Biomod/2012/DNA Maestros:Results

Home Introduction The Project Materials & Methods Results and Discussion Conclusion Team Members Acknowledgements
Validation of the reporter assay for strand displacement on the Icosahedron
Self quenching of fluorophores occurs when two or more fluorophores come closer than a specific distance[1]. A simple situation where self quenching is observed is when the concentration of the fluorescent molecules in solution is high. The proposed mechanism for such an observation is the formation of an intermolecular ground state dimer complex[2], which makes one of the fluorophores in the complex act as a donor and the other as an acceptor. The greater the spectral overlap between the excitation and the emission spectrum of a fluorophore, the higher is the observed self quenching, as seen in cases of homo-FRET[3]. Fluorescein has been extensively studied for concentration dependent self-quenching[4] and we wanted to use such a self-quenching assay as a reporter for strand displacement if it occurred on the DNA icosahedron.
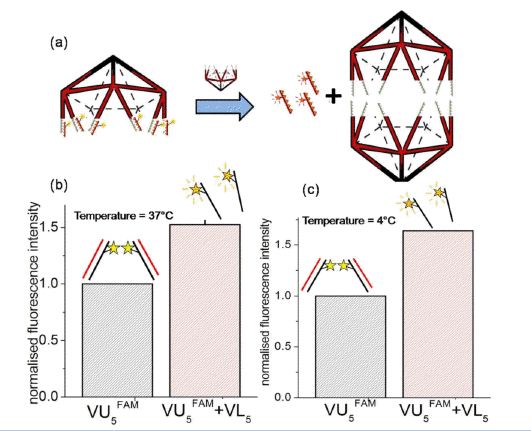
We wanted to use the property that VU5 has 10 identical overhangs, all within a small volume of ~500 nm3. Thus, if multiple FAM (Fluorescein) labelled 10-mer oligomers (FAM-10-mer) were to be hybridized to VU5, this would result in an increased local concentration of the FAM label, creating a self-quenched state for the FAM fluorescence. If one were to now somehow displace the ten FAM-10-mer molecules hybridized to each VU5, the FAM-10-mer molecules would be released in bulk solution which would increase the intermolecular distance between the FAM labels and thereby result in increased fluorescence.
In order to see if this assay worked, we created a complex of VU5 hybridized to ten FAM-10-mer molecules, called VU5FAM (Figure 4a). We first decided to displace the ten FAM-10-mer oligos on VU5FAM by introducing the lower half icosahedron (VL5) whose overhangs are complementary to the upper half icosahedron, VU5FAM in the solution containing 100 nM added TMR as an internal control. We incubated VL5 with VU5FAM at 37°C. Due to the cooperative nature of complex formation by 1:1 VL5 and VU5FAM the major product of the reaction is the full icosahedron[5]. Importantly we observed an increase in the fluorescence intensity of the FAM label upon complexation with VL5 (Figure 4b). Importantly, the intensity of the internal control, 100 nM TMR, remains unchanged whether VU5FAM was complexed to VL5 or not. This increase in fluorescence intensity due to the formation of complete icosahedron confirmed that the displacement of the ten FAM-10-mer oligos was indeed successful. More importantly, we see that the percentage of fluorescence quenching that occurred due to the hybridization of the FAM-10-mer oligos on to VU5 was 38%. Given that the R0 of Fluorescein is 4.2 nm[6], this suggests that the distance between adjacent FAM oligos onVU5FAM is 4.6 nm. The theoretically expected distance as obtained from a model of the icosahedron currently being built in our host laboratory, turns out to be 4.7 ± 3 nm. The same experiment was also conducted at 4°C to minimize possible contributions to the fluorescence due to temperature-induced dissociation of FAM-10-mer oligos from VU5FAM (Figure 4c). These relative changes in fluorescence intensities obtained at 37°C and 4°C are similar, confirming that the contribution to fluorescence intensity due to any pre-melted FAM oligos is negligible.
