A][P] = K_D[AP] }[/math]
Let us assign initial concentrations as [A0] and [P0]
We want to have n% of the promoters bound, thus [AP] = n[P0]
[math]\displaystyle{ ([A_0]-n[P_0])([P_0]-n[P_0]) = nK_D[P_0] }[/math]
[math]\displaystyle{ [A_0]-n[P_0] = \frac{n}{1-n}K_D }[/math]
[math]\displaystyle{ [A_0] = \frac{n}{1-n}K_D+n[P_0] }[/math]
Substituting 0.1nM for KD and 0.1nM for [P0], and let n be 95%...
The concentration of A0 required is 2nM.
Knowing that at eqm:
[math]\displaystyle{ [AHL][LuxR] = K_D[A_0] }[/math]
[math]\displaystyle{ ([AHL_0]-[A_0])([LuxR_0]-[A_0]) = K_D[A_0] }[/math]
[math]\displaystyle{ [LuxR_0] = \frac{K_D[A_0]}{[AHL_0]-[A_0]} + [A_0] }[/math]
The concentration of AHL0 would be 50nM, KD would be 1nM.
The concentration of LuxR0 required is ~3nM.
Note that the KD of AHL-LuxR is only an estimate and may be incorrect.