Biomod/2012/DNA Maestros:Project

Home Introduction The Project Materials & Methods Results and Discussion Conclusion Team Members Acknowledgements
Project Idea
In a nutshell, we have developed a strategy to effect the controlled release of encapsulated cargo from a DNA polyhedron triggered by the binding of a small molecule. The mechanism of the cargo release is generic and step-wise in nature and it depends solely on the predictive and programmable nature of Watson-Crick base pairing. In this project design, there are three major components: the DNA polyhedron which is an icosahedron, the cargo which is FITC-Dextran and the molecular trigger which is cyclic di-GMP (cdGMP) along with its aptamer. The subsequent sections justify the choice of each of these modules in our design
Project Design
The Icosahedron
For any DNA polyhedron to be used for cargo delivery, there are two key requirements: the polyhedron should have maximum encapsulation volume and minimum pore size. Encapsulation volume directly translates into the ‘cargo-loading’ capacity of the nanocage, and the pore-size sets a cut-off on types of material that can be encapsulated. Among all the platonic solids, the icosahedron satisfies both of these criteria to the best possible extent. Also, due to its modular assembly out of halves, it is the only platonic solid which that has been experimentally shown to possess the capacity to entrap free floating cargo from bulk solution[1, 2]. Apart from these properties, the icosahedron possesses minimal amount of DNA and therefore has considerable potential to be used for any future biological applications. Further, due to its modular nature of construction, it is easy to achieve predictable modifications by incorporating minor alterations in the sequences of the constituent 5WJs. Lastly, the yield of the icosahedron is near quantitative, and encapsulation is a highly efficient process. Hence we chose this polyhedron assembled as described by Bhatia et al in 2009[1].
The cargo - FITC Dextran
For our encapsulation and release studies, 10 kDa FITC-Dextran (FD10) has been used as the molecular cargo. Many studies have also characterised FD10 as a biocompatible pH responsive molecule for in-cellulis studies[3, 4]. FD10 incorporates 2-3 molecules of FITC on an average, and the biophysical properties of the FAM label are unaltered. In previous studies published in the host laboratory, successful encapsulation of FD10 within the icosahedron has been demonstrated, without any alteration in the fluorescence properties[5]. Fluorescein, FITC or the FAM label has been extensively studied in literature for its fluorescence properties – it has been shown to be capable of self-quenching, as well as intermolecular FRET[6, 7, 8]. We can use these properties to investigate cargo encapsulation and release using different fluorescence based assays.
The chemical trigger and release mechanism
The basic engineering of the icosahedron is the incorporation of a small-molecule sensitive RNA aptamer. The small molecule we chose was cyclic di-GMP (cdGMP). Apart from conveniences like easy commercial availability, cdGMP is a bioactive, biocompatible, cell-permeable molecule. We used an RNA aptamer for cdGMP which is a part of its cognate riboswitch in V. Cholerae[9].
Most importantly, the aptamer we chose has been shown to have extremely tight binding for cdGMP (Kd≤10nM)[10], and its bound conformation has been elucidated in a crystal structure[10]. Therefore, the structural determinants of cdGMP binding have been characterized to atomistic detail. This provides us a great advantage to integrate the aptamer to the icosahedron using rational design. Though it is known that the aptameric region of the cdGMP riboswitch undergoes a conformational change upon binding cdGMP, and that the bound form is structurally characterized, it is not known whether the binding event could be transduced to achieve a strand displacement.
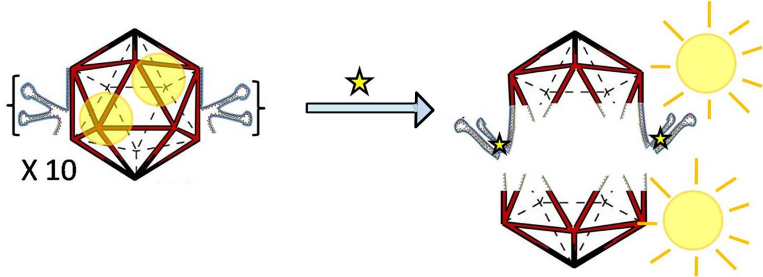
Here we demonstrate that integrating the cdGMP aptamer with a DNA architecture preserves its binding to cdGMP, preserves the architecture and encapsulation abilities of the icosahedral modules, and is also capable of mediating a strand displacement upon addition of cdGMP. All of these have been integrated to create a cdGMP responsive DNA icosahedron that releases its contents by dissociating in the presence of a small molecule.
References
-
Bhatia, D., Mehtab, S., Krishnan, R., Indi, S.S., Basu, A., Krishnan, Y. (2009) Icosahedral DNA nanocapsules via modular assembly. Angew. Chem. Int. Ed. 48, 4134-4137.
-
Bhatia, D., Chakraborty, S., Mehtab, S. and Krishnan, Y. (2012) A method to encapsulate molecular cargo within DNA icosahedra. Methods Mol. Biol. in press.
-
Calamita, HG; Ehringer, WD; Koch, AL; et al. (2001) Evidence that the cell wall of Bacillus subtilis is protonated during respiration. Proc. Natl. Acad. Sci. USA 98, 15260-15263.
-
Bayer, N; Schober, D; Prchla, E; et al. (1998) Effect of bafilomycin A1 and nocodazole on endocytic transport in HeLa cells: Implications for viral uncoating and infection. J. Virol. 72, 9645-9655.
-
Bhatia, D., Surana, S., Chakraborty, S., Koushika, S.P. and Krishnan, Y. (2011) A synthetic, icosahedral DNA-based host-cargo complex for functional in vivo imaging. Nat. Commun. 2, 339
-
Egawa, Y.; Hayashida, R.; Seki, T. et al. (2008) Fluorometric determination of heparin based on self-quenching of fluorescein-labeled protamine Talanta 76, 736-741.
-
Fudala, R., Mummert, M. E.; Gryczynski, Z.; et al. (2012) Lifetime-based sensing of the hyaluronidase using fluorescein labeled hyaluronic acid J. Photochem. Photobiol. 106, 69-73.
-
Bradshaw, J. M., Kwan, J. K., Ling, A. E. H. & Shaw, (2012) D. FRET-based binding assay. Apr, http://www.freepatentsonline.com/8153390.html
-
Smith KD, Lipchock SV, Ames TD, Wang J, Breaker RR, Strobel SA. (2009) Structural basis of ligand binding by a c-di-GMP riboswitch. Nat. Struct. Mol. Biol. 16(12): 1218-23.
-
Kulshina, N., Baird, N.J., & Ferré-D'Amaré, (2009) A.R. Recognition of the bacterial second messenger cyclic diguanylate by its cognate riboswitch. Nat. Struct. Mol. Biol. 16, 1212-1217.