Aptamer recognition of cyclic-di-GMP causes strand displacement on VU5apt
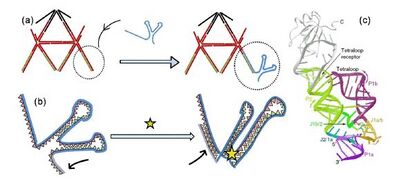
In order to create an icosahedron that could be split into two halves by the additon of cdGMP, we had to first make an icosahedron where each of the ten links that hold VU5 and VL5 could dissociate upon addition of cdGMP. In order to achieve this, we created an upper half icosahedron VU5apt that was modified at each of its ten single stranded overhangs with the cdGMP aptamer. We modified constituent oligonucleotides U2 and U3 of the Uapt 5WJ such that the 10 bases at their 5´ end are complementary to the 10 bases at the 3´ end of the cdGMP aptamer (Figure 5).
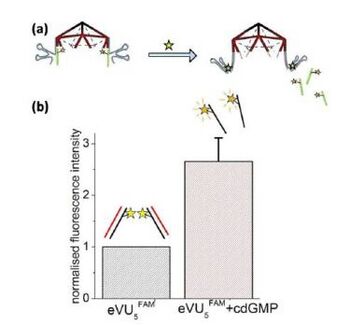
The cdGMP aptamer is actually part of a riboswitch where it is embedded in a longer sequence that forms an expression platform[8]. Although the conformational rearrangement is well documented, and structures of the free and cdGMP bound states of the cdGMP aptamer are well characterized, whether the 3’ end can actually bring about strand displacement is unknown. In order to test this hypothesis we designed an experiment similar to that described in Figure 4. Here, the ten overhangs of the upper half icosahedron modified with the RNA aptamer (VU5apt), are also hybridized to an analogous 5' FAM functionalized 10 bp strand (FAM-10-mer-b) to give eVU5FAM. In the presence of 100 μM cdGMP, the ten aptamer domains on eVU5FAM would undergo a conformational change. If this conformational change resulted in the displacement of the ten FAM-10-mer-b molecules on eVU5FAM, due to increase in intermolecular distance between solution-dispersed FAM labels, we should observe. an increased in the fluorescence intensity of FAM. Importantly we observe that this is indeed the case, with the fluorescence intensity increasing nearly three-fold relative change in the fluorescence intensity upon cdGMP addition. This also suggests that the local environment provided by the cdGMP aptamer in eVU5FAM could further quench the FAM labels on hybridized FAM-10-mer-b molecules in addition to the self-quenching effect.
Cyclic-di-GMP triggered release of encapsulated cargo from a DNA icosahedron mediated by strand displacement
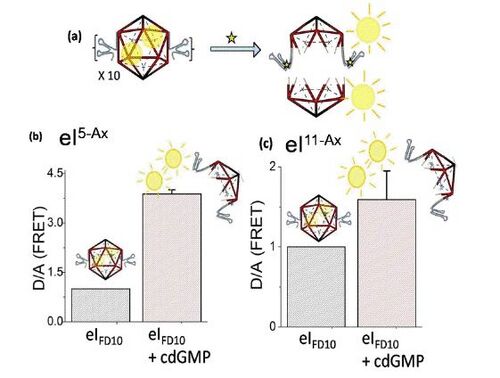
Encapsulation of cargo within a DNA icosahedron was done by assembling the engineered, small-molecule sensitive icosahedron in the presence of excess of cargo molecules. Choosing a fluorescently labelled cargo for encapsulation allowed the monitoring of the release process. In this case, FITC-Dextran of molecular weight 10 kDa (FD10) was encapsulated within the icosahedron and the cargo-loaded icosahedra were purified from free FD10 by adapting a previously published protocol that is also described in detail here[9]. The encapsulated FD10 can act as donor fluorophores in a FRET experiment if there exists proximal acceptor fluorophores, say on the body of the DNA icosahedron. Fluorescein and Alexa 647 is a well-documented FRET pair[10]. Therefore the aptamer-modified, FD10 loaded, icosahedral DNA scaffold was also functionalized with five Alexa 647 fluorophores at well-defined locations to give eIFD105AX.
If the encapsulated FD10 undergoes a resonance energy transfer with Alexa-647 fluorophores, then the release of FD10 molecules into the bulk solution upon addition of cdGMP, should cause an increase in fluorescence of the FAM labels on FD10. It has been shown that, for this particular polyhedron and FD10, encapsulation results in an average of two FD10 molecules per DNA icosahedron[9]. According to the manufacturer’s specifications, on average one FD10 molecule carries 2-3 FAM molecules. Thus the system has 5-6 donor molecules and 5 acceptor molecules which make it suitable for FRET. When fluorescence of eIFD105AX was recorded in absence and presence of 100 µM cdGMP, it was observed addition of cdGMP caused a significant enhancement of the Donor to Acceptor Fluorescence intensity (D/A) ratio (Fig 7). This is consistent with the decrease in donor fluorescence on FD10 molecules due to their proximity to the Alexa 647 functionalized DNA icosahedron, that is subsequently lost due to their release into the bulk solution upon addition of cdGMP. This resultant loss of FRET is reflected in the gain of donor fluorescence. This was recapitulated on a completely distinct FD10- loaded, cdGMP sensitive DNA icosahedron carrying eleven Alexa fluorophores on the DNA scaffold, namely eIFD1011AX.
This indicates that addition of cyclic di GMP, that binds to the RNA aptamer of eI FD10 complexes causes a conformational change that results in a strand displacement reaction that displaces the cognate lower half of the icosahedron from its engineered upper half. This splits the icosahedron into its two constituent halves that in turn releases its encapsulated cargo.
References
-
Zhuang, X., Ha, T., Kim, H. D., Centner, T., Labeit, S. & Chu, S. Fluorescence quenching: A tool for single-molecule protein-folding study. Proc. Natl. Acad. Sci. USA 97, 14241-14244 (2000)
-
Kim, T. W., Park, J. H. & Hong, J. I. Self-quenching Mechanism: the Influence of Quencher and Spacer on Quencher-fluorescein Probes. Bull. Korean Chem. Soc. 28, 1221-1224 (2007)
-
Gautier, I., Tramier, M., Durieux, C., Coppey, J., Pansu, R.B., Nicolas, J.-C., Kemnitz, K. & Coppey-Moisan, M. Homo-FRET Microscopy in Living Cells to Measure Monomer-Dimer Transition of GFP-Tagged Proteins. Biophys. J. 80, 3000–3008 (2001)
-
Lakowicz, J. R. in Principles of Fluorescence Spectroscopy, Vol 1, Third Ed., Springer, p 69.
-
Bhatia, D., Mehtab, S., Krishnan, R., Indi, S.S., Basu, A. & Krishnan, Y. Icosahedral DNA nanocapsules via modular assembly. Angew. Chem. Int. Ed. 48, 4134-4137 (2009).
-
A. Kawski, Excitation energy transfer and its manifestation in isotropic media, Photochem. Photobiol. 38, 487–508 (1983)
-
Smith, K. D., Lipchock, S. V., Ames, T. D., Wang, J., Breaker, R. R. & Strobel S. A. Structural basis of ligand binding by a c-di-GMP riboswitch. Nat. Struct. Mol. Biol. 16, 1218-1223
-
Sudarsan, N., Lee, E. R., Weinberg, Z., Moy, R.H., Kim, J. N. , Link, K. H. & Breaker R. R. "Riboswitches in eubacteria sense the second messenger cyclic di-GMP. Science 321 411–3 (2008).
-
Bhatia, D., Surana, S., Chakraborty, S., Koushika, S.P. & Krishnan, Y. A synthetic, icosahedral DNA-based host-cargo complex for functional in vivo imaging. Nat. Commun., 2, 339 (2011)
-
Bradshaw, J. M., Kwan, J. K., Ling, A. E. H. & Shaw, D. FRET-based binding assay. Apr, http://www.freepatentsonline.com/8153390.html (2012)

